Clear Sky Science · en
Optimization and comparison research on adaptive modularization BERT-SPAN model for named entity recognition of SARS-CoV-2 virus proteins
Why finding protein names in papers matters
Every week, thousands of new scientific papers about COVID-19 and its virus, SARS-CoV-2, appear online. Hidden in these dense articles are crucial clues about the virus’s proteins—molecular machines that help it infect our cells and that are prime targets for drugs and vaccines. But no human can read everything. This study tackles a simple but powerful question: can we teach computers to automatically spot and list the virus’s proteins in text, quickly and accurately enough to help researchers keep up?
Turning crowded text into usable data
When scientists write about SARS-CoV-2, they use a confusing mix of full names, abbreviations, and nicknames for its proteins. The same protein might appear as “spike protein,” “S,” or buried inside phrases like “receptor-binding domain of spike.” To make sense of such language at scale, computers first need a “corpus” – a carefully labeled collection of sentences where experts have highlighted each protein mention. The authors built such a resource by screening around 200,000 coronavirus abstracts from PubMed and then hand-labeling every SARS-CoV-2 protein mention in relevant sentences. After cleaning and cross-checking, they ended up with 7,159 sentences and 12,660 labeled protein mentions that serve as ground truth for training and testing algorithms.
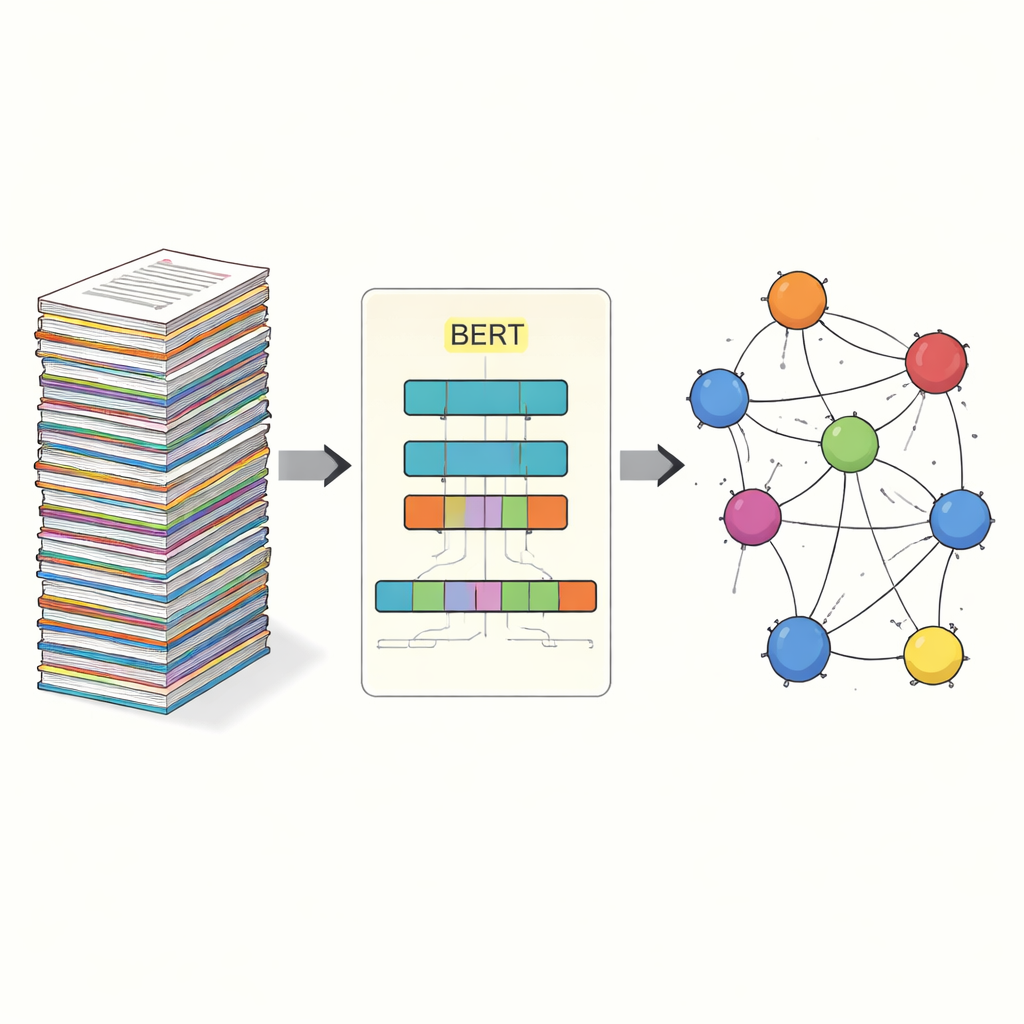
Teaching machines to read virus protein names
The core task in this study is called named entity recognition: getting a computer to pick out specific items—in this case, viral proteins—from free-form text. Earlier approaches relied on manually crafted rules or lists of known terms, which break down when naming is irregular or evolves. Newer systems use deep learning, where a model learns patterns directly from examples. The team builds on BERT, a language model that reads text in both directions at once and learns rich representations of words based on their context. On top of BERT, they use a “SPAN” approach: instead of deciding, word by word, whether each token is inside an entity, the model directly predicts the start and end positions of each protein mention. This makes it easier to handle tricky cases like nested or multi-word names.
Adding smart building blocks to the model
To further sharpen the system, the authors introduce an adaptive modular feedforward layer, which can be thought of as several small expert modules sitting between BERT and the final span detector. Different modules specialize in different kinds of patterns—for example, surface features such as hyphens and mixed letters and numbers, or subtle contextual cues that signal when a short letter like “S” is actually a protein name. An adaptive weighting mechanism then decides, for each sentence, which modules to trust more, combining their outputs into a stronger overall signal before the span prediction step. This design lets the model flexibly focus on the most informative cues for each specific piece of text.
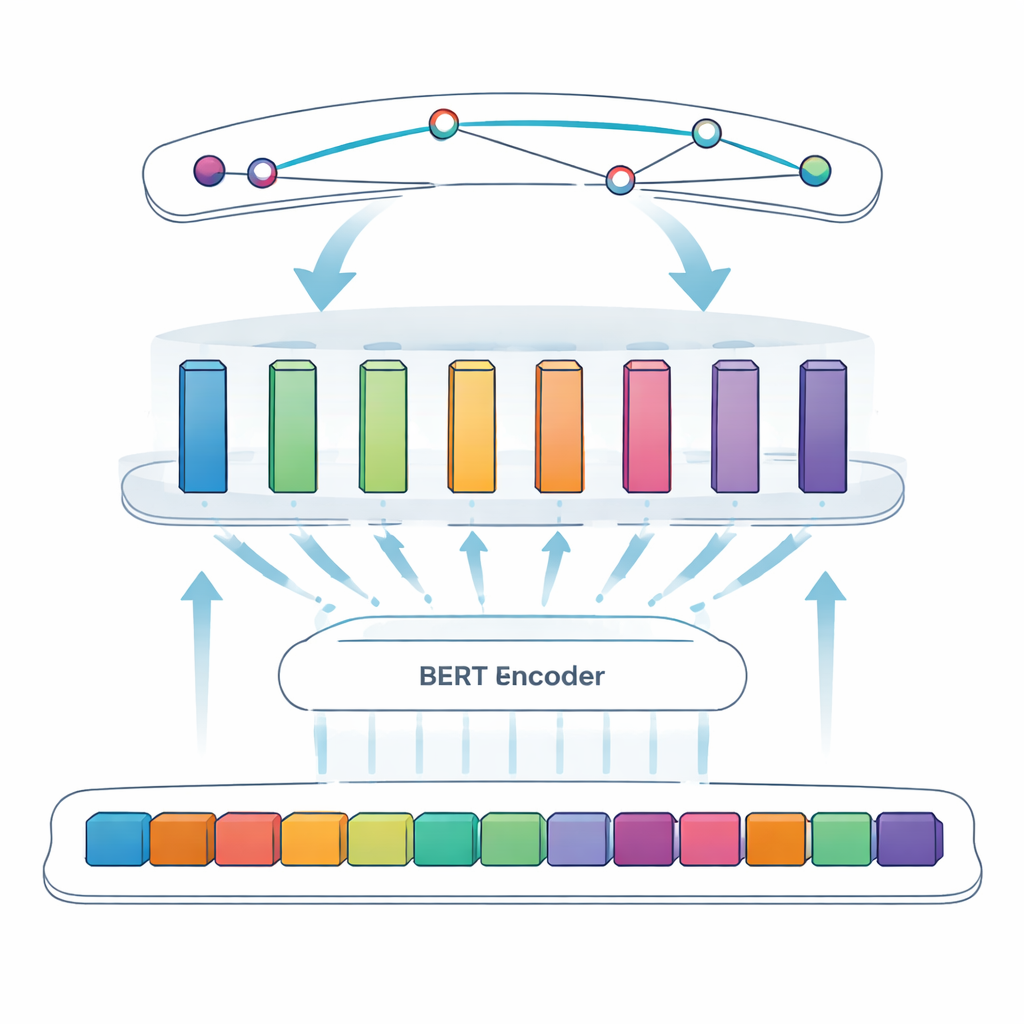
Putting the approach to the test
The researchers rigorously compared their adaptive modular BERT–SPAN model with several established systems, including a popular BiLSTM–CRF model and a standard BERT–SPAN version without modular enhancements. Using standard measures of accuracy, they found that the traditional BiLSTM–CRF model reached an F1 score (a balance of precision and recall) of about 80.8 percent on their SARS-CoV-2 protein dataset. Switching to BERT–SPAN alone raised performance to 91.85 percent. Adding the adaptive modular layer pushed the F1 score further to 93.66 percent, a meaningful gain in such a mature field. Careful “ablation” experiments—removing pieces of the architecture one at a time—showed that both modularization and adaptive weighting contributed to the improvement, especially by reducing boundary mistakes and confusion between similar abbreviations.
How this helps future research
For a lay reader, the key outcome is that the authors built both a high-quality dataset and a smarter reading engine that can reliably recognize SARS-CoV-2 proteins in scientific text. By turning messy language into structured lists of entities, their system makes it much easier to build knowledge graphs, map which viral proteins interact with which host molecules, and track how evidence about particular targets is accumulating. In practical terms, this means that computers can shoulder more of the tedious text-mining work, freeing human experts to focus on interpreting results and designing experiments. The study shows that pairing powerful language models with modular, adaptive components is a promising recipe for sifting through the biomedical literature deluge—not just for coronavirus research, but for many future challenges where science outpaces our ability to read.
Citation: Shi, X., Fei, Z., Nkhata, A. et al. Optimization and comparison research on adaptive modularization BERT-SPAN model for named entity recognition of SARS-CoV-2 virus proteins. Sci Rep 16, 14207 (2026). https://doi.org/10.1038/s41598-026-44317-7
Keywords: SARS-CoV-2 proteins, named entity recognition, biomedical text mining, BERT language model, knowledge graphs