Clear Sky Science · en
High-throughput multi-organ proteomics workflow for drug efficacy and toxicity analysis
Why Faster Protein Maps Matter for Medicine
Our bodies are made of thousands of different proteins that shift in response to disease and drugs. Reading those changes across many organs at once is key to understanding why a treatment works, when it fails, and what side effects it may cause. This study introduces a rapid way to measure proteins in hundreds of tissue samples per day and applies it to a cancer drug called L-asparaginase, revealing how the drug reshapes biology throughout the body.
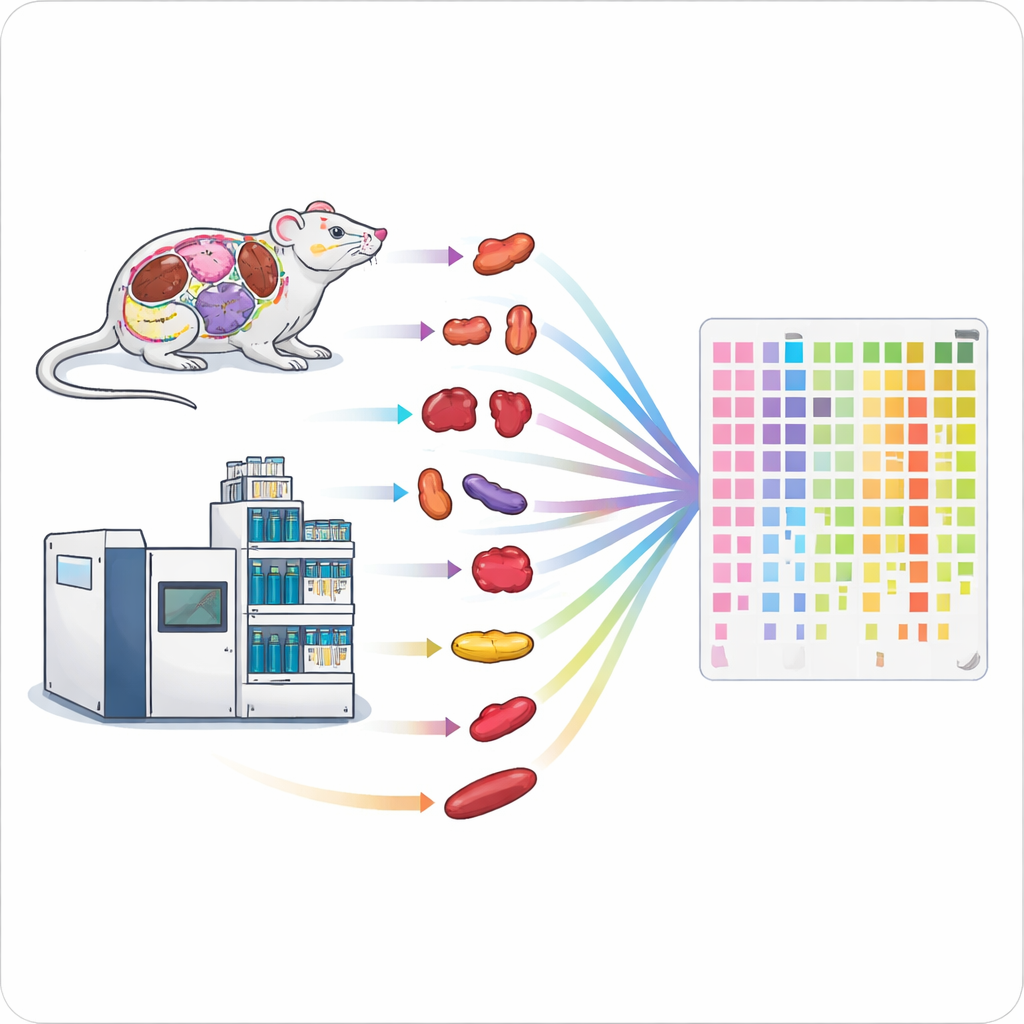
Speeding Up the Protein Assembly Line
Traditional methods for measuring proteins are slow and labor‑intensive, making it hard to study whole organisms or large patient groups. The authors combined two high‑performance technologies—a fast type of mass spectrometry and short liquid chromatography runs—into a single streamlined workflow. By carefully tuning settings such as how long ions are collected and which mass ranges are scanned, they achieved deep protein coverage in just one to two minutes of measurement per sample. Their optimized setup could identify roughly 6,200 human proteins in a one‑minute run and about 7,500 in two minutes, far surpassing earlier high‑throughput methods.
From Tissue to Data in a Single Day
Collecting data quickly is only half the battle; preparing the samples can be an even bigger bottleneck. To address this, the team designed a parallel sample‑handling pipeline for solid tissues. They used automated bead‑based homogenizers to break open up to 24 tissue pieces at once, followed by ultrasound‑assisted digestion of proteins into peptides in 96‑well plates and rapid desalting using specialized tips. This high‑throughput pipeline allowed 96 tissues to be processed in about five hours, compared with several days using a standard workflow. Tests in mouse brain, kidney, and lung showed that the fast method recovered similar numbers of proteins and produced highly comparable quantitative results, but with more than seven‑fold higher throughput.
Watching a Cancer Drug Act Across the Body
To demonstrate what this platform can reveal, the researchers turned to L‑asparaginase, an enzyme drug used for childhood leukemia that is difficult to repurpose for adults because of side effects and drug resistance. They treated mice with standard L‑asparaginase, a modified version with reduced activity on a related amino acid, or a control solution, and then collected 13 different organs over an eight‑day course. In total, they quantified more than 11,000 proteins across 507 samples. Patterns in the protein data cleanly separated tissues such as brain, liver, and fat depots and showed that the standard drug, but not the mutant version, triggered large shifts in protein levels across many organs. Measurements of small molecules in blood and tissues confirmed that the standard drug strongly depleted the amino acid asparagine, while the mutant caused only brief changes, helping explain why it is less effective.
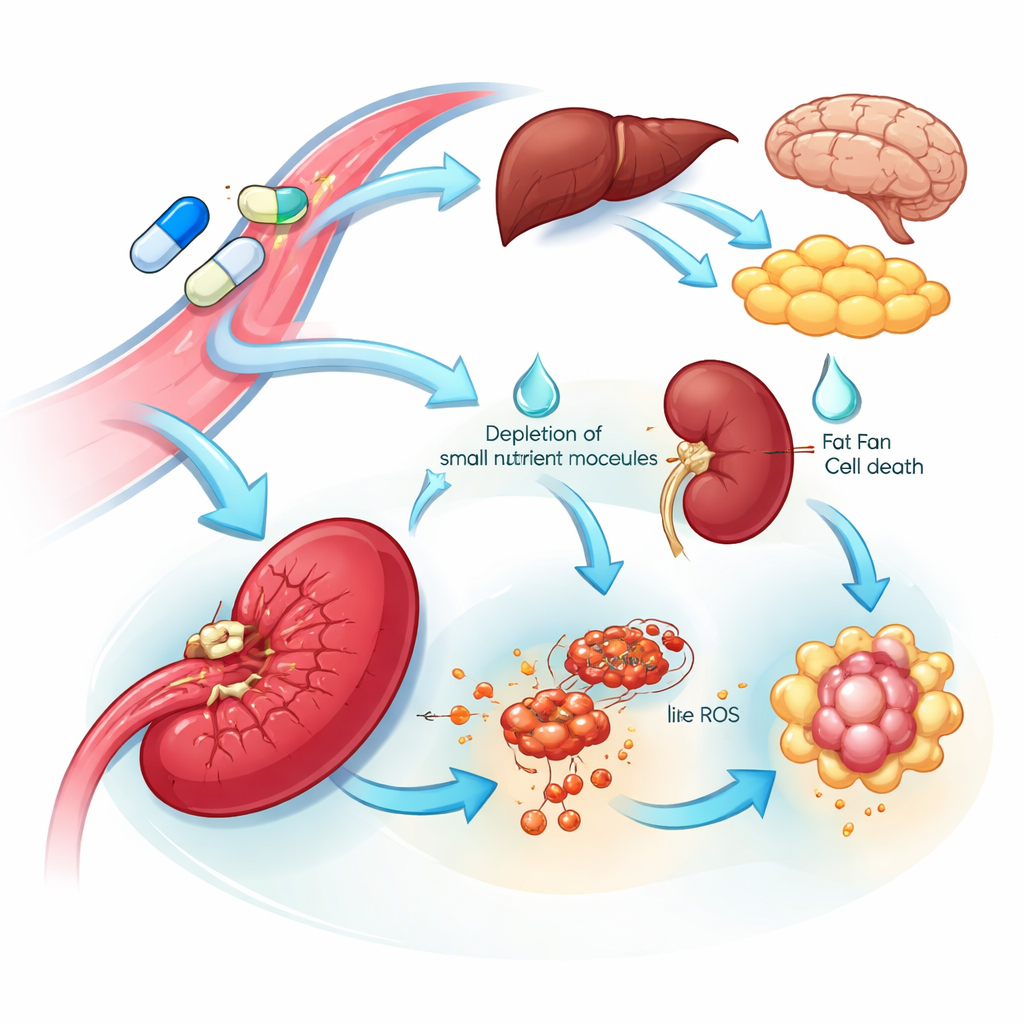
Clues to Organ Vulnerability and Side Effects
The multi‑organ view uncovered both vulnerable targets and potential sources of toxicity. Many tissues responded to L‑asparaginase by increasing the enzyme that makes asparagine, suggesting a built‑in resistance program that helps restore this nutrient. In contrast, the spleen showed a striking collapse in this enzyme and a drop in related amino acids and DNA building blocks, along with signals consistent with iron‑driven cell death. These changes suggest that spleen‑related cells may be particularly sensitive to the drug. Across nearly all organs, the standard drug—but not the mutant—suppressed proteins involved in blood clot regulation and cholesterol handling. This broad, coordinated downshift mirrors known clinical problems with the drug, including blood‑clotting disorders and high blood triglycerides, and points to specific blood proteins that might be monitored or targeted to reduce these risks.
What This Means for Future Treatments
By pairing a rapid, scalable protein‑measurement pipeline with a whole‑body drug study, the authors created a kind of molecular atlas for L‑asparaginase: where it acts, how organs adapt or fail to adapt, and which pathways link its benefits to its harms. For non‑specialists, the key message is that it is now possible to scan thousands of proteins across many organs in just a day, turning complex drug effects into detailed, testable maps. Such workflows could accelerate the design of safer cancer therapies, guide dose and schedule decisions, and help identify combination treatments that preserve anticancer power while blunting dangerous side effects.
Citation: Xiong, Y., Tan, L., Chan, Wk. et al. High-throughput multi-organ proteomics workflow for drug efficacy and toxicity analysis. Nat Commun 17, 3505 (2026). https://doi.org/10.1038/s41467-026-69471-4
Keywords: proteomics, drug toxicity, cancer therapy, mass spectrometry, systems biology