Clear Sky Science · en
Molecular recognition and induced dimerization of hnRNP A2/B1 truncations by G-quadruplex single strand DNA
How Cells Spot Viral DNA
Viruses leave traces of their genetic material inside our cells, and the body must recognize these traces quickly to mount a defense. This study explores how one abundant cellular protein, hnRNP A2/B1, may help sense unusual DNA shapes that are common in many viruses. By understanding how this protein changes its form when it binds to such DNA, researchers hope to lay the groundwork for new antiviral and anticancer strategies.
A Shape-Sensing Protein in the Nucleus
HnRNP A2/B1 is best known as a helper protein that manages RNA, the working copy of genetic information. It lives mainly in the cell nucleus and participates in a host of essential tasks, from splicing RNA messages to controlling their stability. Recent work, however, has revealed a second life for this protein: it can act as a sensor for foreign DNA from invading viruses. When viral DNA enters the nucleus, hnRNP A2/B1 participates in turning on innate immune signals that trigger production of antiviral molecules called interferons.
Unraveling an Aggregated Protein
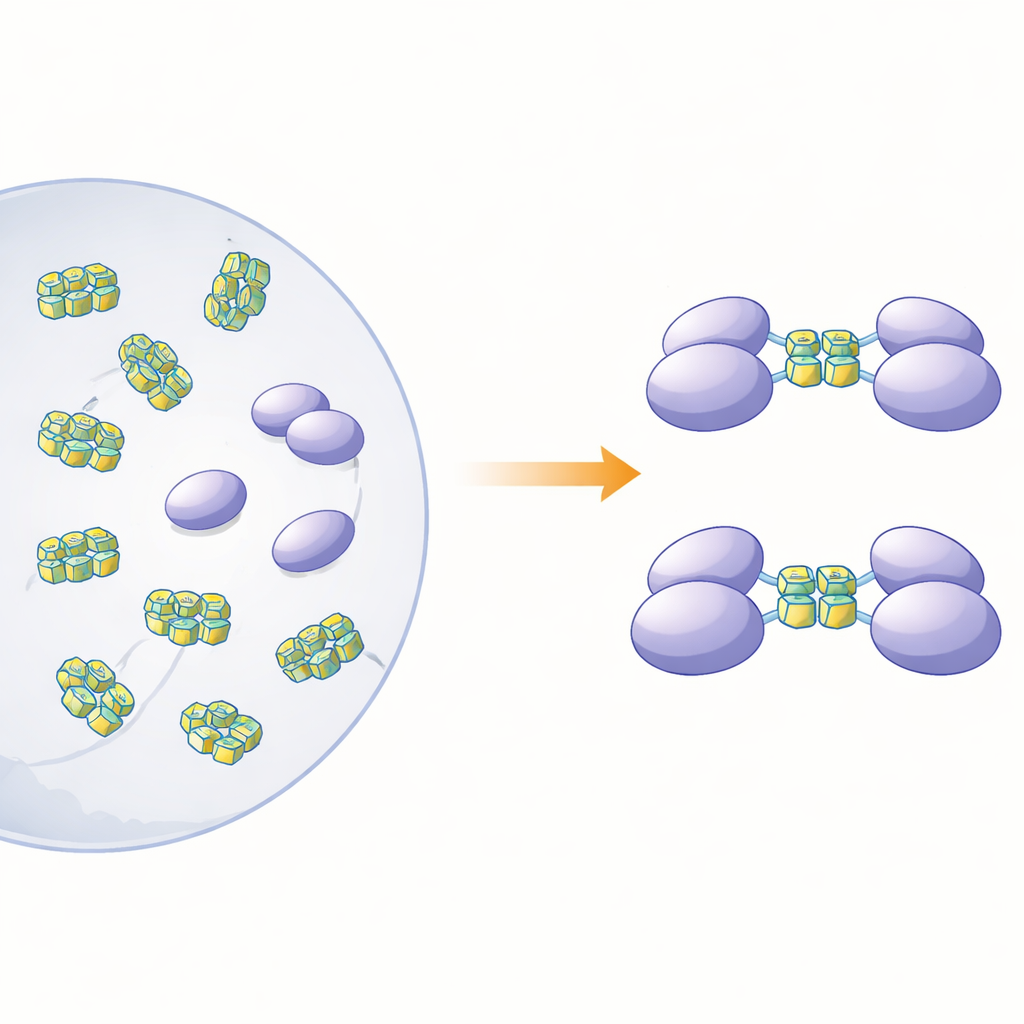
To clarify how this protein’s physical form relates to its sensing role, the authors produced full-length hnRNP A2/B1 and several shortened versions in bacteria. When they examined the full-length protein in solution, they found that it did not behave as neat, separate particles. Instead, it formed large, irregular clumps—"soluble amorphous aggregates"—that were unstable and prone to further precipitation. In contrast, three truncated versions lacking different stretches at the protein’s tail end remained as tidy, single molecules (monomers) in solution. Computer-based structure predictions suggested that roughly three-quarters of hnRNP A2/B1 is made of floppy, disordered segments, especially near its ends. These flexible regions appear to promote aggregation and help explain why it has been so difficult to grow crystals of the protein for high-resolution structural studies.
Viral-Like DNA That Makes Proteins Pair Up
The researchers then asked which kinds of DNA the monomeric fragments would bind, and what that binding would do to the protein’s state. They focused on single-stranded DNA segments that can fold into G‑quadruplexes—compact stacks formed by guanine-rich sequences that bend the strand into a tight, U-shaped structure. Such shapes are sprinkled throughout the genomes of many viruses and cancer-related genes. Using calorimetry and gel-based binding tests, the team showed that one fragment, spanning the so-called RRM and RGG regions (residues 15–250), binds strongly to single-stranded DNA but not to the corresponding double-stranded form. Size-based separation and ultracentrifugation experiments revealed a striking effect: when this fragment encountered G‑quadruplex DNA of 12 or 22 bases, two copies of the protein fragment joined together to form a dimer, but non‑quadruplex DNA did not trigger this pairing.
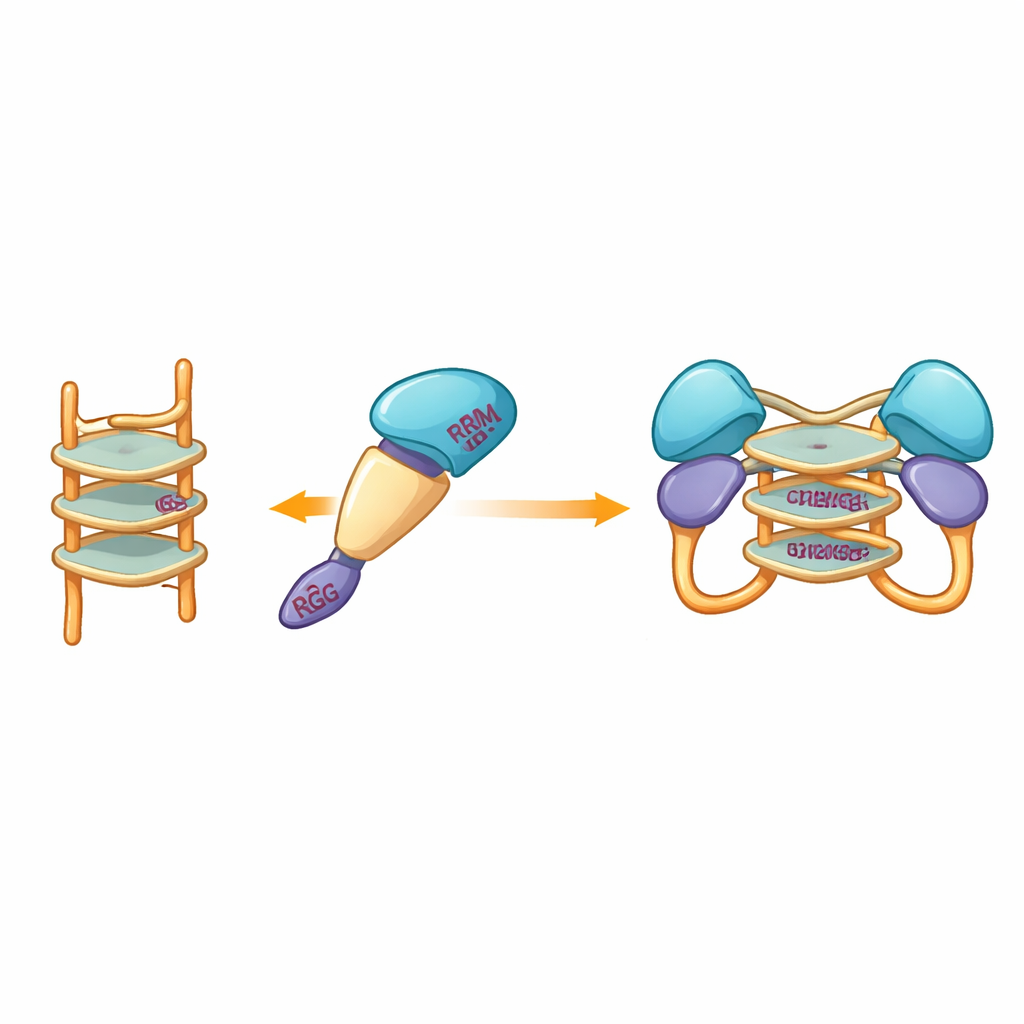
Peering into a Flexible Interface
Because experimental crystallography attempts repeatedly failed, the team turned to computational modeling to visualize how the protein fragment might clasp a G‑quadruplex. Models built with modern prediction tools suggested that the more rigid central domains of hnRNP A2/B1 cradle the stacked DNA structure, while surrounding disordered segments remain mobile. Docking simulations highlighted a set of specific amino acids from several domains that could form hydrogen bonds with the G‑quadruplex, stabilizing a dimeric complex in which two protein molecules share the same DNA segment. Notably, these interacting residues differ from those used by close relatives of hnRNP A2/B1, hinting that this protein has evolved a unique way of recognizing G‑quadruplex DNA.
From Protein Pairing to Antiviral Defense
Taken together, the findings show that while full-length hnRNP A2/B1 tends to clump into unstable aggregates, carefully trimmed versions behave as orderly single molecules that can pair up when they encounter G‑quadruplex DNA. This controlled dimer formation, driven specifically by guanine-rich single-stranded sequences abundant in many viral genomes, suggests a plausible physical step in how the protein senses invading DNA and helps switch on antiviral signaling pathways. Although these results were obtained in test tubes and still need to be confirmed in living cells and infection models, they provide a clearer, more concrete picture of how changes in protein assembly may be tied to the cell’s early warning system against viruses—and offer a roadmap for designing small molecules that could tune this response in future therapies.
Citation: Shahatibieke, D., Tang, X., Zheng, X. et al. Molecular recognition and induced dimerization of hnRNP A2/B1 truncations by G-quadruplex single strand DNA. Sci Rep 16, 10970 (2026). https://doi.org/10.1038/s41598-026-44646-7
Keywords: hnRNP A2/B1, G-quadruplex DNA, DNA sensing, antiviral immunity, protein dimerization