Clear Sky Science · en
Comprehensive analysis of aberrant alternative splicing and RNA binding proteins regulators associated with myocardial ischemia reperfusion injury in mice
Why heart restart can still hurt the heart
When someone has a heart attack, doctors rush to reopen the blocked artery and restore blood flow. Yet this life saving step can also injure the heart in a delayed wave of damage called reperfusion injury. This study in mice looks deep inside heart cells at how tiny RNA messages are cut and edited when blood flow returns, and how those changes might worsen or ease the damage. Understanding this hidden layer of control could open new paths to protect the heart after a heart attack.
Hidden messages inside heart cells
Every cell uses RNA as a working copy of genetic information. Before these RNA messages are used to make proteins, they can be cut and rejoined in different ways, a process known as alternative splicing. That means a single gene can give rise to several protein versions with distinct effects. The authors asked how this editing process changes in mouse hearts exposed to a controlled blockage and reopening of a key coronary artery, a standard model of heart attack followed by treatment. They reanalyzed an existing RNA sequencing dataset that compared heart tissue from injured mice with sham operated controls.
Wide shifts in RNA editing after injury
The team found that reperfusion injury was accompanied by sweeping shifts in RNA editing patterns across the genome. They detected hundreds of splicing events whose usage differed between healthy and injured hearts, especially at the starting end of RNA segments. Many of the affected genes belonged to important cell signaling routes that decide whether heart muscle cells grow, recycle their components, or self destruct. In particular, genes in the mTOR and MAPK pathways, which help control cell metabolism, stress responses, and inflammation, showed strong changes in how their RNA was cut and assembled. 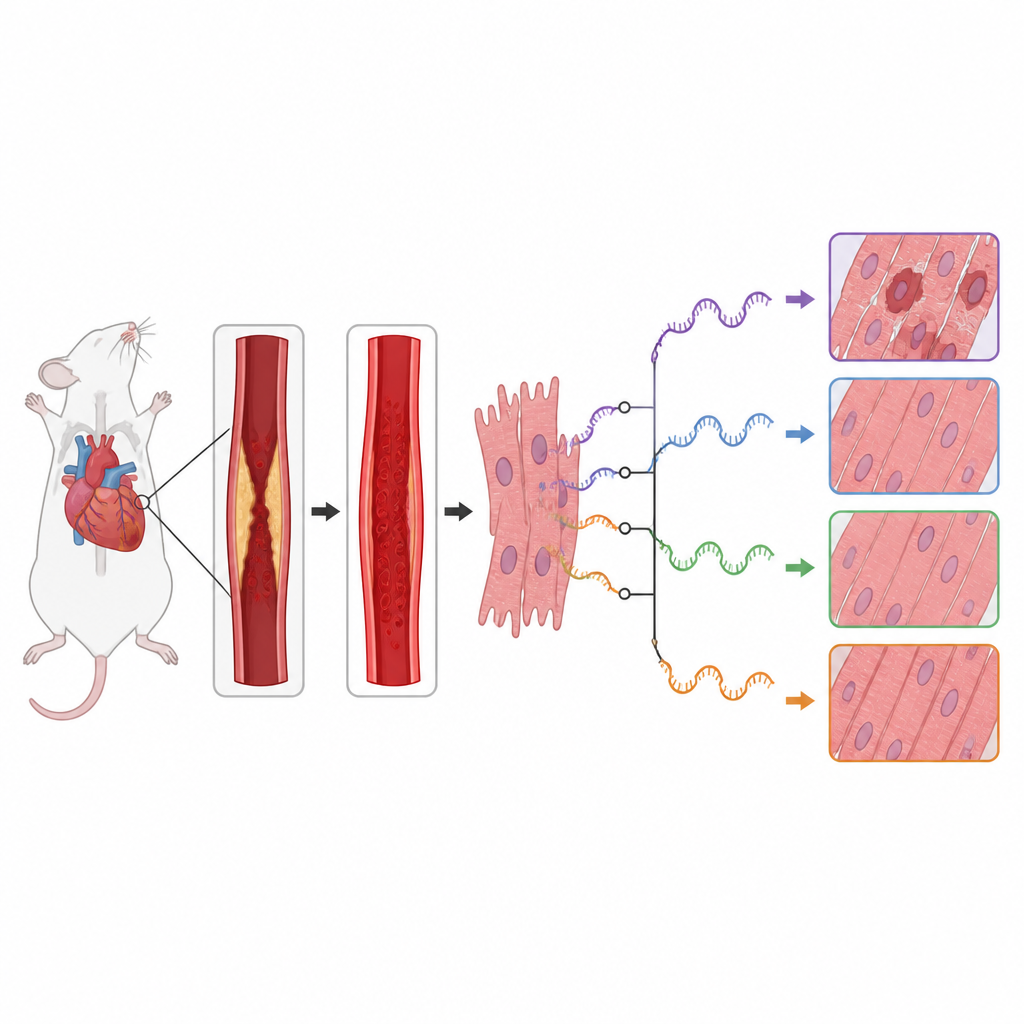
Proteins that steer RNA cutting
RNA binding proteins act as traffic controllers that decide where each RNA is sliced and stitched. By comparing gene activity in injured and control hearts, the researchers identified more than a hundred such proteins whose levels changed after injury. Many were linked to blood vessel growth and responses to low oxygen, two processes central to heart recovery. One protein, LMNA, stood out because its levels closely tracked the editing of a specific RNA from a gene called Atp6v1h, which helps regulate a cellular pump tied to mTOR signaling. The spliced form of this RNA was more common in injured hearts, hinting that LMNA may help fine tune energy sensing during the stress of reperfusion.
Zooming in on key genes and confirming in animals
The authors highlighted several genes whose edited RNA forms may tilt heart cells toward survival or damage. For example, a hypoxia responsive factor called Eif4e2, an inflammation related regulator Traf6, the insulin receptor Insr, and a stress sensitive regulator Nr4a1 all produced RNA variants that were more frequent after injury. Others, such as Map4k4, showed reduced use of particular splice forms. To test whether these RNA shifts were real beyond computer analysis, the team created their own mouse model of reperfusion injury. They confirmed that many of the predicted gene variants and RNA binding proteins changed in the same direction at both the RNA and protein levels, strengthening the case that these are genuine features of the injured heart. 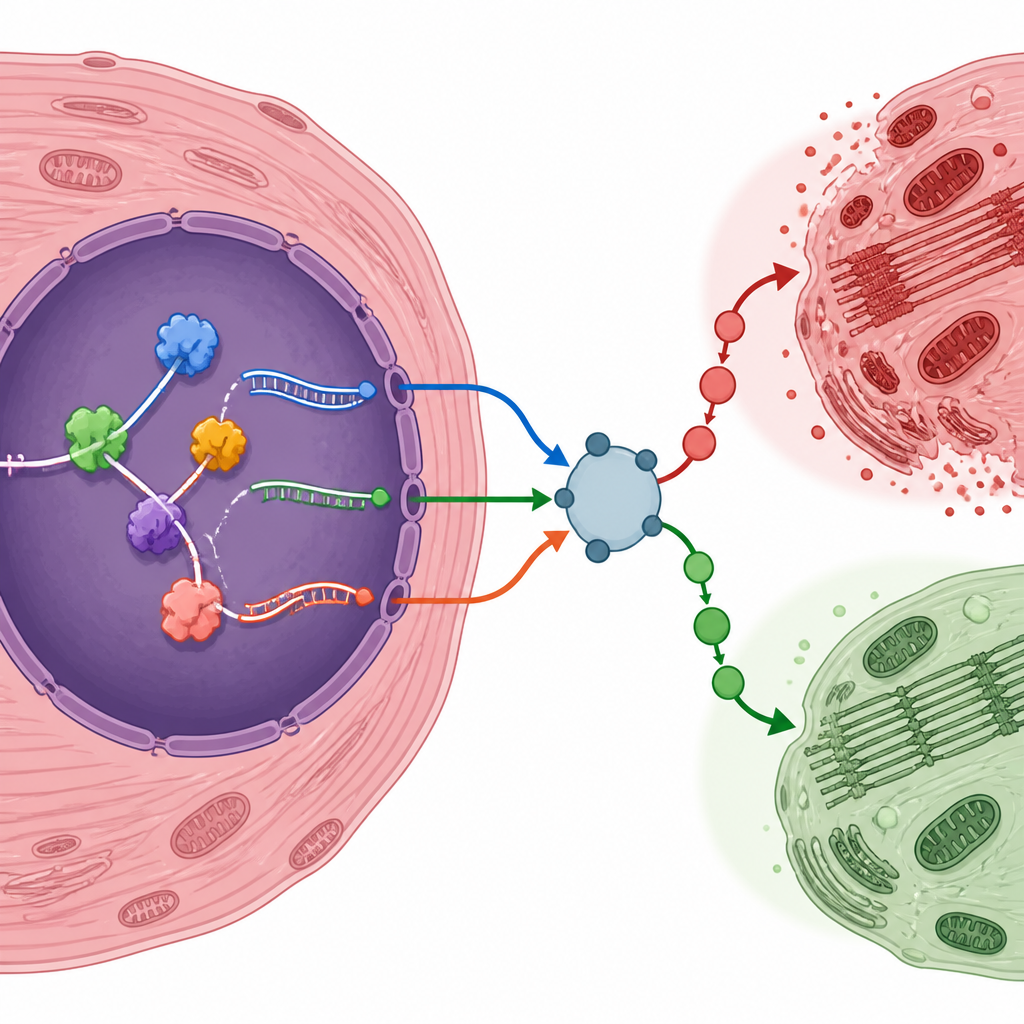
What this means for future heart care
In plain terms, this work shows that after blood flow returns to the heart, cells rapidly rewire their internal messages by recutting RNA in new ways, guided by a set of regulatory proteins. These edited messages tweak major decision making pathways that govern cell stress, cleanup, and inflammation. While the study was done in mice and does not yet prove cause and effect, it maps out a network of candidate RNA switches and their controllers that may shape how much permanent damage follows a heart attack. In time, therapies that gently adjust RNA binding proteins or specific splice patterns could complement current treatments, offering a more precise way to shield the heart when its blood supply is restored.
Citation: Zhou, D., Tan, Y., Jiang, B. et al. Comprehensive analysis of aberrant alternative splicing and RNA binding proteins regulators associated with myocardial ischemia reperfusion injury in mice. Sci Rep 16, 15156 (2026). https://doi.org/10.1038/s41598-026-45174-0
Keywords: myocardial ischemia reperfusion injury, alternative splicing, RNA binding proteins, mTOR pathway, MAPK signaling