Clear Sky Science · en
Linker histone H1 represses H3 tail acetylation induced by H4 tail acetylation and alters its dynamics
Why tiny protein tails matter for our DNA
Inside every cell, nearly two meters of DNA are packed into a microscopic nucleus. This feat of folding is handled by spools of proteins called histones, which help decide whether genes are switched on or off. The new study explored how small chemical tags on these histones, and the presence of another protein called linker histone H1, work together to either open up DNA for gene activity or keep it locked down. Understanding this balance sheds light on how cells maintain long‑term memory of which genes should stay active or silent.
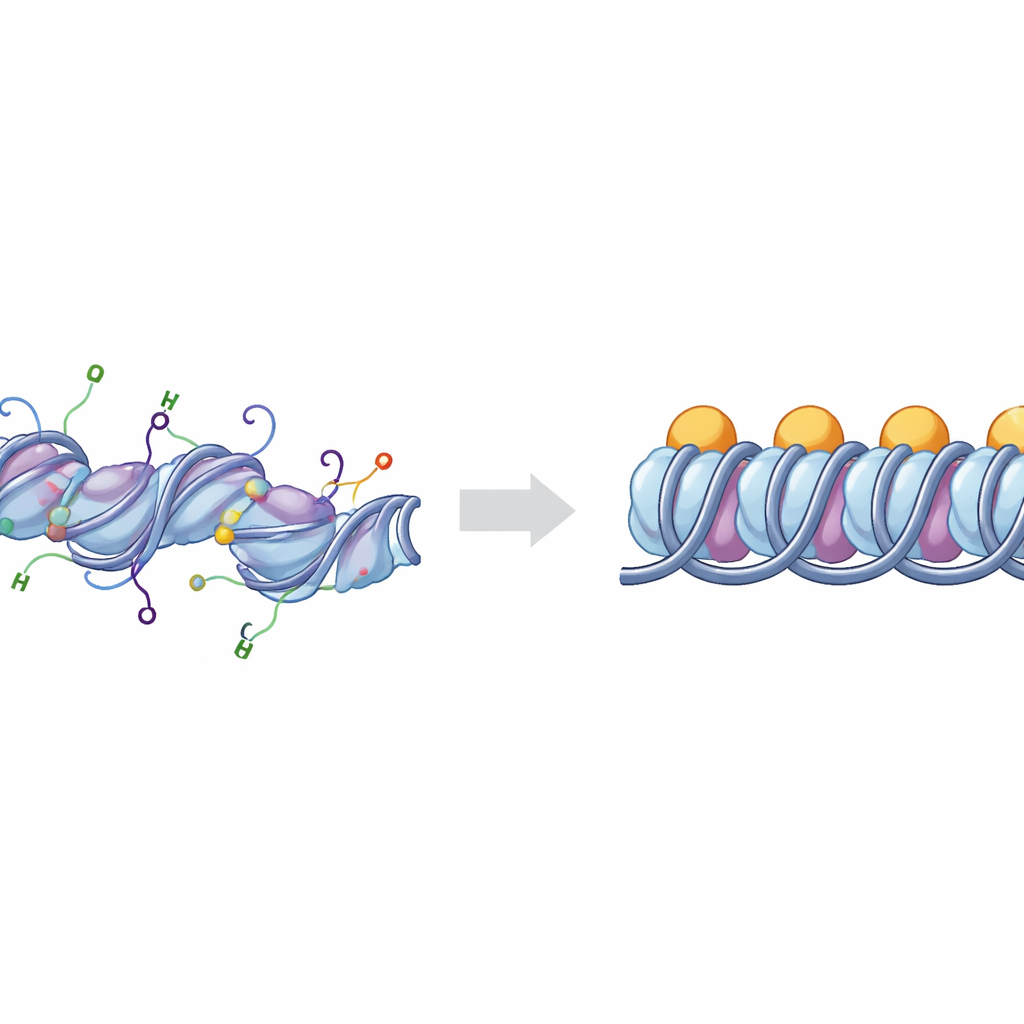
The basic building blocks of DNA packing
DNA in our chromosomes is wound around clusters of histone proteins to form units known as nucleosomes. When an extra protein, linker histone H1, binds at the points where DNA enters and exits the nucleosome, the structure is called a chromatosome. Groups of chromatosomes can stack and fold to create tightly packed regions of DNA that are generally less active. Cells fine‑tune this packing with chemical marks placed on flexible “tails” that extend from the histones. Two of the most important tails belong to histones H3 and H4, whose chemical changes are strongly linked to whether DNA regions behave as active, open "euchromatin" or silent, compact "heterochromatin."
How one tail’s change affects another
Earlier work showed that adding four acetyl groups to the H4 tail—at positions known collectively as H4‑4Kac—tends to make the neighboring H3 tail easier to acetylate as well, but that was studied mainly without linker histone H1. The new research asked what happens when H1 is present, a more realistic situation in living cells. Using cryo‑electron microscopy, the authors solved high‑resolution structures of chromatosomes carrying the acetylated H4 tail. Surprisingly, the overall shape of the core DNA and histone proteins remained almost unchanged compared with unmodified chromatosomes. The H1 protein still sat at the same central position, and the wrapped DNA looked nearly identical, suggesting that any important effects must come from changes in how the floppy histone tails move and touch the DNA.
Following moving parts with simulations and magnets
To capture these fast, subtle motions, the team combined computer simulations with nuclear magnetic resonance (NMR) spectroscopy. Coarse‑grained molecular dynamics simulations modeled how the H3 and H4 tails brush against the wrapped DNA and the extra stretches of DNA that link nucleosomes together. In chromatosomes, the presence of H1 naturally makes the two H3 tails behave differently: one tends to touch mainly the core‑wrapped DNA, while the other also contacts the exposed linker DNA. When H4 is acetylated, its own tail peels away from the DNA, becoming more flexible and solvent‑exposed. This release slightly shifts how both H1’s long tail and the H3 tails explore the DNA surface, but the simulations suggested only modest average changes in contact patterns.
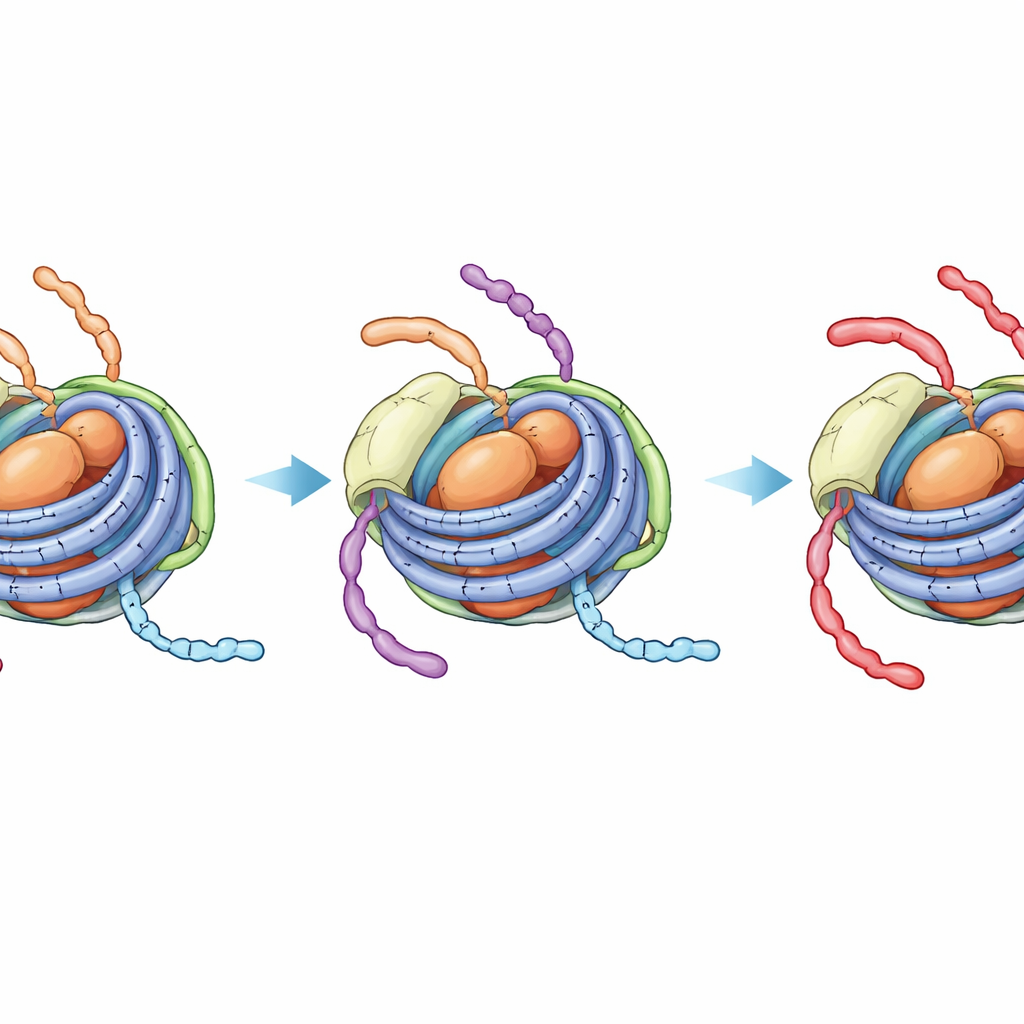
A hidden, tighter state that blocks further change
NMR spectroscopy, which senses the local environment of specific atoms, revealed a more striking effect. In unmodified chromatosomes, signals from the H3 tail show a mixture of two dynamic states—one resembling the more open nucleosome (with contact to linker DNA) and one resembling the tighter core‑only nucleosome particle. Over the timescale of an enzyme reaction, these states interchange, so the enzyme that acetylates position K14 on the H3 tail can still access linker‑exposed segments. In contrast, when the H4 tail carries four acetyl groups and H1 is present, the H3 tail signals collapse into a single, robust state that looks like the tightly bound core‑DNA form. Direct NMR monitoring of the acetylation reaction confirmed that H3 K14 is then modified far more slowly, at a rate nearly indistinguishable from the closed nucleosome core without any linker DNA at all.
What this means for gene on/off switches
Taken together, the results suggest that acetylation of the H4 tail, in the presence of linker histone H1, paradoxically helps to lock the neighboring H3 tails against the core DNA rather than freeing them for further change. By promoting this stable, DNA‑hugging state and greatly slowing additional acetylation of H3, H1 appears to counteract the spread of open, active chromatin and instead favor more compact, silent regions. In everyday terms, the study shows that one set of chemical marks and a single extra protein can decide whether nearby stretches of DNA remain open for business or get folded away for long‑term storage, adding a new layer of understanding to how cells preserve their identity over time.
Citation: Furukawa, A., Echigoya, K., Blazquez, S. et al. Linker histone H1 represses H3 tail acetylation induced by H4 tail acetylation and alters its dynamics. Commun Biol 9, 496 (2026). https://doi.org/10.1038/s42003-026-09926-y
Keywords: chromatin structure, histone acetylation, linker histone H1, nucleosome dynamics, epigenetic regulation