Clear Sky Science · en
Endogenous protein tagging coupled with a CRISPR screening approach identifies UBE3C as a potential MYC oncogene regulator
Why this study matters
Cancer doctors have long known that a powerful "master switch" protein called MYC helps tumor cells grow, divide, and resist treatment. In multiple myeloma, a cancer of antibody‑producing blood cells, MYC is often stuck in the "on" position. But directly blocking MYC has proven technically difficult. This study takes a different tack: instead of trying to drug MYC itself, the researchers systematically searched for other genes that control how much MYC a myeloma cell makes. Their work reveals new molecular levers—especially a protein called UBE3C—that might be easier to target with future therapies.
Turning MYC into a living light sensor
To watch MYC inside cancer cells without disturbing their normal wiring, the team engineered a multiple myeloma cell line so that its own MYC protein carries a built‑in green fluorescent tag. In these cells, the brighter the green signal, the more MYC is present. Because the tag is fused to the cell’s natural MYC, its activity and control remain as close as possible to real disease biology. The authors carefully checked that this fluorescent MYC still behaves like the original protein: it responds to a known MYC‑lowering drug, can be reduced by cutting the MYC gene with gene‑editing tools, and continues to interact with its usual binding partner in the nucleus. This validated cell line became the basis for their large‑scale search.
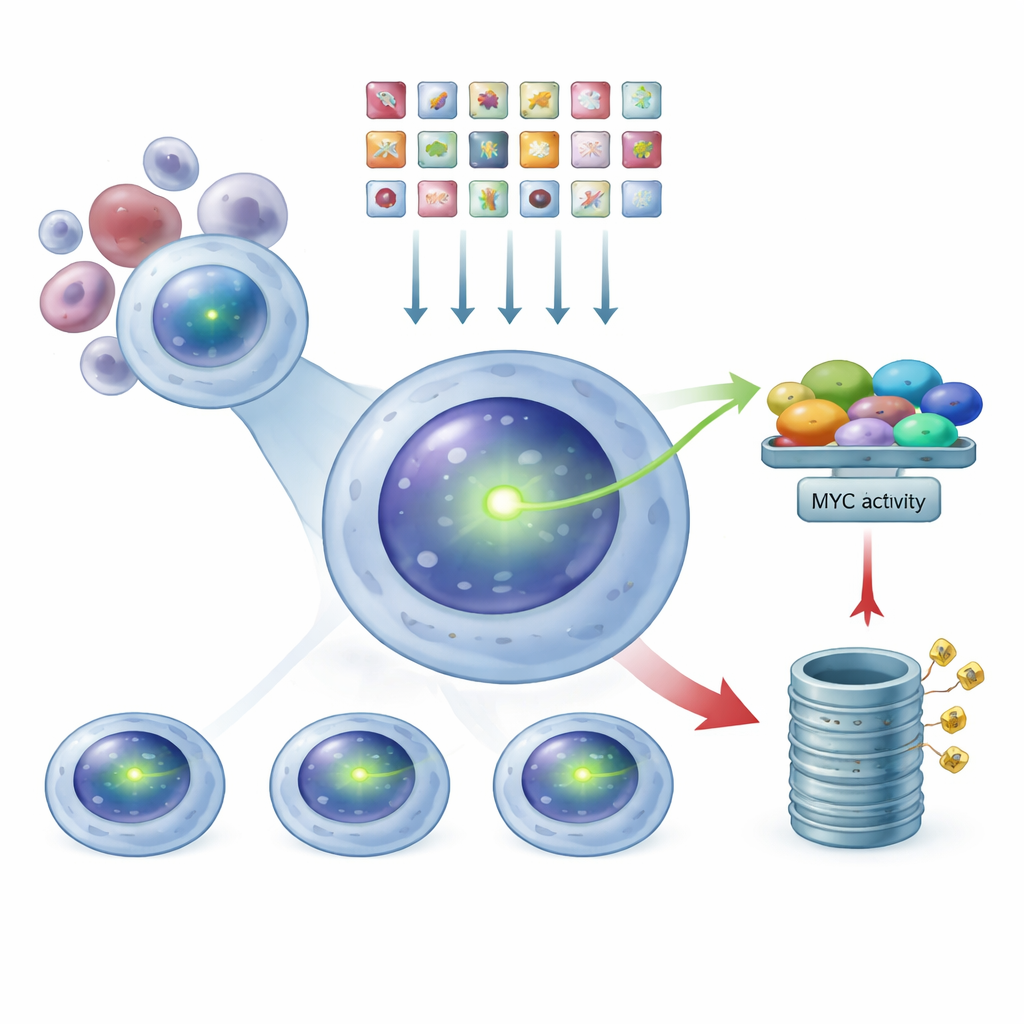
Using CRISPR as a genome‑wide dimmer switch
Armed with this glowing reporter system, the researchers used CRISPR technology as a kind of genome‑wide dimmer switch. They introduced a library of tens of thousands of guide RNAs, each designed to knock out one human gene, into the MYC‑tagged myeloma cells. Each cell received, on average, just one genetic hit. After allowing time for these edits to take effect, they used a cell‑sorting machine to separate the population into three groups based on green brightness: MYC‑low, intermediate, and MYC‑high. By sequencing which guide RNAs were enriched in each group, they could work backwards to infer which knocked‑out genes caused MYC levels to fall or rise.
Two main control hubs: transcription and protein disposal
The pattern that emerged from this screen revealed two major classes of MYC regulators. When components of a large molecular assembly called the Mediator complex were disrupted, MYC levels dropped, placing these factors in the "activator" camp—they normally help cells turn the MYC gene on. In contrast, when parts of the cell’s protein‑shredding machinery, the proteasome, or enzymes that attach or remove small degradation tags were lost, MYC levels climbed. This indicates that MYC is normally kept in check by active breakdown, not just by controlling how much is made. Known players such as IRF4, which boosts MYC, and FBXW7, which promotes its destruction, showed up exactly where expected, giving confidence that the screen was capturing genuine biology rather than noise.
Zooming in on UBE3C as a specific MYC brake
Among the many candidates, one stood out: UBE3C, an enzyme that helps label proteins for destruction and transiently associates with the proteasome. Knocking out UBE3C caused a striking rise in MYC levels in the reporter cells, far stronger than related enzymes UBE3A and UBE3B, which had little effect. The authors then tested unmodified multiple myeloma cell lines and found that disabling UBE3C there also boosted native, untagged MYC protein. This points to UBE3C as a particularly important negative regulator of MYC in myeloma cells. At the same time, the study suggests that MYC’s abundance is shaped jointly by how hard the transcriptional machinery pushes its gene and how actively the protein is marked and fed into the cellular "recycling bin."
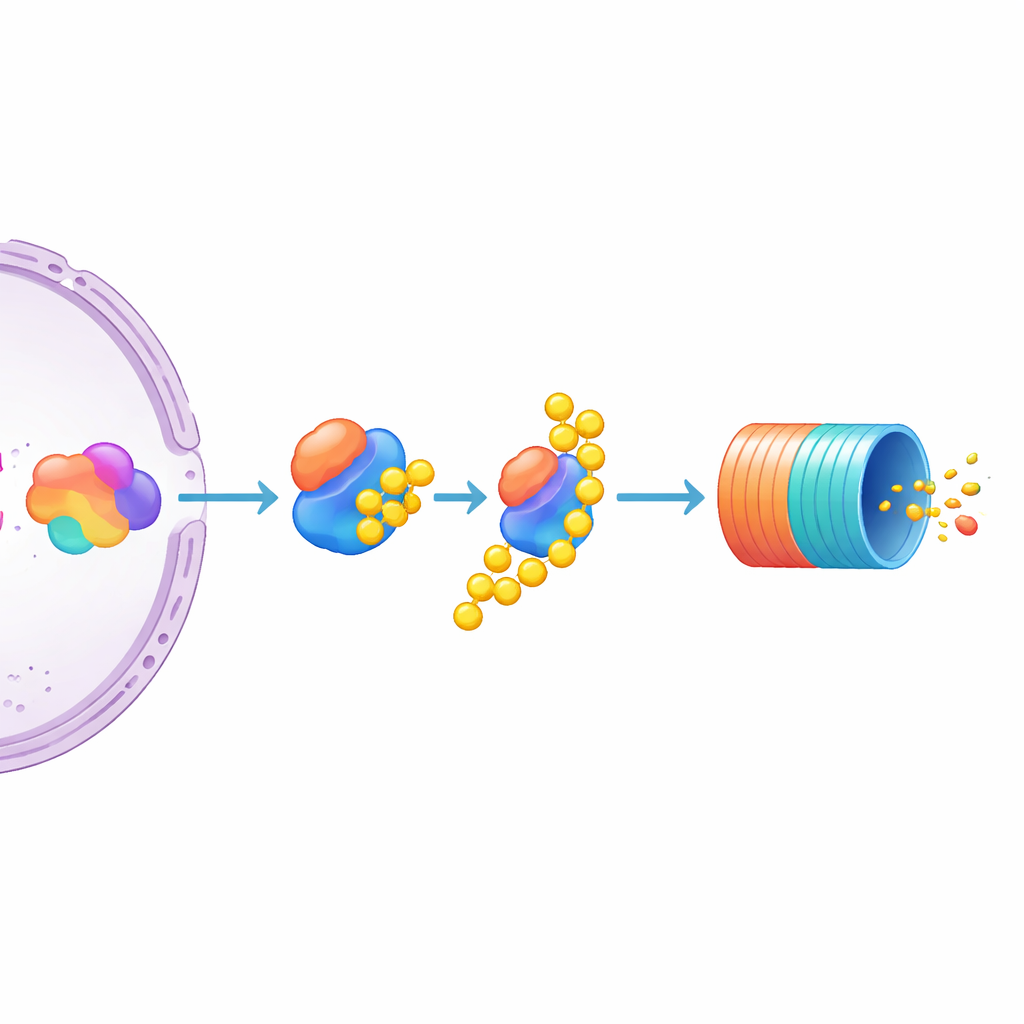
What this means for future cancer treatments
For non‑specialists, the key takeaway is that MYC itself may remain a slippery drug target, but the network that controls it is becoming clearer. By converting MYC into a fluorescent reporter and pairing it with a genome‑wide CRISPR search, the authors built a powerful map of genes that either turn MYC up or down inside multiple myeloma cells. Their data highlight Mediator components as part of the engine that drives MYC production and, more strikingly, identify UBE3C as a likely switch that helps keep MYC protein from accumulating. In the long run, drugs that modulate regulators like UBE3C—or other steps in MYC’s production and disposal—could offer more precise ways to quiet this master switch and slow down MYC‑driven cancers without needing to hit MYC head‑on.
Citation: Seibert, M., Kurrle, N., Kaleab, S. et al. Endogenous protein tagging coupled with a CRISPR screening approach identifies UBE3C as a potential MYC oncogene regulator. Sci Rep 16, 12194 (2026). https://doi.org/10.1038/s41598-026-47974-w
Keywords: MYC regulation, multiple myeloma, CRISPR screen, ubiquitin proteasome, UBE3C