Clear Sky Science · en
A multi-scale adaptive filtering and AtRes_SRU–transformer synergy for breast cancer histopathology classification
Why this matters for patients and doctors
Finding breast cancer early can be the difference between a simple treatment and a life‑threatening disease, yet looking at tissue under the microscope is slow, subjective work. This paper presents a new computer system that helps pathologists sort breast tissue images into harmless and dangerous cases more accurately and efficiently. By carefully cleaning the images and combining several types of artificial intelligence, the system aims to support earlier diagnosis, fewer missed cancers, and fewer unnecessary alarms.
Cleaning the picture before making a call
Microscope images of breast tissue are far from perfect. They contain random speckles, uneven lighting, and color variations from staining, all of which can confuse even advanced algorithms. The authors introduce a front‑end step called multi‑scale adaptive filtering, which acts like a smart, gentle noise filter. Instead of blurring everything equally, it looks at the image at several levels of detail and adapts to local contrast and texture. The result is that distracting noise and staining artifacts are reduced, while important edges—such as boundaries between groups of cells—are preserved. This gives later stages of the system a clearer, more reliable view of each tissue patch.
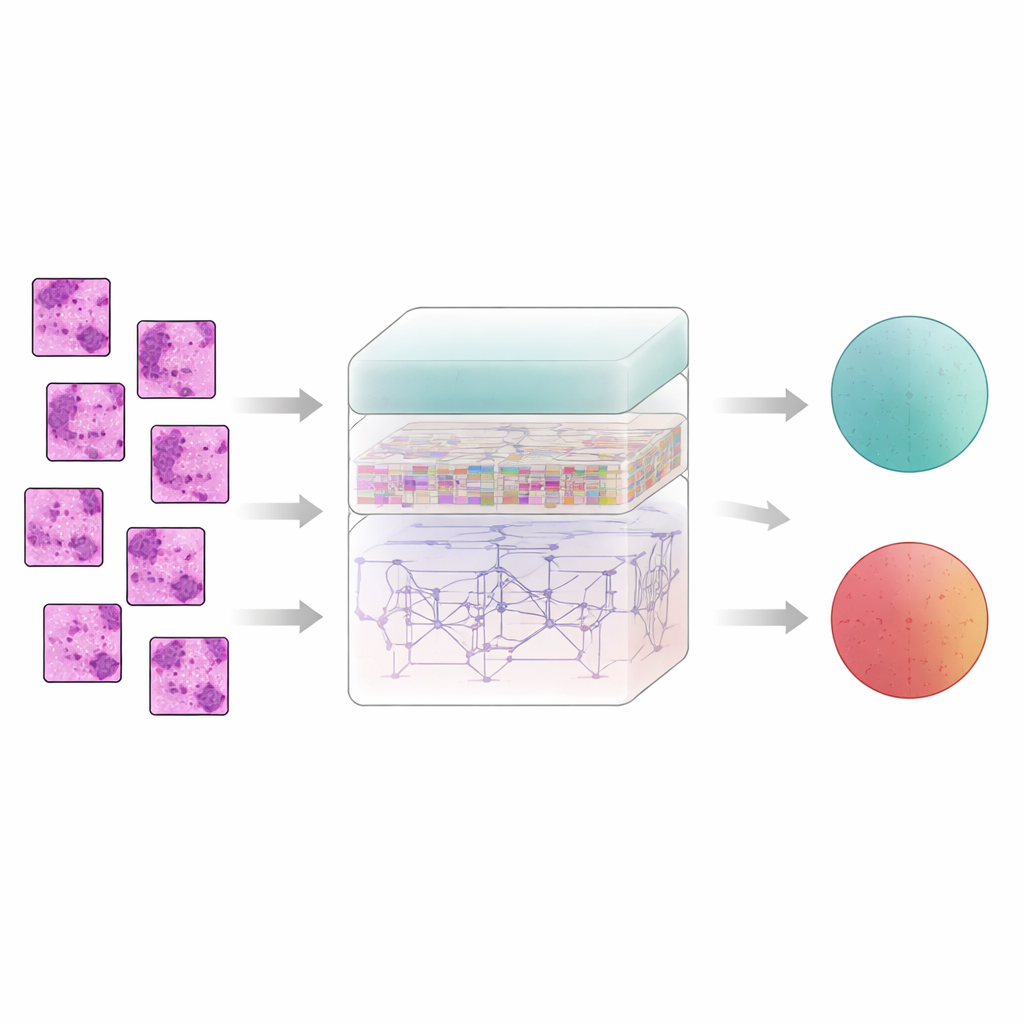
Teaching the computer to follow tissue patterns
Breast tissue does not tell its story one pixel at a time; meaning emerges from how neighboring regions relate to each other. To capture this, the system slices each filtered image into small patches and feeds them into a feature extractor called AtRes_SRU. This part combines a deep residual network, which is good at spotting shapes and textures, with a lightweight recurrent unit that treats adjacent patches like steps in a sequence. Although nothing is moving in time, arranging patches in order lets the model learn how structures flow across the slide—where cell clusters start and end, how glands bend or break, and how abnormal regions sit within more regular tissue. An attention mechanism further nudges the system to focus on the most suspicious regions, such as irregular nuclei or unusually dense cell clusters.
Seeing the whole slide, not just tiny pieces
While local patterns matter, cancer often reveals itself through larger‑scale organization: how scattered abnormal zones are, how wide they spread, and how they interact with surrounding tissue. To grasp this broader picture, the authors add a transformer module, a technology originally developed for language models. Here, each patch becomes a token, and the transformer learns how every patch relates to every other one. A form of absolute position encoding tells the model exactly where each patch came from in the original slide, preserving spatial layout. On top of this, a densely connected decision layer reuses features from many depths of the network, helping the system keep both fine details and global context in mind when it finally decides whether a sample is benign or malignant.
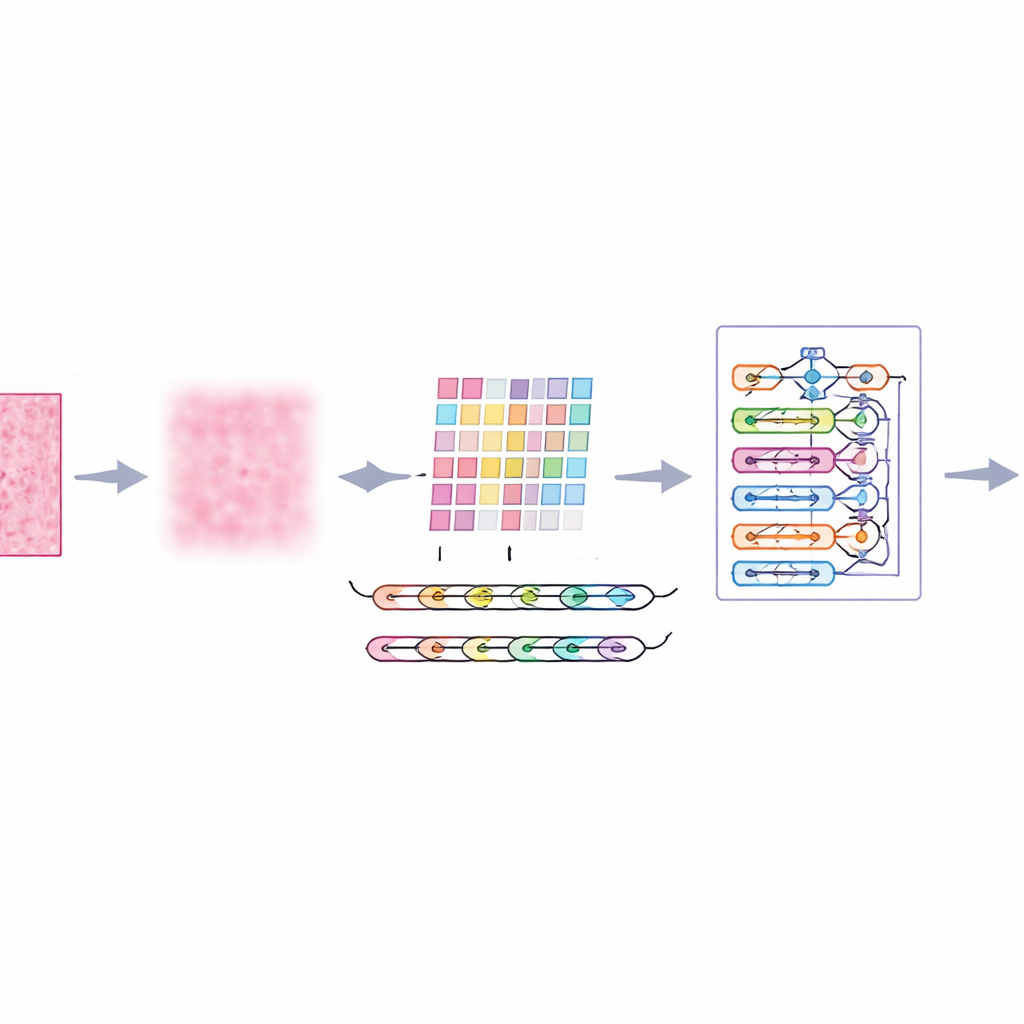
Putting the system to the test
The researchers trained and evaluated their "ImTranNet‑TriCore" framework on a large public collection of breast histopathology images, which includes more than a quarter million small patches from real patients, scanned at several levels of magnification. The dataset is challenging: tissue appearance varies widely across slides, and malignant regions can be subtle or occupy only a small part of an image. Despite this, the new system achieved about 98% accuracy in telling cancerous from non‑cancerous patches and maintained strong performance across all zoom levels. It consistently outperformed a range of widely used methods, including conventional convolutional networks, attention‑enhanced models, and hybrid designs that mix deep learning with traditional machine learning.
From numbers to trust in real clinics
High accuracy alone is not enough for medicine; doctors need to understand why a system reaches its decisions. The authors therefore examined which image regions most influenced the model’s choices using heat‑map and feature‑importance tools. They found that the system tends to highlight the same structures pathologists look at: the shapes and boundaries of cell nuclei, the organization of glands, and dense clusters of abnormal tissue. At the same time, the model keeps false alarms low and shows smoothly calibrated confidence scores, important for avoiding both missed cancers and unnecessary biopsies. While the method still struggles with some very fine‑grained differences at extreme magnification, the study suggests that combining careful image cleaning, local pattern tracking, and whole‑slide context can bring automated breast cancer assessment much closer to the reliability clinicians need.
Citation: Saravana Kumar, N.M., Kandala, M.K., Kaveri, P.R. et al. A multi-scale adaptive filtering and AtRes_SRU–transformer synergy for breast cancer histopathology classification. Sci Rep 16, 14387 (2026). https://doi.org/10.1038/s41598-026-41153-7
Keywords: breast cancer histopathology, medical image analysis, deep learning, transformer models, computer-aided diagnosis