Clear Sky Science · en
Pre-assembly of biomolecular condensate seeds drives RSV replication
How some virus particles get a head start
Respiratory syncytial virus (RSV) is a major cause of lung infections in babies, older adults and people with weakened immune systems. This study asks a deceptively simple question with big implications: when a single RSV particle lands on a cell, what decides whether that infection will take off or quietly stall? By watching individual viral genomes in living cells, the authors uncover hidden differences between virus particles that explain why some infections explode while others fizzle out.
Hidden viral droplets inside infected cells
Once inside a cell, many RNA viruses, including RSV, build tiny, membrane-free “viral factories.” These droplet-like structures concentrate viral components and are the main sites where new viral genomes and proteins are made. But there is a catch: such droplets usually form only when protein levels are already high, while early in infection viral proteins are scarce. Using advanced fluorescence microscopy, the researchers tracked single RSV genome complexes, called vRNPs, from the moment they entered cells. They found that viral factories do not appear out of nowhere; instead, they grow out of individual incoming vRNPs that gradually swell and fuse into larger droplets.
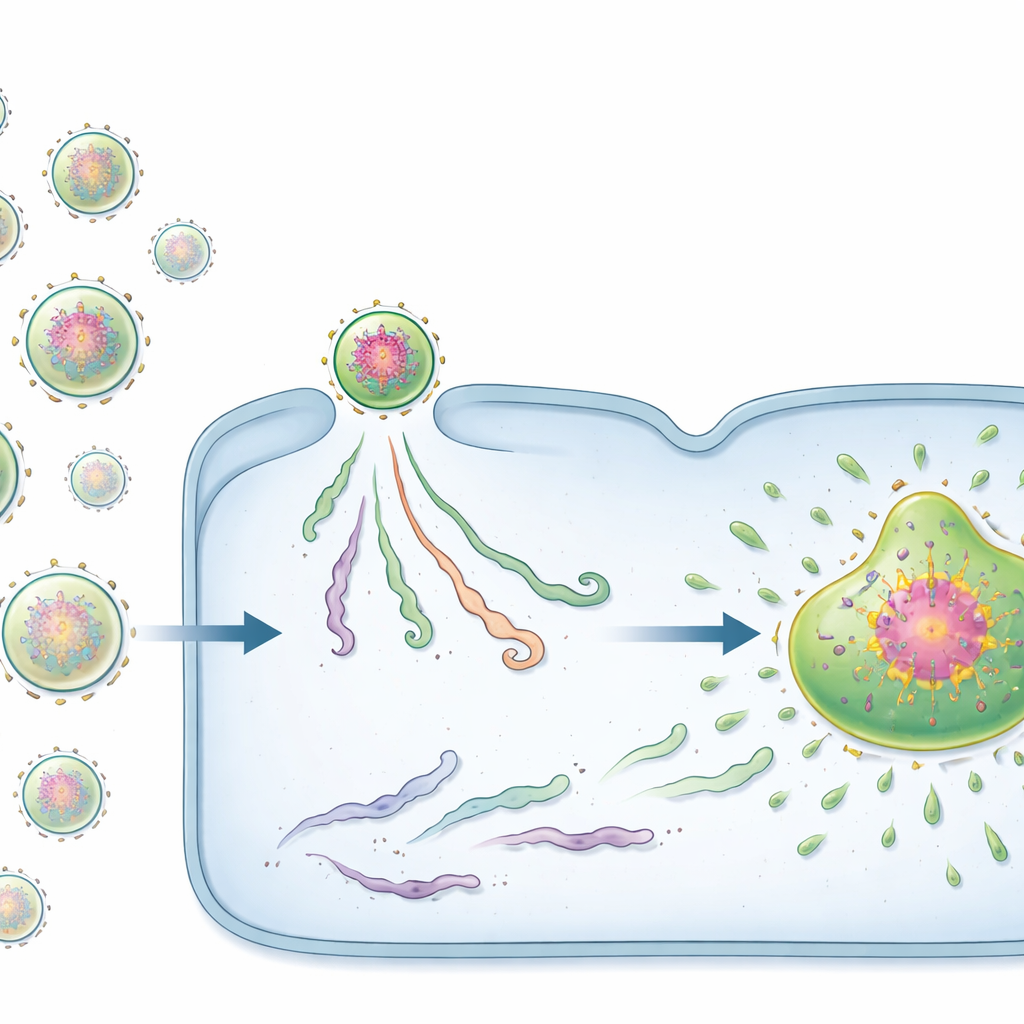
Two kinds of viral starters: seeds and passengers
Surprisingly, not all incoming vRNPs behaved the same. With specially engineered fluorescent probes, the team discovered two distinct states. A minority of vRNPs carried dense clusters of viral proteins already assembled around the genome. These protein-rich complexes, which the authors call pre‑replication centres (PRCs), acted as powerful “seeds” for viral factories. Cells infected by even a single PRC almost always went on to form a factory and produce large numbers of progeny genomes. In contrast, most vRNPs lacked this preloaded protein network and were termed passive vRNPs. Infections started by passive vRNPs frequently stalled: the viral material stayed small and isolated, and no viral factory or progeny genomes appeared.
Why seeds succeed where passengers fail
By measuring viral activity in real time, the authors showed that PRCs begin transcribing viral genes quickly and at far higher rates than passive vRNPs. Detailed imaging and biochemical analyses revealed why. PRCs carry substantially more copies of key viral proteins—especially the polymerase and its co‑factor—per genome than passive vRNPs, even before entering the cell. Once in the cytoplasm, these proteins stay bound to PRCs but fall off passive vRNPs. PRCs also act like sticky scaffolds that pull in additional viral proteins and even other vRNPs. As proteins accumulate, PRCs cross the threshold needed to condense into a droplet‑like viral factory, where genome replication finally switches on. This creates a feed‑forward loop: more proteins make a better seed, which captures still more proteins and rapidly grows into a factory.
Built‑in differences between virus particles
The study further shows that RSV particles released from infected cells are not all equal. Some virions package multiple vRNPs, and many contain either mostly PRCs or mostly passive vRNPs. Virions enriched in PRCs are far more likely to initiate a productive infection in a new cell. This “virion heterogeneity” was seen across lab strains, clinical isolates and virus collected directly from infected people, suggesting it is a natural feature of RSV biology. The number of genomes per particle mattered much less than whether at least one of them was a PRC. Thus, a small subset of well‑equipped particles probably drives most successful infections, while others act as underpowered passengers.
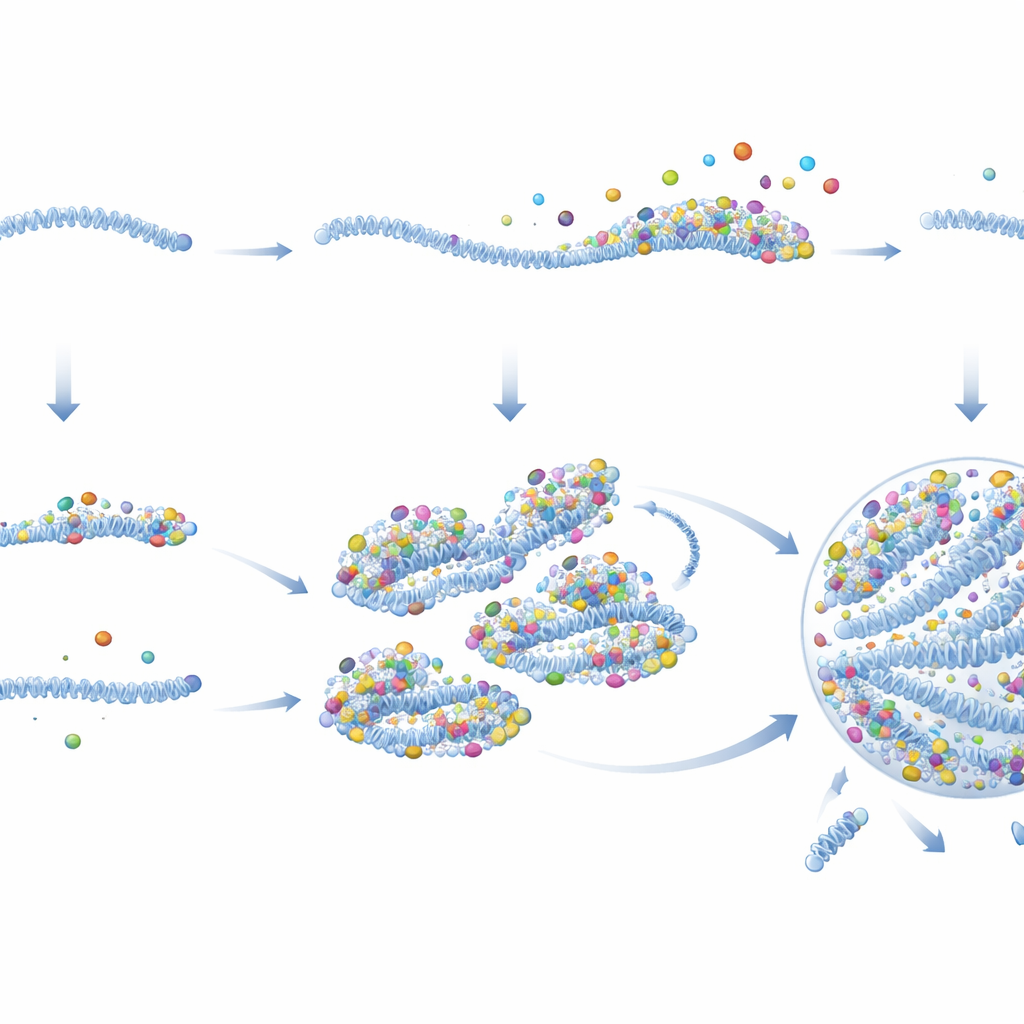
Why this matters for infection and beyond
In simple terms, the authors conclude that RSV solves an apparent chicken‑and‑egg problem—needing high protein levels to build viral factories, but needing factories to make proteins—by pre‑assembling protein‑rich seeds inside some virions before they leave an infected cell. When these seeded genomes enter a new cell, they immediately start producing proteins and gathering more around themselves, quickly nucleating a viral factory and launching replication. Passive genomes, by contrast, rarely escape this low‑activity state unless extra viral proteins are supplied. This work not only clarifies why RSV infections can vary so much from cell to cell, but also suggests new strategies: drugs that disrupt these seeds or their ability to condense could blunt infection at its very first steps, and similar principles may govern how many other disease‑related biomolecular droplets form inside our cells.
Citation: Ratnayake, D., Galloux, M., Boersma, S. et al. Pre-assembly of biomolecular condensate seeds drives RSV replication. Nature 652, 189–200 (2026). https://doi.org/10.1038/s41586-025-10071-5
Keywords: respiratory syncytial virus, viral factories, biomolecular condensates, single-virus imaging, infection heterogeneity