Clear Sky Science · en
Structure and mechanism of the HECT ligase HECTD3
How Cells Tag Their Proteins for Action
Inside every cell, a tiny signaling system decides which proteins are repaired, recycled, or rewired for new tasks. This study focuses on one of the enzymes that runs this system, a protein called HECTD3. HECTD3 has been linked to cancer, inflammation, and viral infection, but how it works at the atomic level has remained unclear. By combining powerful imaging and biochemical tools, the authors reveal how HECTD3 is built, how it chooses its molecular partners, and how it decorates other proteins with small tags that can change their fate.
A Cellular Tagging Machine with Many Roles
Cells often modify their proteins with a small molecule called ubiquitin, which can act like a colored tag in a sorting warehouse. Strings of these tags, arranged in different ways, send different messages: some mark a protein for destruction, others tune its activity or location. Enzymes called E3 ligases are responsible for picking which proteins are tagged and how. HECTD3 belongs to a branch of E3 ligases known to form a temporary chemical link with ubiquitin before passing it on to a target. It is unusually active in several cancers and has been tied to processes such as programmed cell death, immune signaling, and viral replication, yet its structure and working logic were largely unknown.
A Constantly On Enzyme with a Unique Shape
The researchers first asked whether HECTD3 behaves like a switch that can be turned on or off. Many related ligases keep themselves in check by folding into self‑blocking shapes or by pairing up into larger assemblies. Using biochemical tests and a technique that measures the size of proteins in solution, the team found that HECTD3 mostly exists as a single copy and does not appear to shut itself down through self‑inhibition or stable clustering. Both its core catalytic section and the full‑length protein are active, with a clear preference for working with one family of helper enzymes that hand over ubiquitin. This means HECTD3 is essentially “ready to go” and likely controlled in cells by factors other than simple self‑blocking.
Revealing a New Protein Architecture
To understand how HECTD3 is built, the authors used cryo‑electron microscopy to capture images of single molecules frozen in thin ice. They determined structures of the protein alone and in a state where a ubiquitin molecule is firmly attached to its active site. These snapshots show that HECTD3 is compact rather than ring‑shaped, with a classic two‑lobed catalytic region at one end and a previously mysterious N‑terminal region at the other. The N‑terminal half turns out to form two entirely new folds, which the authors name D3NA and D3NB, arranged as a helical platform supporting a substrate‑binding DOC module. When ubiquitin is loaded, the catalytic lobes adopt a particular L‑shaped arrangement, and specific acidic residues near the active site help prepare incoming protein side chains to accept the tag. The work shows that, although HECTD3 follows the general rules of its enzyme family, it uses a distinct combination of structural elements to do so.
How HECTD3 Tags Itself and Its Targets
Next the team examined what kinds of ubiquitin patterns HECTD3 creates. Rather than building long chains of one specific type, the enzyme mainly attaches multiple single ubiquitin units and short chains, with evidence for some chains that favor a linkage known to promote protein breakdown. This suggests HECTD3 is better at starting tagging events than at extending them into long signals. The authors then turned to a proposed target protein called PARP1, a DNA repair factor already known to influence cancer therapy responses. In test‑tube reactions, HECTD3 efficiently modified PARP1, decorating several lysine sites in its DNA‑binding half with tags that include chains associated with protein removal. By using chemical cross‑linking and mass spectrometry, they mapped contact points between HECTD3 and PARP1, confirming that the DOC module and the catalytic region both help position the substrate. Mutating two exposed amino acids on the DOC surface sharply reduced PARP1 tagging without harming the core chemistry, highlighting them as key anchors for substrate recognition.
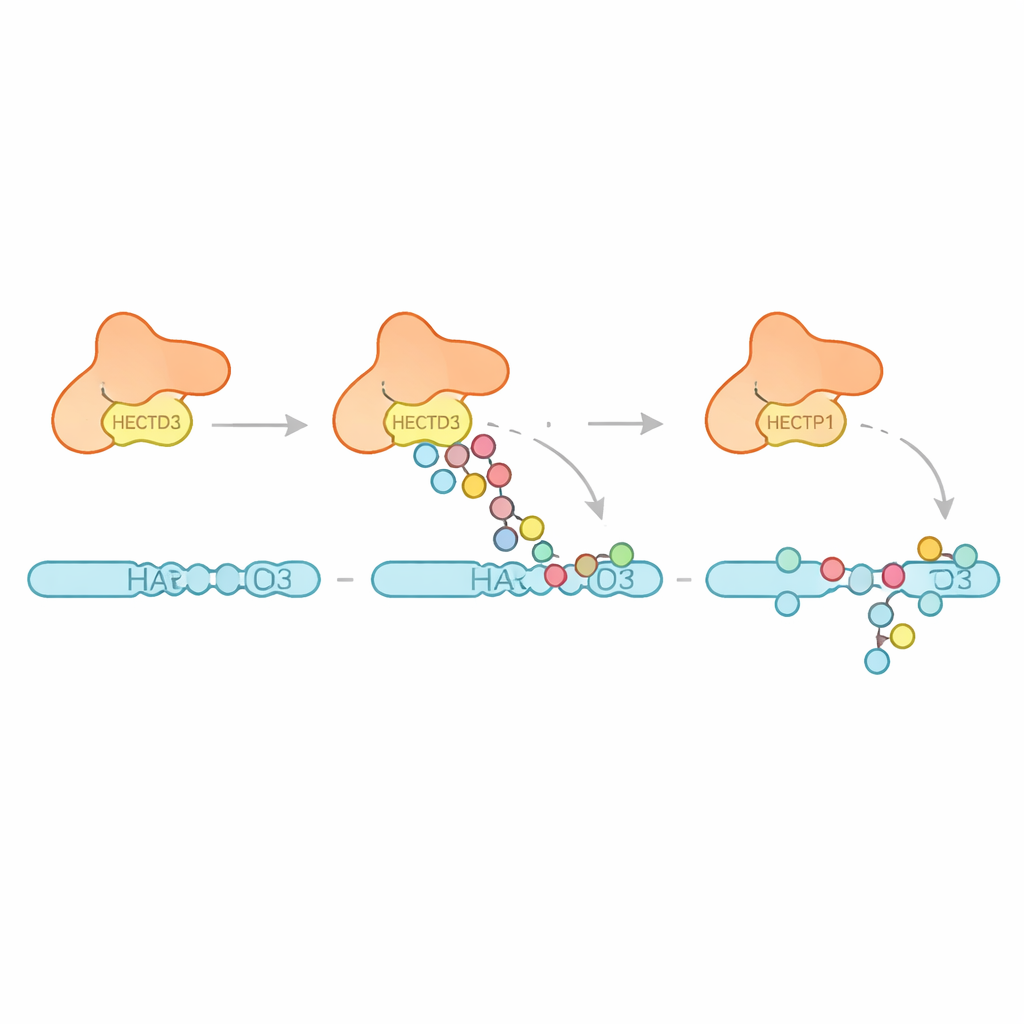
Why This Matters for Disease and Treatment
Together, these findings paint HECTD3 as a constantly active but structurally sophisticated tagging machine that mainly kicks off ubiquitin marks and relies on a flexible, unique N‑terminal region to recognise specific partners like PARP1. Because HECTD3 activity is elevated in several cancers and shapes how DNA repair proteins are handled, understanding its structure and mechanism opens doors to designing molecules that interfere with its docking surfaces or catalytic center. Such targeted interventions could, in principle, shift how cells decide the fate of critical proteins in cancer, infection, and inflammatory disease.
Citation: Huber, J., Esposito, D., Maslen, S. et al. Structure and mechanism of the HECT ligase HECTD3. Nat Commun 17, 2783 (2026). https://doi.org/10.1038/s41467-026-69520-y
Keywords: ubiquitin ligase, HECTD3, protein tagging, PARP1, cancer biology