Clear Sky Science · en
AMR-GNN: a multi-representation graph neural network framework to enable genomic antimicrobial resistance prediction
Why predicting drug resistance matters
Antibiotic-resistant infections are one of the major medical threats of our time, killing more than a million people each year. Doctors urgently need faster ways to tell which antibiotics will work for a given infection, but traditional lab tests can take several days. This study introduces a new artificial intelligence framework, called AMR-GNN, that reads the full DNA sequence of bacteria and predicts whether they will be resistant or susceptible to different antibiotics, potentially paving the way for same-day guidance at the bedside.
From slow cultures to digital DNA tests
Today, most hospitals still rely on culture-based tests: bacteria are grown in the lab and exposed to various drugs to see which ones stop their growth. While reliable, this approach is slow and labor-intensive. At the same time, sequencing entire bacterial genomes has become cheaper and easier, generating huge amounts of detailed information. The challenge is that bacterial DNA is extremely high-dimensional, containing millions of building blocks, and there is no single agreed-upon way to turn this genetic code into a format that computers can easily use to predict drug resistance. Earlier tools often focused on a few known resistance genes or simple patterns, which works well when resistance is due to a single mutation but fails when many subtle changes interact.
Combining different genomic views into one picture
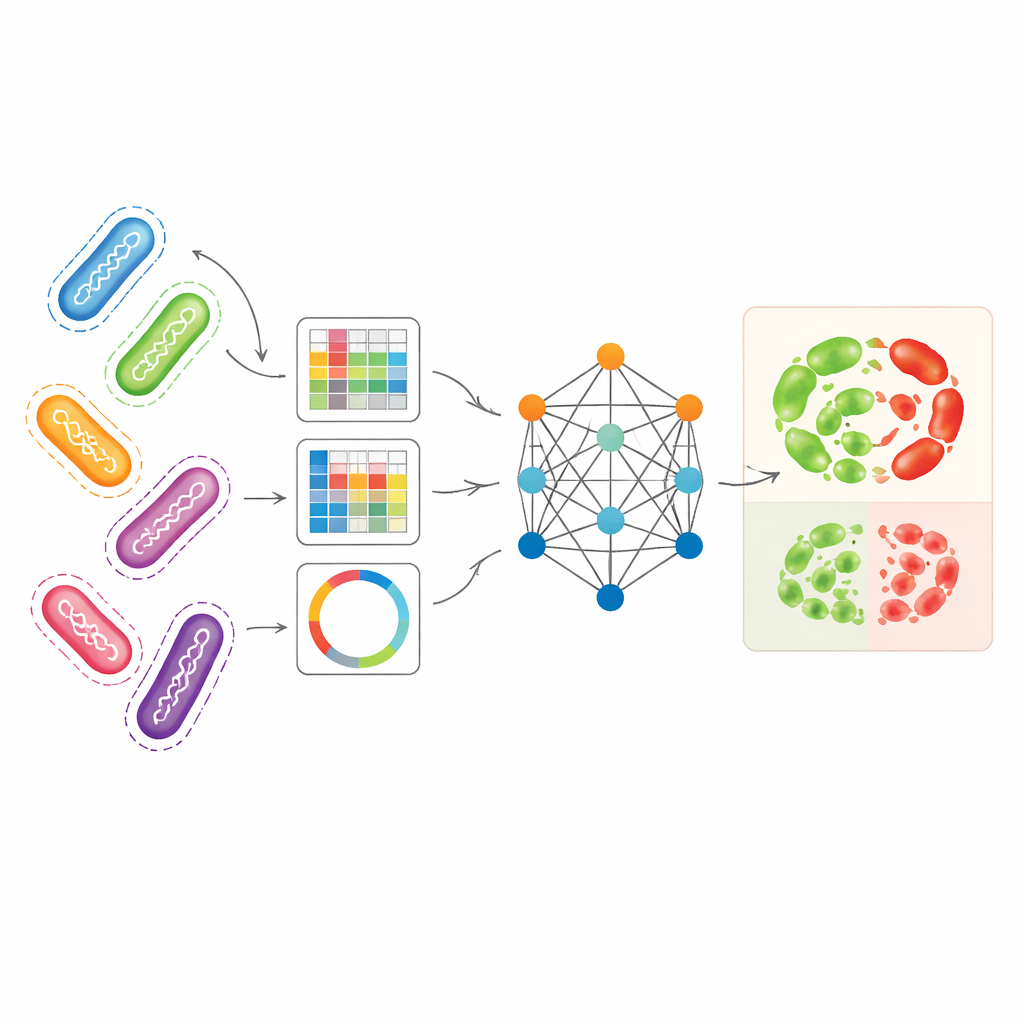
The researchers set out to harness several complementary ways of representing bacterial genomes at once. They used Pseudomonas aeruginosa—a hospital-acquired pathogen with particularly complex resistance patterns—as their main test case. One representation, called unitigs, captures recurring DNA fragments without relying on a reference genome. Another representation tracks small DNA changes at specific positions, while a third converts selected resistance-related genes into image-like patterns that summarize how short stretches of DNA are arranged. On their own, these representations already allowed standard machine-learning models to predict resistance with reasonable accuracy, especially unitigs for certain antibiotics. But each view misses part of the biological story, and using them separately underutilizes the richness of the underlying genome data.
How the graph-based model learns from related strains
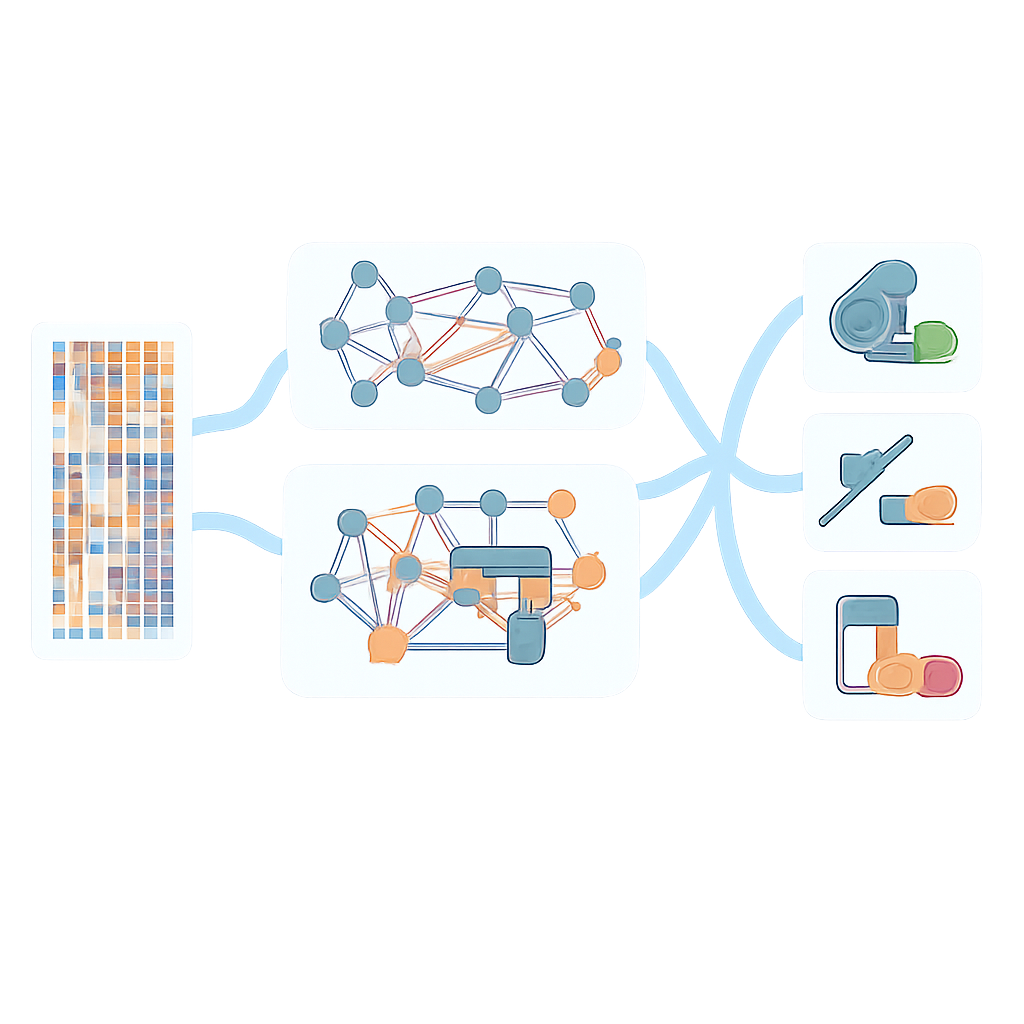
AMR-GNN uses a form of deep learning known as a graph neural network, which treats each bacterial isolate as a point (a node) and connects isolates that are genetically similar with links (edges). In this setup, the detailed unitig profile of each isolate forms its main feature vector, while the other genomic views define how isolates are connected in the graph. The model then passes information along these connections, allowing it to learn from patterns shared among related genomes. To avoid being misled by simple clonal relationships—where bacteria are closely related but differ in resistance for reasons the model should uncover—the authors deliberately removed edges that linked isolates from the same genetic lineage group. This "decoupling" step forces the network to pay more attention to the specific DNA features associated with resistance, rather than relying on broad lineage labels as shortcuts.
Stronger predictions across bacteria and drugs
When the team compared AMR-GNN to simpler models that relied on single genomic views, the graph-based approach improved performance for nearly all of the 12 antibiotics tested in P. aeruginosa, with the biggest gains for drugs that were previously hardest to predict. The model also generalized better to independent test datasets, though performance still dropped somewhat outside the training data, highlighting the need for larger and more diverse collections of genomes. Beyond P. aeruginosa, the researchers applied AMR-GNN to more than 23,000 genomes from other major pathogens, including Escherichia coli, Klebsiella pneumoniae, Staphylococcus aureus, and Enterococcus faecium, across many clinically important antibiotics. In most species–drug combinations, the framework reached very high accuracy and outperformed widely used rule-based tools that depend on curated lists of known resistance genes.
Making black-box models more explainable
An important concern for clinical use is whether such an AI system can provide insight into why it makes a particular prediction. The team tackled this by applying interpretability methods that trace which DNA features contribute most to the model’s decisions. For drugs where the model performed best, AMR-GNN highlighted many known resistance genes and mutations, such as classic targets of fluoroquinolone antibiotics. It also pointed to less well-understood genes whose changes were strongly associated with higher drug concentrations needed to stop bacterial growth, suggesting new candidates for laboratory follow-up. This ability to both predict resistance and flag potential biological drivers helps bridge the gap between pure prediction and mechanistic understanding.
What this means for future patient care
In essence, this work shows that combining multiple DNA "views" inside a graph-based deep-learning model can substantially improve the prediction of antibiotic resistance from bacterial genomes. AMR-GNN is presented as a flexible, interpretable framework that can be extended to other data types, such as gene activity measurements or clinical information. While more work is needed—especially larger, geographically varied datasets and prospective clinical trials—the approach brings us closer to a future where a bacterial genome sequence, obtained directly from a patient sample, could quickly guide doctors toward the right drug and help slow the spread of resistant infections.
Citation: Nguyen, HA., Peleg, A.Y., Wisniewski, J.A. et al. AMR-GNN: a multi-representation graph neural network framework to enable genomic antimicrobial resistance prediction. Nat Commun 17, 3555 (2026). https://doi.org/10.1038/s41467-026-69934-8
Keywords: antimicrobial resistance, graph neural networks, bacterial genomics, machine learning, antibiotic susceptibility prediction