Clear Sky Science · en
Molecular anatomy of PLK1 master docking motifs
How cells keep their division engines on track
Every time a cell divides, it must copy and separate its chromosomes with stunning precision. A key helper in this process is an enzyme called PLK1, which acts like a traffic controller for dividing chromosomes. This study asks a simple but important question: how does PLK1 know exactly where to go, and how does it turn on at the right place and time? By mapping its docking sites in atomic detail, the authors reveal how a few special "master" landing pads give PLK1 both accuracy and power.
Targeted landing spots on chromosomes
PLK1 carries a protein segment called the polo-box domain that acts as a docking module. This domain recognizes short phosphorylated stretches on other proteins, allowing PLK1 to park precisely where its activity is needed during cell division. Earlier work showed that two proteins at the chromosome attachment site, BUB1 and CENP-U, are especially important: if their docking sites are removed, PLK1 no longer appears at those structures. The authors suspected that these "master" docking motifs might share special design rules that distinguish them from the many weaker, more temporary docking sites scattered around the cell.
A relay that builds a super-strong docking site
The team focused on CENP-U, which sits in the inner part of the chromosome’s attachment point. CENP-U has two nearby potential docking motifs, centered on the amino acids called T78 and T98. Biochemical and mass spectrometry experiments showed that a first enzyme, CDK1, readily tags T98 with a phosphate group. This initial tagging attracts PLK1, which then phosphorylates the less favorable T78 site in a "relay" process. Structural data and binding measurements revealed that, once modified, the region around T78 becomes an exceptionally strong landing pad for PLK1, binding about 100 times tighter than the T98 site.
One master site is better than two
At first glance, the presence of two phosphorylated motifs suggested that a pair of PLK1 molecules might bind side by side and stabilize each other. Crystal structures indeed showed two polo-box domains sitting on a single CENP-U chain. However, detailed binding measurements told a different story: adding the second site did not significantly strengthen overall binding and could even make it slightly weaker. Instead, the authors found that the single T78-centered site, once created, is strong enough on its own to serve as the master docking point. Its strength comes from a large contact surface that plugs into several pockets on PLK1, rather than from dimerization.
Shared design of powerful docking motifs
By extending the CENP-U fragment and comparing it with other known PLK1 partners such as BUB1, BUBR1, and PRC1, the researchers saw a recurring pattern. Master motifs use a phosphorylated amino acid to engage a central pocket, but they also place hydrophobic side chains into nearby grooves, including a previously underappreciated "cryptic" pocket and a neighboring hydrophobic cavity. Computer models based on AlphaFold suggest that several high-impact PLK1 partners wrap around the polo-box in a similar way, occupying multiple pockets at once. This extended footprint appears to be a hallmark of master docking motifs that can hold PLK1 nearby for long enough to modify many local targets.
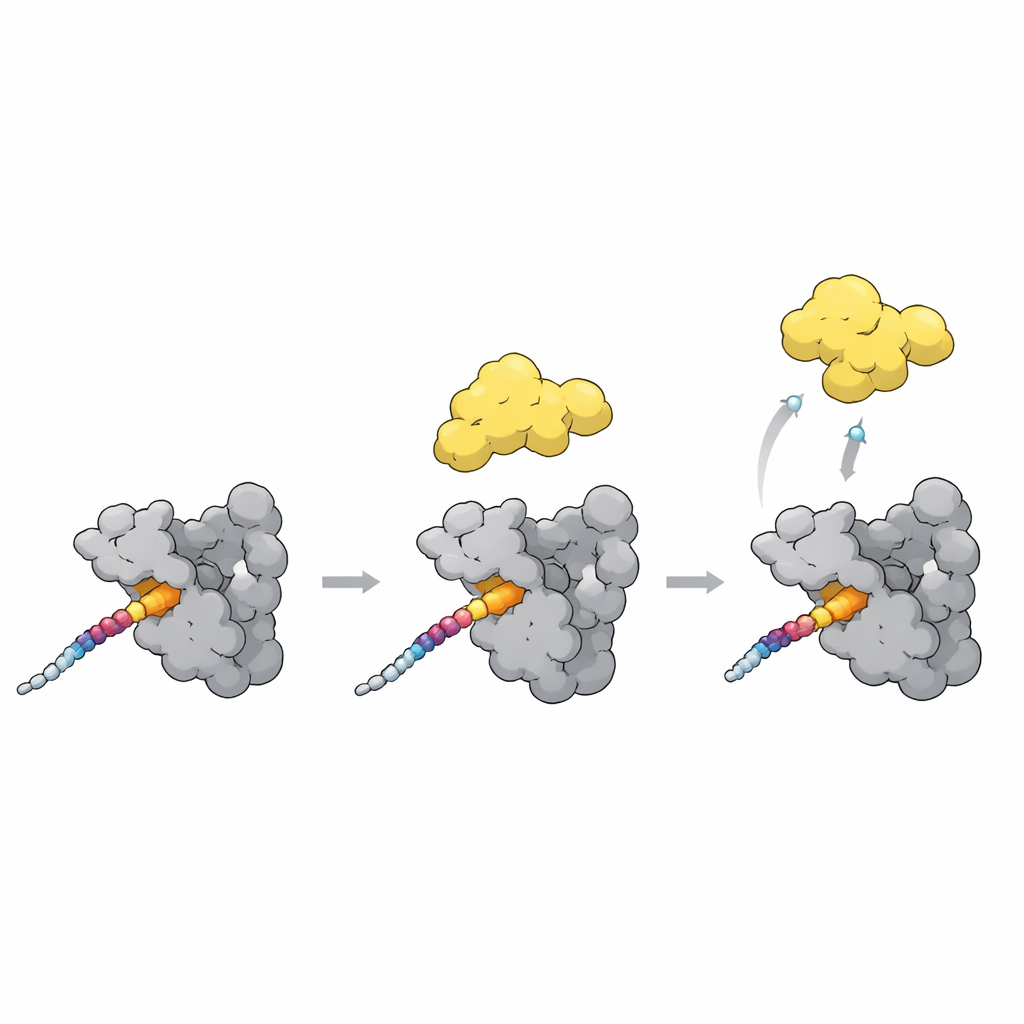
Switching PLK1 from closed to open
A final piece of the puzzle is how docking turns PLK1’s activity up or down. In a closed state, the polo-box domain nestles against the kinase part of PLK1 and keeps it partly shut. The new structures, together with AlphaFold models, point to a flexible loop in the polo-box as a key sensor. When a strong docking motif like the T78 site on CENP-U binds across the polo-box surface, this loop shifts and clashes with the linker that holds the kinase close. That clash likely nudges PLK1 into a more open, active shape. Intriguingly, a natural inhibitor protein binds the same pockets but stabilizes the closed form instead, showing that similar contacts can either unlock or clamp down on PLK1 depending on the exact arrangement.
Why this molecular map matters
In simple terms, this work shows that PLK1 does not need to double up to work effectively at chromosomes. Instead, a single, carefully built master docking site, created in two steps by CDK1 and PLK1, can grab PLK1 very tightly and flip it into an active state. Similar extended docking motifs on other proteins appear to use the same trick. By revealing the molecular anatomy of these powerful landing pads, the study explains how cells achieve precise control over a key division enzyme and offers blueprints for future drugs that might tune PLK1 activity more selectively.
Citation: Ren, L., Esposito-Verza, A., Gasper, R. et al. Molecular anatomy of PLK1 master docking motifs. Nat Commun 17, 4228 (2026). https://doi.org/10.1038/s41467-026-73038-8
Keywords: PLK1, CENP-U, cell division, kinetochore, protein docking