Clear Sky Science · en
Structural insight into hierarchical DNMT3A autoinhibition and its dysregulation in disease
How cells mark their DNA
Every cell in our body carries the same DNA, yet heart cells behave very differently from brain cells. One way cells manage this feat is by placing tiny chemical tags on DNA, helping to turn genes on or off. This article explores how a key tagging enzyme, DNMT3A, is kept under tight control—and how small changes in its structure can misdirect these tags and contribute to human disease.
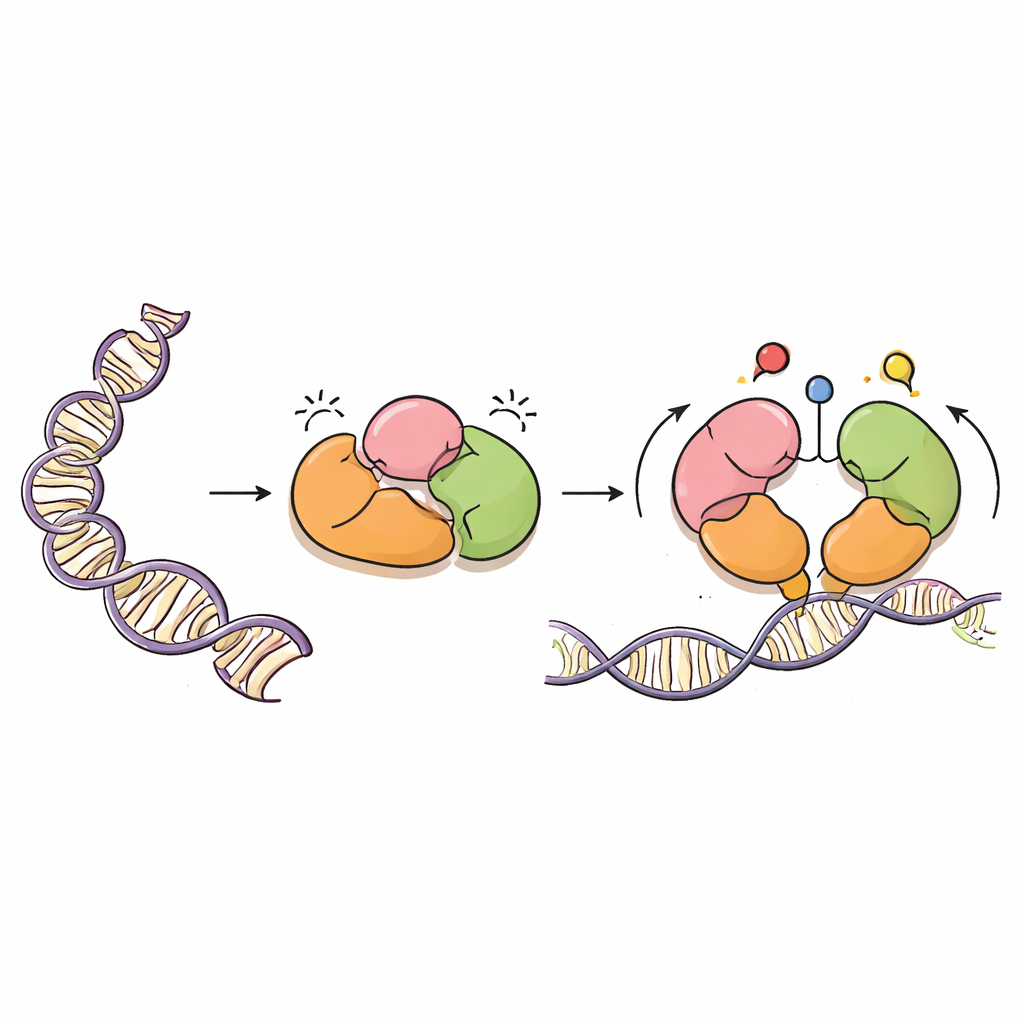
A careful painter of DNA
DNMT3A is an enzyme that adds methyl groups—simple carbon-based marks—to specific DNA letters, usually at sites called CpG dinucleotides. These marks are vital for normal development, imprinting of parental genes, and the long-term stability of our genomes. But like a powerful paint sprayer, DNMT3A must be precisely guided. It works alongside a partner protein, DNMT3L, and carries several "reader" regions that sense nearby proteins called histones, which package DNA into chromatin. These reader regions help DNMT3A know where it should work and where it should stay quiet.
Built-in brakes inside the enzyme
Using cryo–electron microscopy, the authors solved the three-dimensional structure of a major DNMT3A form (DNMT3A2) bound to DNMT3L. They discovered that DNMT3A contains layered self-braking, or autoinhibition. Two reader regions, called PWWP and ADD, fold back onto the enzyme’s catalytic core in a way that physically blocks the part of the protein that recognizes CpG sites, as well as pockets that normally sense chemical marks on histones. This compact arrangement prevents DNMT3A from accessing DNA, keeping its methylating activity low until the right signals appear.
Histone signals release the brakes
The team overlaid their structure with previously known structures of DNMT3A and related proteins bound to histone fragments. This comparison showed that when histone tails carry certain marks—specifically unmodified lysine-4 (H3K4me0) and dimethylated lysine-36 (H3K36me2)—they compete with DNMT3A’s internal contacts. Binding of these histone marks pries the PWWP and ADD regions away from the catalytic core and from a flexible loop that recognizes CpG sites. Molecular dynamics simulations, which model protein motion over time, revealed that once these internal clamps are loosened, the CpG-recognition loop becomes more mobile and exposed, better positioned to engage DNA. In essence, DNMT3A switches from a closed, resting shape to a more open, active shape only when it senses a suitable chromatin environment.
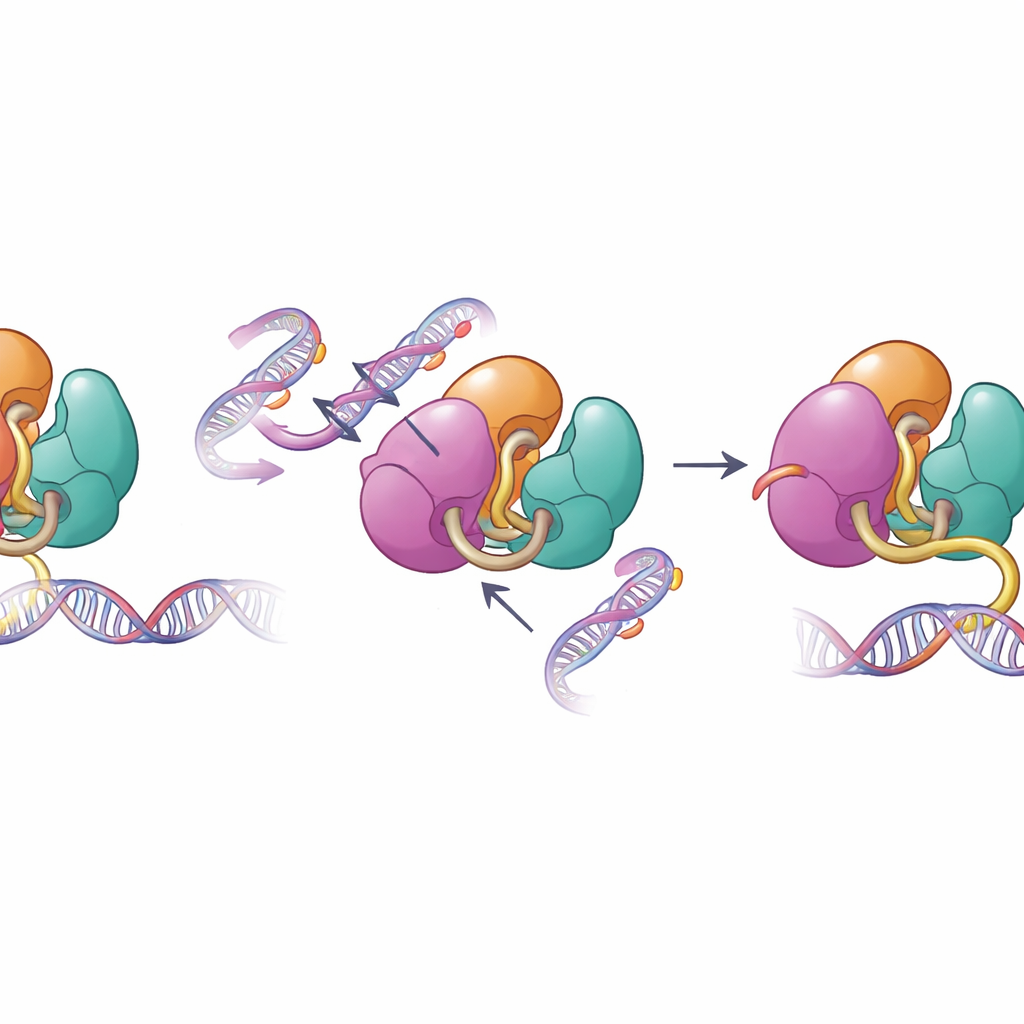
When structural safeguards fail
The same surfaces that mediate autoinhibition are frequently altered in human disease. The authors examined disease-linked changes in DNMT3A found in blood cancers and developmental disorders, including mutations in the PWWP and ADD regions. Biochemical tests showed that these mutations weaken the internal contacts that keep DNMT3A in check, boosting its ability to bind DNA and methylate it in test-tube experiments. When the team looked at methylation patterns in mouse stem cells carrying one of these mutations, they found that DNMT3A became less responsive to the normal histone signal H3K36me2 and started methylating stretches of the genome that would usually remain less marked. This loss of targeting precision helps explain why such mutations are associated with widespread, misplaced DNA methylation in patients.
Why this matters for health
Overall, the study reveals DNMT3A as a finely tuned machine whose activity is governed by nested layers of self-restraint, released only in the right chromatin setting. By mapping exactly how these internal brakes work—and how disease-causing mutations disable them—the work provides a structural blueprint for understanding abnormal DNA methylation in cancer and developmental syndromes. In the long run, insights into these control points may guide efforts to design drugs that gently restore proper DNMT3A regulation, correcting misapplied DNA marks without shutting down this essential enzyme entirely.
Citation: Lu, J., Vig, E., Chen, J. et al. Structural insight into hierarchical DNMT3A autoinhibition and its dysregulation in disease. Nat Commun 17, 2901 (2026). https://doi.org/10.1038/s41467-026-69563-1
Keywords: DNA methylation, epigenetics, DNMT3A, chromatin regulation, cancer mutations