Clear Sky Science · en
Membrane editing with proximity labeling reveals regulators of lipid homeostasis
How Cells Keep Their Fatty Coats in Balance
Every cell in your body is wrapped in a thin, fatty coat called a membrane. These membranes are more than simple barriers: their exact mix of lipids helps control how cells grow, communicate, and respond to stress. One especially powerful lipid, phosphatidic acid (PA), acts like a molecular alarm bell. Bursts of PA can switch key signaling pathways on or off, so cells must raise and lower its levels very quickly and very precisely. This study reveals how cells sense sudden surges of PA and recruit specific proteins to bring this potent lipid back under control.
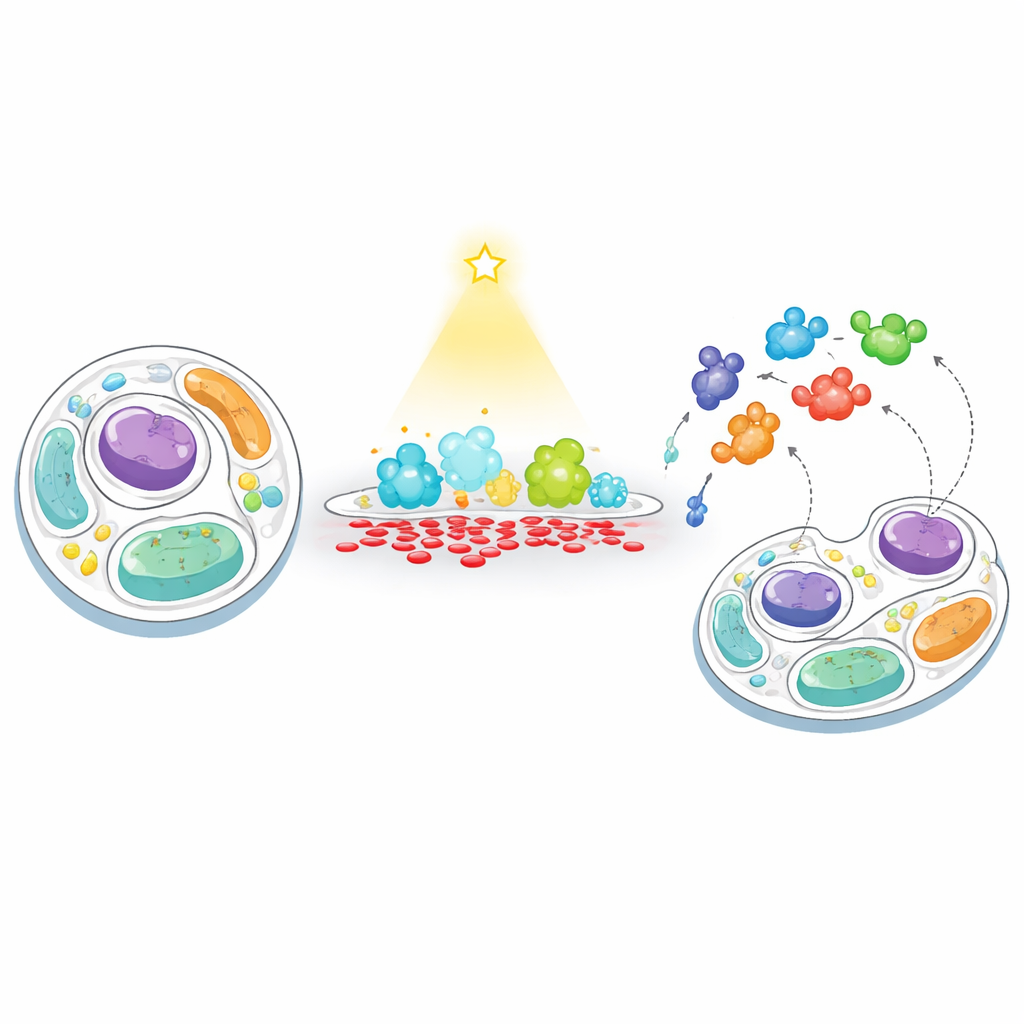
A New Way to “Feed” and “Fish” in Cell Membranes
The researchers developed a two-step strategy they call “feeding–fishing” to track how cells respond when a membrane’s PA level is artificially pushed out of balance. First, they “feed” a chosen membrane with PA using a light-controlled enzyme, an engineered version of phospholipase D that can be switched on at will with blue light. By targeting this enzyme to either the cell’s outer surface or to lysosomes (the cell’s recycling centers), they could trigger short, localized bursts of PA production. Second, they “fish” for nearby proteins by attaching a biotin-tagging enzyme (TurboID) to the same membrane. Any protein that wanders near gets tagged and later captured and identified by high-throughput mass spectrometry, revealing which proteins are drawn to, or repelled from, PA-rich regions.
Who Shows Up When PA Spikes?
Applying this method to human cells, the team identified thousands of proteins near the plasma membrane and lysosomes, then pinpointed those whose abundance changed when PA levels rose. Many of the strongest responders were already known to handle PA or related lipids, validating the approach. Enzymes called lipins, which break down PA into another signaling lipid, diacylglycerol (DAG), were enriched on PA-loaded membranes, while a DAG-making enzyme that produces PA showed up less often, hinting at a built-in feedback system. The best-known PA transfer protein, Nir2, also moved to both PA-rich plasma membranes and lysosomes, underscoring its central role in shuttling PA between organelles. The analysis further revealed changes in proteins linked to mitochondria and cholesterol handling, suggesting that PA surges ripple across many aspects of cell metabolism.
Hidden Couriers That Move and Remove PA
To test which recruited proteins actually help clear excess PA, the scientists focused on several lipid transfer proteins—molecular couriers that move fats between membranes. By overproducing these couriers and tracking a fluorescent PA sensor, they found that Nir2 and a protein called SCP2 markedly reduced PA buildup at the plasma membrane and lysosomes. SCP2, previously known for moving cholesterol and fatty acids, also showed PA-carrying activity in a purified liposome assay, expanding its job description. Other transfer proteins, including PDZD8 and TEX2, did not directly lower PA levels but appeared to tighten physical contacts between the endoplasmic reticulum and lysosomes, likely helping route lipids for later processing. In contrast, an oxysterol-binding protein, ORP1L, unexpectedly caused PA to rise further, in a way that depended on its cholesterol-binding site, hinting at a more complex cross-talk between PA and cholesterol flows.
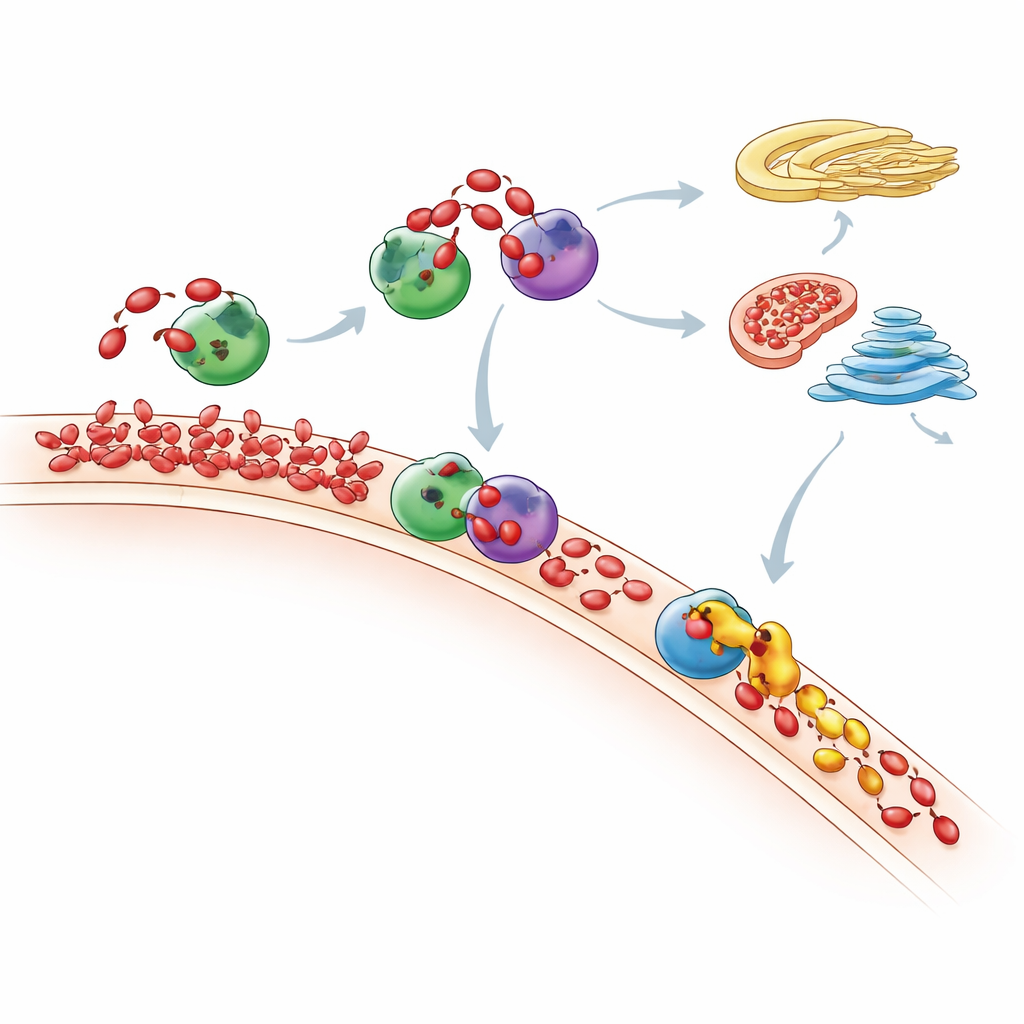
Where Extra PA Ultimately Goes
Measuring the full lipid landscape with sensitive mass spectrometry, the researchers tracked how the cell’s entire phospholipid mix changed after PA “feeding.” They saw that PA spikes are short-lived: over time, many PA molecules are converted either into DAG and then common membrane lipids through the Kennedy pathway, or into other lipids via the CDP-DAG route. Mitochondria appear to play a key role, using an enzyme called TAMM41 to help convert incoming PA into phosphatidylglycerol, a building block for mitochondrial membranes. When Nir2 was depleted, newly made PA no longer stayed where it was produced but accumulated instead in the Golgi region, showing that disrupting one courier can reroute lipid traffic through the cell and alter important signaling pathways such as mTOR, a major growth regulator.
Why This Work Matters for Cell Health
By combining precise “membrane editing” with proximity tagging, this study provides a powerful blueprint for discovering how cells detect and correct imbalances in their lipid makeup. For PA, the work maps out a network of enzymes and transport proteins that rush to overloaded membranes, move PA to safer locations, and transform it into other lipids, all while keeping key signaling circuits in check. Because similar lipid disturbances are linked to cancer, metabolic disease, and neurodegeneration, the feeding–fishing approach could be used to uncover new drug targets across many conditions where membrane lipids silently shape cell behavior.
Citation: Tei, R., Li, XL., Luan, L. et al. Membrane editing with proximity labeling reveals regulators of lipid homeostasis. Nat Chem Biol 22, 728–739 (2026). https://doi.org/10.1038/s41589-025-02104-x
Keywords: phosphatidic acid, lipid homeostasis, membrane proteomics, lipid transfer proteins, optogenetic membrane editing