Clear Sky Science · en
Structural basis of GAIN domain autoproteolysis and cleavage-resistance in the adhesion G-protein coupled receptors
How Cell Surface Receptors Act Like Tiny Self-Cutting Machines
Our cells are covered with elaborate receptor proteins that sense the outside world. Some of these, called adhesion G protein–coupled receptors, behave like tiny self-cutting machines: a piece of the protein can slice itself, changing how the receptor signals. This study asks a deceptively simple question with big implications for brain function and drug design: why do some of these receptors readily cut themselves, while closely related ones stubbornly resist?
A Hidden Switch on Sticky Cell Receptors
Adhesion G protein–coupled receptors (aGPCRs) help cells stick to their surroundings and talk to neighboring cells. A hallmark of these receptors is a bulky outer region that includes the so‑called GAIN domain. Buried inside this domain lies a short “tethered agonist” segment that can switch the receptor on when exposed. In many aGPCRs, a tiny section of the protein chain in the GAIN domain can cut itself at a specific site, splitting the receptor’s outer portion into an N‑terminal and a C‑terminal fragment. The cutting occurs next to a particular building block sequence that includes a histidine and a serine or threonine and is thought to act as a built‑in chemical knife.
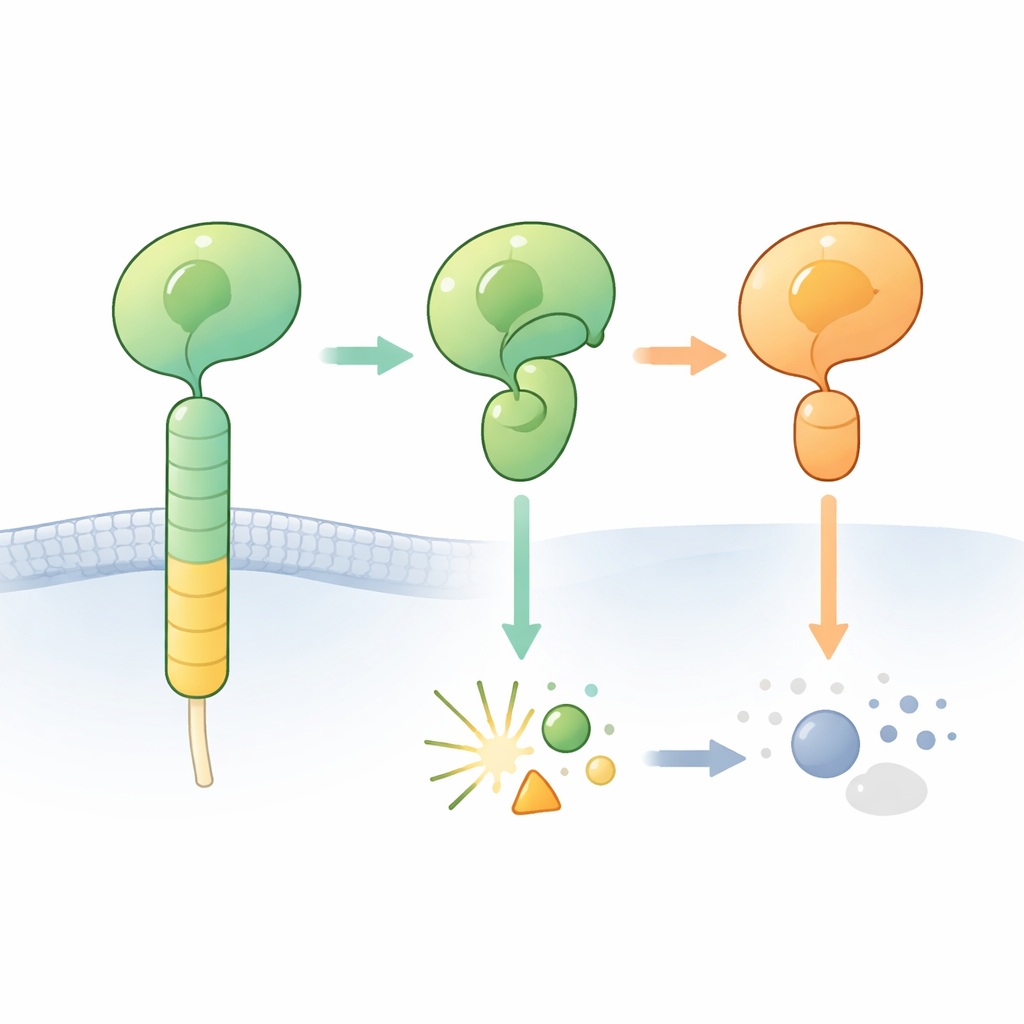
When the Expected Cut Never Happens
The researchers zoomed in on one brain receptor, ADGRB2 (also known as BAI2), whose GAIN domain carries the “right” histidine–leucine–serine sequence but is intriguingly hard to cut. They determined a high‑resolution crystal structure of the hormone receptor–like and GAIN regions from human ADGRB2. Surprisingly, the cleavage site sits in an open groove that water molecules can reach, but the key histidine and serine side chains are oriented in a way that is not ready for chemistry: they would need to twist into different positions to start the reaction. Heating the purified protein or adding chemicals that normally push stalled self‑cutting reactions to completion did not trigger cleavage, confirming that ADGRB2 is naturally cleavage‑resistant under these conditions.
Subtle Ring Interactions Steer the Chemical Strike
To understand what makes other receptors more “cut‑happy,” the team compared ADGRB2 to family members that do cleave efficiently and ran extensive molecular dynamics simulations, letting the atoms jiggle and shift on the computer. A key insight emerged: in cleavage‑competent receptors, the histidine at the cutting site nestles against a nearby phenylalanine or tyrosine side chain in a T‑shaped ring‑to‑ring interaction. This gentle nudge locks the histidine into a preferred orientation that lines it up to pull a proton off the serine or threonine, turning that group into a powerful attacker of the peptide bond. In ADGRB2, this phenylalanine is replaced by a serine, and the backbone region nearby is distorted by a bulkier side chain, so the histidine spends far less time in the “attack‑ready” pose. Simulations also showed that in efficient cutters, the peptide bond to be broken is often slightly twisted, storing strain that makes it easier to snap.
Flexible Loops as Adjustable Levers
Beyond this ring interaction, the GAIN domain contains floppy loop regions, or “flaps,” that frame the cutting groove. These loops vary widely between receptors and are not strongly conserved in evolution, yet they turn out to be crucial for full activity. By swapping flap segments between a normally cleavage‑competent receptor (ADGRL1) and ADGRB2, the authors could dial cleavage rates up or down. Introducing phenylalanine‑like residues in the right spot restored some self‑cutting ability to a normally inactive cousin, ADGRB3, especially when combined with a histidine at the catalytic position. Conversely, mutating the stabilizing phenylalanine in the active receptor or grafting in ADGRB2‑like flaps weakened cleavage. These experiments show that both precise local chemistry and long, mobile loops cooperate to push the protein into a strained, reactive state.
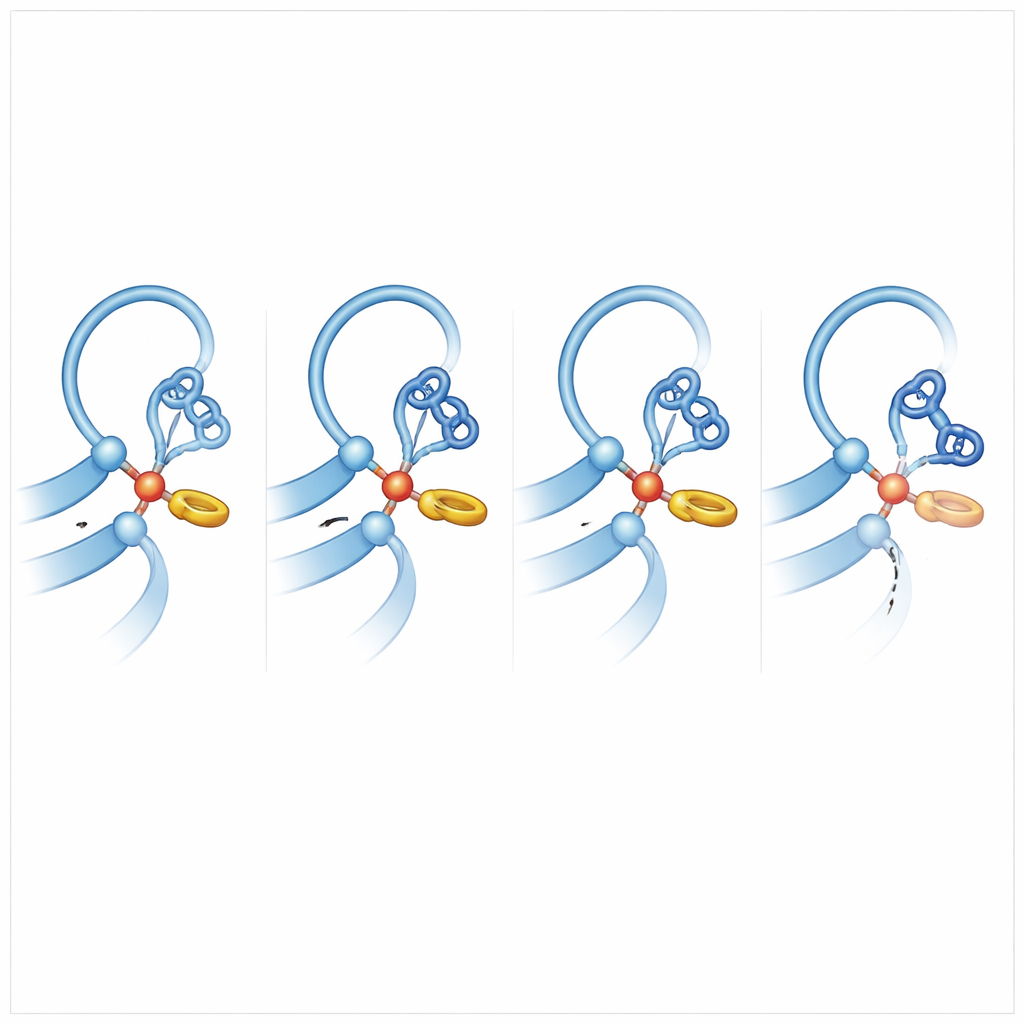
Why Some Receptors Stay Intact
Looking across nearly 150 mammalian versions of ADGRB2, the team found that most lack the ideal histidine at the cutting site, and their flaps and nearby residues are highly variable. Together with the measured half‑life of about 100 days for cleavage in purified ADGRB2, this suggests that many BAI2‑like receptors are designed to avoid rapid self‑cutting. Instead, they may signal either without ever cleaving or by using very slow or context‑dependent cleavage in specific tissues. More broadly, the work shows that self‑processing in these receptors is not a simple on–off switch controlled by a three‑letter motif. Rather, it emerges from a delicate balance of side‑chain packing, backbone strain and mobile loops that collectively tune whether the GAIN domain acts as a built‑in knife or remains safely locked. Understanding this balance may eventually allow researchers to design drugs that shift receptors between stable and self‑cutting states, changing how cells sense mechanical forces and chemical cues.
Citation: Pohl, F., Seufert, F., Chung, Y.K. et al. Structural basis of GAIN domain autoproteolysis and cleavage-resistance in the adhesion G-protein coupled receptors. Nat Commun 17, 3259 (2026). https://doi.org/10.1038/s41467-026-71225-1
Keywords: adhesion GPCR, GAIN domain, autoproteolysis, protein structure, molecular dynamics