Clear Sky Science · en
A nutrient bottleneck controls antibiotic efficacy in structured bacterial populations
Why this matters for infections
Antibiotics are often tested in well-mixed test tubes, yet many real infections involve dense bacterial communities clinging to tissues or medical devices. In these crowded settings, food and drugs seep in from the outside, creating sharp chemical gradients that lab tests rarely capture. This study asks a simple but powerful question: can the way nutrients flow through these structured bacterial populations control how well antibiotics work—and even help pockets of bacteria survive and bounce back?
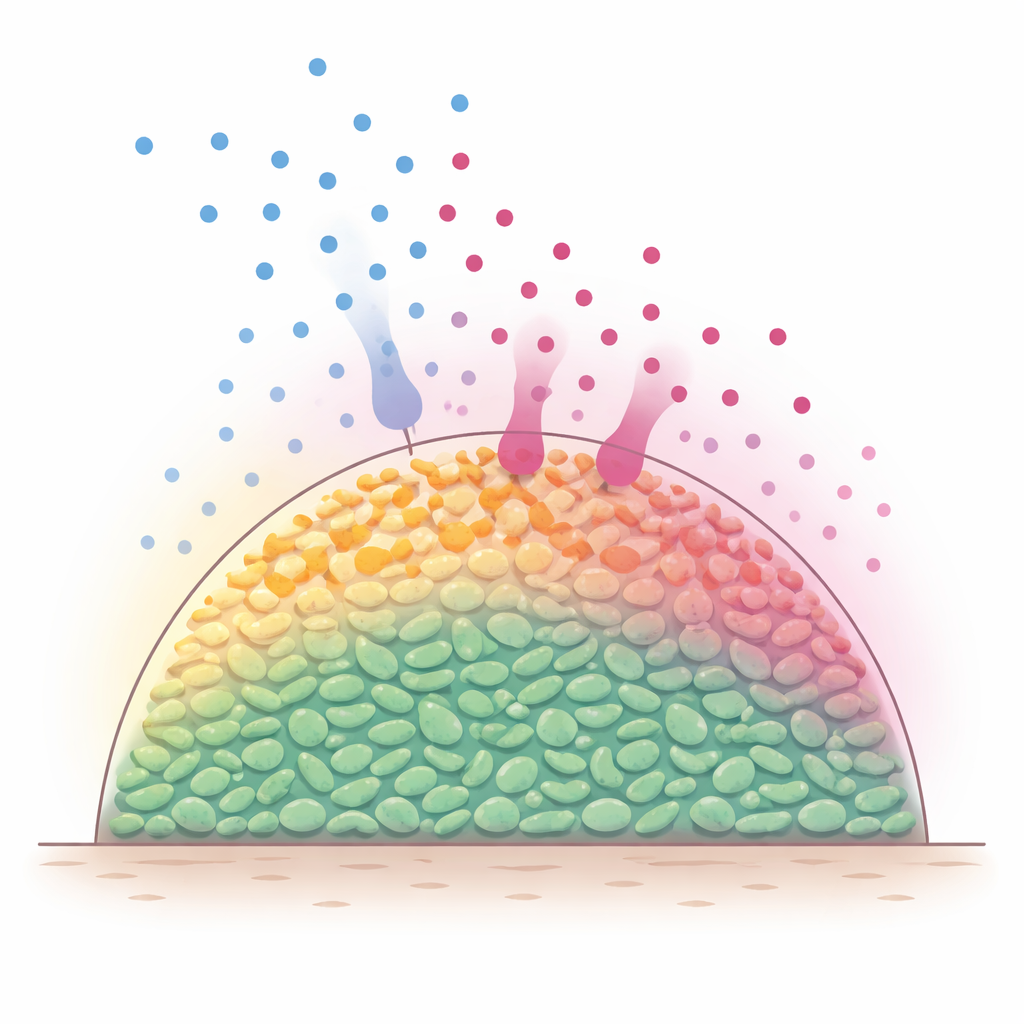
Hidden layers inside bacterial communities
In natural settings, bacteria frequently live in multicellular assemblies, sometimes called biofilms, where cells pack together in three dimensions. Nutrient molecules, such as sugars, diffuse in from the outside and are quickly eaten by the outermost cells. As a result, the surface layers stay metabolically active, while deeper cells become starved and sluggish. Because many antibiotics are most effective against actively growing bacteria, scientists have long suspected that these built-in nutrient gradients create a shielded interior that is harder to kill. However, it has been difficult to directly watch what happens inside such dense populations as nutrients and antibiotics move and react over time.
Building a controllable mini-biofilm
To tackle this challenge, the researchers trapped E. coli bacteria inside transparent hydrogel matrices—soft, water-filled materials whose pores hold cells in place but still allow small molecules to diffuse. One side of each gel block contained bacteria but no nutrients or drugs; the other side held a reservoir of sugar (glucose), antibiotic (fosfomycin), or both. By removing a divider between the two halves, the team could let molecules slowly diffuse while using fluorescence microscopy to distinguish living cells from dead ones. This setup mimics a cross-section of a biofilm, with the outside world on one side and the protected interior on the other, but with much tighter experimental control than in natural samples.
When food unlocks lethal action
Surprisingly, when the antibiotic alone diffused into the bacterial region—even at doses far above standard inhibitory levels—most cells remained alive for a full day. The picture changed dramatically when a modest amount of glucose was added along with the antibiotic. In that case, the team observed a remarkably sharp “death front” sweeping inward from the exposed side: an advancing layer where living cells abruptly gave way to dead ones. The speed of this front depended strongly on nutrient supply and consumption but was largely insensitive to how much extra antibiotic was added once it exceeded a certain threshold. More sugar sped up the front; more cells, which collectively ate the sugar faster, slowed it down. Similar behavior appeared with several other nutrients and antibiotics, suggesting a general principle rather than a quirk of a single drug.
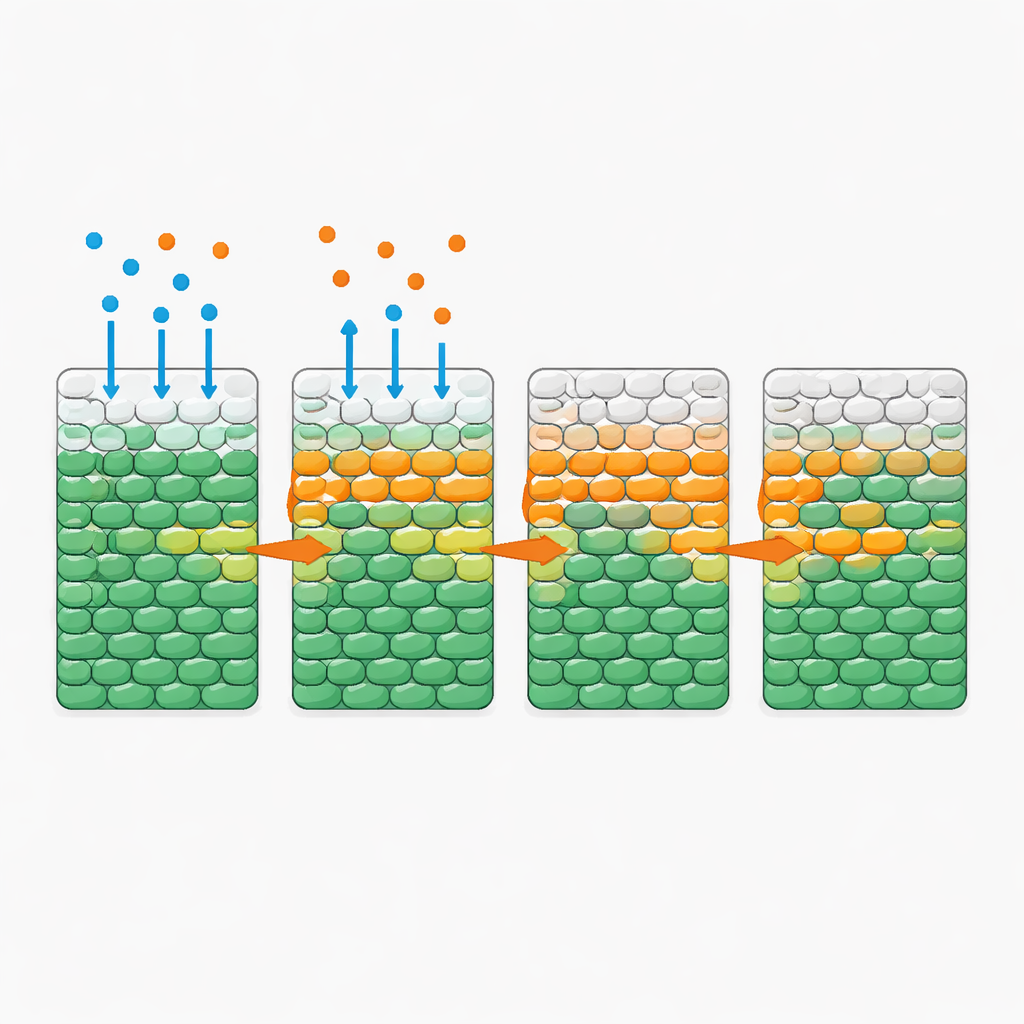
Simple rules behind a complex pattern
To understand these dynamics quantitatively, the authors built a mathematical model that tracks three ingredients: bacteria, nutrient, and antibiotic. In the model, nutrient and antibiotic diffuse through the population; cells consume nutrient to grow; and once exposed to enough antibiotic and nutrient for long enough, they first multiply and then die. All of the parameters were measured independently in well-mixed cultures, so the model required no tuning to the spatial experiments. Yet its simulations reproduced the observed death fronts and their dependence on nutrient level, cell density, and consumption rate. Analysis of the model revealed a key dimensionless quantity that compares the time it takes cells to grow and die under antibiotic exposure with the time it takes them to deplete their local nutrient supply. When nutrient is exhausted quickly, inner cells remain starved and thus largely protected; when consumption is slow, nutrient penetrates deeply and the front can sweep through the entire population.
How survival and regrowth emerge
Bacterial populations are rarely uniform in their sensitivity to drugs, even when the cells share the same genetic background. The team used their platform to explore what happens when some cells are more resistant than others. At intermediate antibiotic doses, the death front still formed and moved inward, but after it passed, scattered microcolonies of more resistant cells began to regrow in its wake, feeding on nutrients released into now-emptier regions. Lowering the antibiotic dose or increasing the starting cell density amplified this regrowth. In structured populations, many such resistant pockets can persist side by side, in contrast to well-mixed cultures where typically a single winner dominates. This pattern could give infections more chances to evolve higher resistance or rebound after treatment.
What this means for treating infections
This work shows that nutrient availability is not just a background detail but a central bottleneck that can either enable or limit antibiotic killing in structured bacterial populations. Rather than simply asking whether a drug can reach an infection site, clinicians and researchers may also need to ask how food sources flow through the community and how fast cells consume them. The study’s framework, including its simple metric for when nutrient consumption outpaces killing, offers a way to predict when antibiotic treatment will likely stall at a protective shell. It also suggests that combining antibiotics with strategies that alter nutrient penetration or disrupt spatial structure could make existing drugs more effective, and highlights why dense, biofilm-like infections can be so stubborn even when lab tests say a drug should work.
Citation: Hancock, A.M., Dill-Macky, A.S., Moore, J.A. et al. A nutrient bottleneck controls antibiotic efficacy in structured bacterial populations. Nat Commun 17, 3337 (2026). https://doi.org/10.1038/s41467-026-69625-4
Keywords: antibiotic resistance, bacterial biofilms, nutrient gradients, fosfomycin, microbial population dynamics