Clear Sky Science · en
The GENEVA platform models tumor mosaicism to reveal variations of responses to KRAS inhibitors and identify improved drug combinations
Why this matters for future cancer treatments
Targeted cancer drugs are designed to strike specific genetic weaknesses in tumors, yet patients with seemingly similar cancers can respond very differently. This article introduces GENEVA, a new experimental platform that recreates that real-world diversity in the lab and in mice, then tracks how thousands of individual cancer cells respond to treatment over time. By doing so, the researchers uncover why some cells die, why others endure, and how to choose smarter drug combinations that could keep cancers from bouncing back.
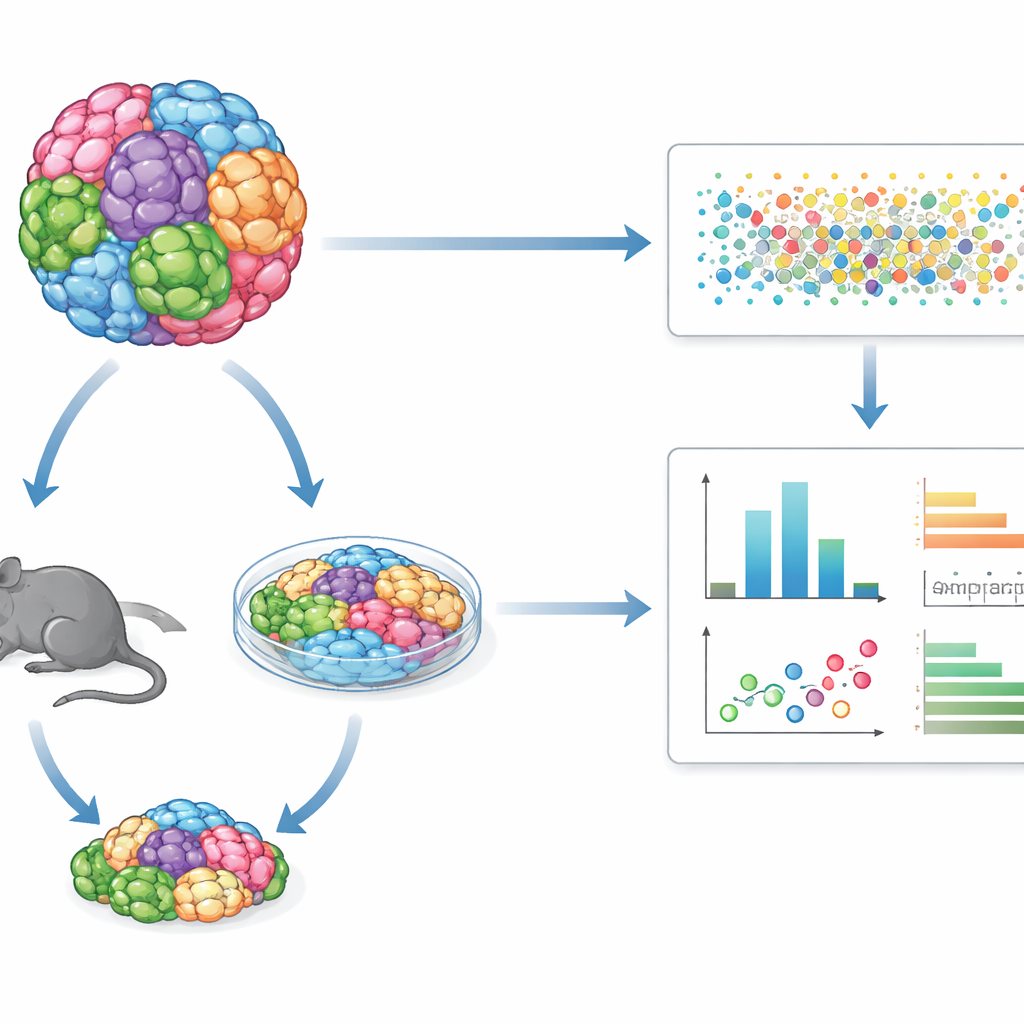
Many tumor types in a single shared test
Instead of studying one cancer cell line at a time, GENEVA mixes dozens of genetically distinct human cancer cell lines or patient-derived tumor samples into a single “mosaic” culture or tumor. These mixed populations are grown either in three-dimensional lab cultures or as tumors in mice. After treatment with a drug, the team uses single-cell RNA sequencing and genetic barcodes to read out, cell by cell, which genetic background each cell came from and what molecular state it is in. This lets them measure not only how many cells survive, but how their internal programs change, revealing patterns of sensitivity and resistance across many tumor types in one experiment.
Probing a new class of KRAS-blocking drugs
To showcase the power of GENEVA, the researchers focused on a recently approved group of drugs that block a cancer-driving mutation called KRAS G12C. These drugs have brought real benefits for some people with lung and other cancers, but responses are often short-lived. Using GENEVA, the team pooled KRAS-mutant and non-mutant cell lines and treated them with multiple KRAS G12C inhibitors at different doses, both in dishes and in mice. As expected, KRAS G12C cells were generally more sensitive, but even within that group some cells or lines persisted, hinting at hidden survival strategies that standard assays would miss.
Power surge in the cell’s power plants
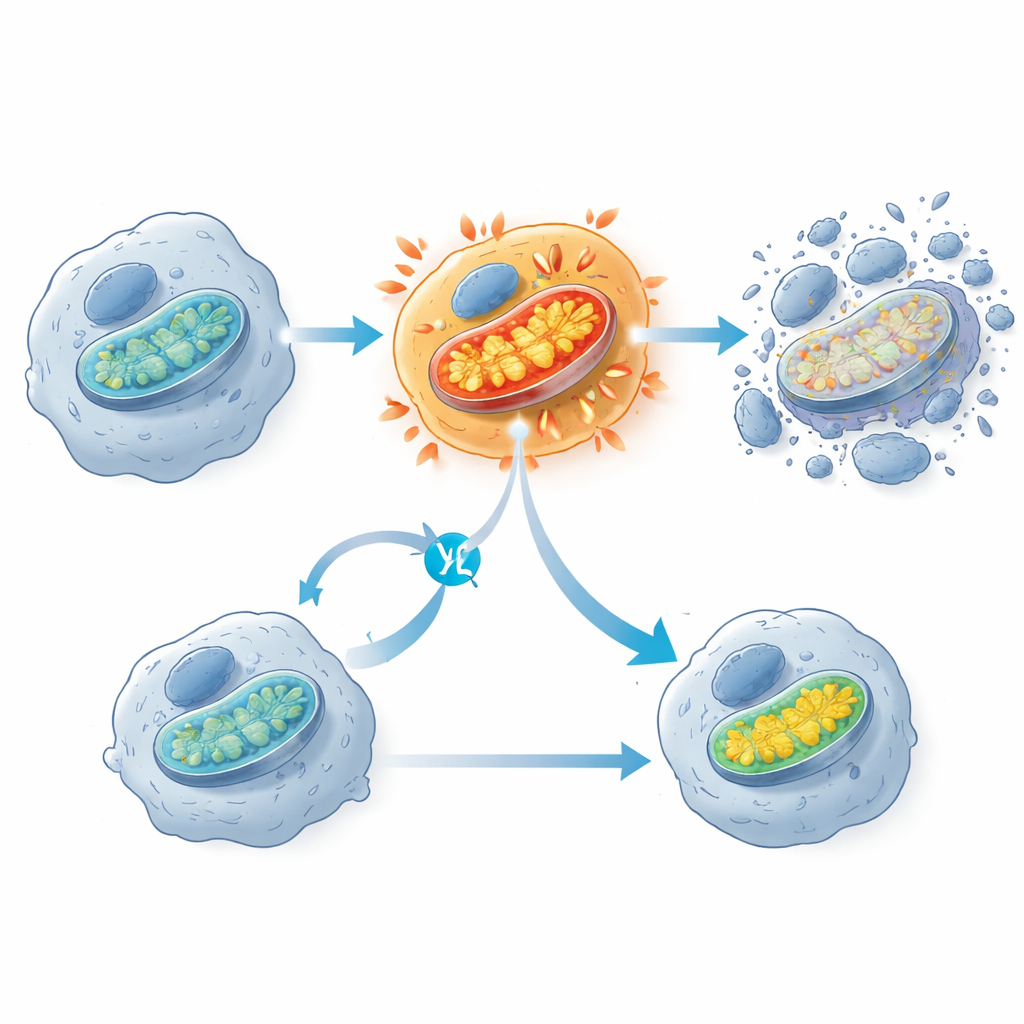
A striking finding was that KRAS G12C inhibitors trigger an early burst of activity in mitochondria, the structures often described as the cell’s power plants. Shortly after drug exposure, sensitive cancer cells showed a spike in oxygen consumption and mitochondrial membrane potential, followed hours later by activation of cell-death enzymes. Cells that weathered treatment tended to have fewer mitochondrial components and dampened mitochondrial gene activity. When the researchers partially blocked the mitochondrial electron transport chain with another drug, it actually protected KRAS G12C-mutant cells from dying, suggesting that this sudden mitochondrial overdrive is a key step that pushes cells over the edge. The team also observed signs of oxidative damage to fats in cell membranes, but blocking this process did not prevent cell death, indicating it is a side effect rather than the main kill switch.
Uncovering hidden escape routes and better pairings
Because GENEVA profiles thousands of surviving cells, it can highlight genes and pathways that are turned up specifically in drug-tolerant KRAS G12C cells. The study flagged several such routes, including signaling through mTOR, enzymes that alter DNA packaging, and programs linked to changes in cell shape and movement known as epithelial–mesenchymal transition (EMT). Many of these features appeared most strongly in tumors growing in mice, where the surrounding tissue microenvironment is present. Guided by these signals, the team tested combination treatments. Pairing a KRAS G12C inhibitor with an mTOR inhibitor or with a blocker of TGF-β (a driver of EMT) produced stronger and more lasting tumor shrinkage in mice than any single drug alone.
From single cells to smarter precision therapies
In plain terms, this work shows that how a cancer cell manages its energy and shape can dictate whether a targeted KRAS drug will cure it or merely slow it down. The GENEVA platform makes it possible to watch these decisions unfold across many tumor types and conditions at once, revealing common weak spots and escape paths. By identifying that a mitochondrial “power surge” helps kill KRAS G12C-mutant cells, and that pathways like mTOR and EMT let some cells slip away, the study points to concrete drug combinations that may prolong responses in patients. More broadly, GENEVA offers a way to test future cancer drugs in realistic, genetically diverse models long before clinical trials, increasing the odds that precision therapies will work for more people.
Citation: Yu, J.X., Suh, J.M., Popova, K.D. et al. The GENEVA platform models tumor mosaicism to reveal variations of responses to KRAS inhibitors and identify improved drug combinations. Nat Cancer 7, 522–537 (2026). https://doi.org/10.1038/s43018-026-01130-5
Keywords: KRAS G12C inhibitors, tumor heterogeneity, mitochondrial metabolism, drug resistance, combination cancer therapy