Clear Sky Science · en
Unlocking the potential of computational phenotypic drug discovery: methods, challenges, and future directions
Why this new way of finding medicines matters
Most new medicines fail somewhere between the lab and the clinic, even after years of work and huge costs. This article explains a rising alternative called phenotypic drug discovery, which looks at how compounds change the behavior of whole cells or organisms rather than aiming at one pre‑chosen molecule. With the help of artificial intelligence and large biological datasets, this approach could uncover treatments for complex diseases that have resisted traditional methods.
From lock‑and‑key thinking to watching living systems
For decades, drug discovery has followed a "mechanism‑first" script: pick a single protein thought to drive disease, design or search for chemicals that bind to it, and then hope those interactions translate into real health benefits. Powerful tools such as protein structure prediction, computer‑aided design, and virtual screening have sped up this search. Yet many carefully chosen targets and promising molecules still fail to help patients, because changing one molecule does not always fix a tangled disease network. This gap between success in test tubes and success in people has triggered renewed interest in strategies that focus on what actually happens in living systems.
A compound‑first way to spot helpful changes
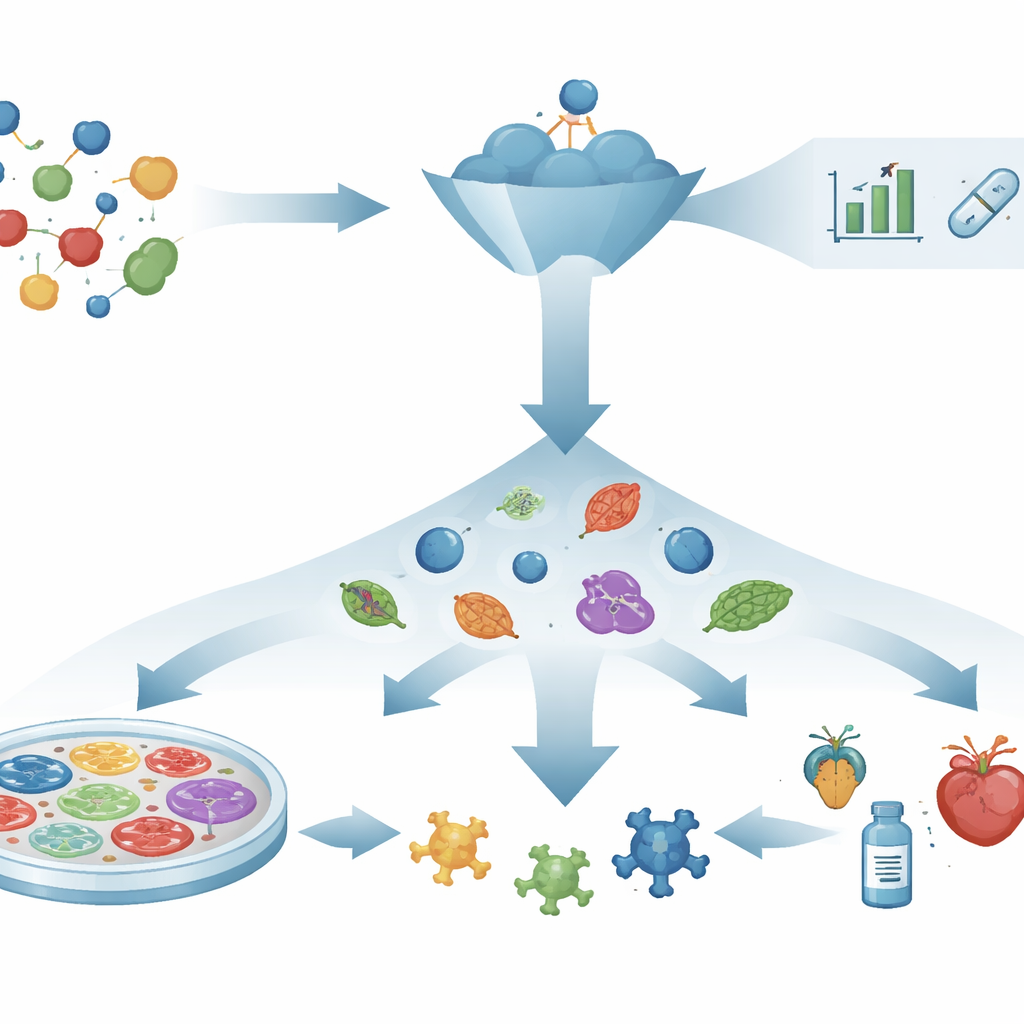
Phenotypic drug discovery flips the script. Instead of starting with a known target, it asks a more direct question: does a compound push a sick system back toward health? Researchers expose cells, tissues, or small organisms to many compounds and look for visible or measurable changes, such as restored cell shape, survival of fragile cells, or recovery of normal behavior. Only after a promising effect is seen do scientists investigate how the compound works inside the cell. This target‑agnostic approach has already led to medicines such as treatments for cystic fibrosis and spinal muscular atrophy, whose detailed mechanisms were worked out later. It is especially useful for diseases driven by many pathways at once, where picking a single "master" target is unrealistic.
Turning rich biological readouts into numbers
Modern phenotypic screens generate enormous amounts of data: high‑resolution images of stained cells, readouts of gene activity, maps of protein changes, and even behavior of model organisms. To make sense of this complexity, the field relies on "embeddings"—ways of converting chemistry, images, gene expression, and networks of interactions into compact numerical fingerprints. These fingerprints capture patterns in how drugs look, how they alter cells, and how they connect to genes and diseases. Machine‑learning models, from classic algorithms to deep neural networks and graph‑based methods, then learn to link these fingerprints to outcomes such as toxicity, likely benefit, or similarity to known drugs. In some cases, they can even suggest new compounds or new uses for existing ones.
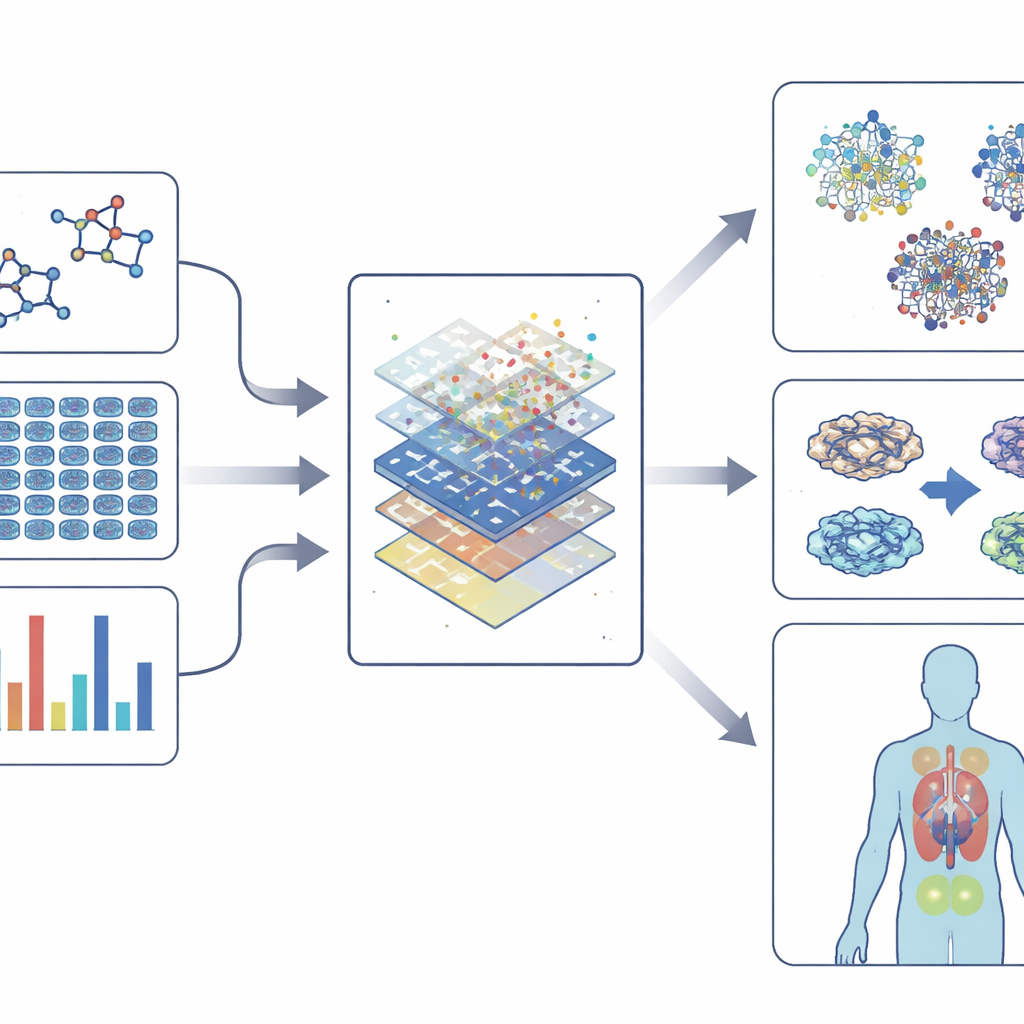
Using smarter models while keeping them understandable
As models become more powerful, they also risk becoming more opaque. Explainable AI tools aim to open this black box by highlighting which features—such as particular gene changes or structural parts of a molecule—drive a prediction. This transparency helps researchers trust the models, design safer compounds, and understand why a phenotypic effect occurs, not just that it does. At the same time, single‑cell and multi‑omics technologies allow scientists to see how different cell types in a tissue respond to a drug, bringing the field closer to personalized treatment choices.
Hurdles on the road to better medicines
Despite its promise, phenotypic drug discovery faces major challenges. High‑quality datasets are still patchy and inconsistent across labs, making it hard to train models that generalize well. Complex phenotypic assays can be difficult to reproduce, and moving from a hit in cells or animals to a safe, effective human therapy often exposes hidden flaws. Untangling how a phenotypic hit works—the process known as target deconvolution—can be slow and uncertain, especially when a compound acts on several pathways at once. Case studies of failed or repurposed drugs underline how missing or misunderstood mechanisms can lead to wasted trials or unexpected side effects.
Where this approach is headed next
The article concludes that the real power of phenotypic drug discovery will come from combining rich biological models with advanced computation. Patient‑derived stem cells, three‑dimensional mini‑organs, and detailed imaging can provide more human‑like test systems, while generative AI may propose molecules tailored to correct specific disease signatures, even in rare conditions with limited data. If issues of data quality, validation, and mechanism‑finding can be addressed, this compound‑first, systems‑aware strategy could make drug discovery faster, more efficient, and more closely aligned with how diseases behave in real patients.
Citation: Kumar, S., Pal, A. & Chatterjee, S. Unlocking the potential of computational phenotypic drug discovery: methods, challenges, and future directions. npj Syst Biol Appl 12, 46 (2026). https://doi.org/10.1038/s41540-026-00676-5
Keywords: phenotypic drug discovery, machine learning in drug discovery, drug repurposing, high-content screening, computational pharmacology