Clear Sky Science · en
Deep visual proteomics uncovers nociceptor diversity and pain targets
Why tiny pain-sensing cells matter
Pain is one of the body’s most important warning systems, yet not all pain-sensing nerve cells behave the same way. Some respond to heat, others to pinch or pressure, and some stay silent until injury or inflammation “wake” them up. This study dives deep into the actual proteins inside individual mouse pain-sensing neurons to understand what makes them different from one another—and to pinpoint new molecular switches that might be targeted to relieve chronic pain.
A closer look at the body’s pain messengers
Bundles of sensory nerve cells in the spine, called dorsal root ganglia, send information about touch, temperature, and injury up to the brain. Within these bundles are many neuron subtypes, including nociceptors that specialize in detecting harmful stimuli. Earlier research used RNA, the cell’s instruction manual, to chart this diversity. But RNA only tells part of the story: it hints at which proteins could exist, not which ones are actually present and active. Proteins, not genes, ultimately shape how a neuron fires and how we feel pain. The authors set out to build a detailed protein-level map of different sensory neuron types, including two major pain-sensing subgroups and touch-sensitive mechanoreceptors.
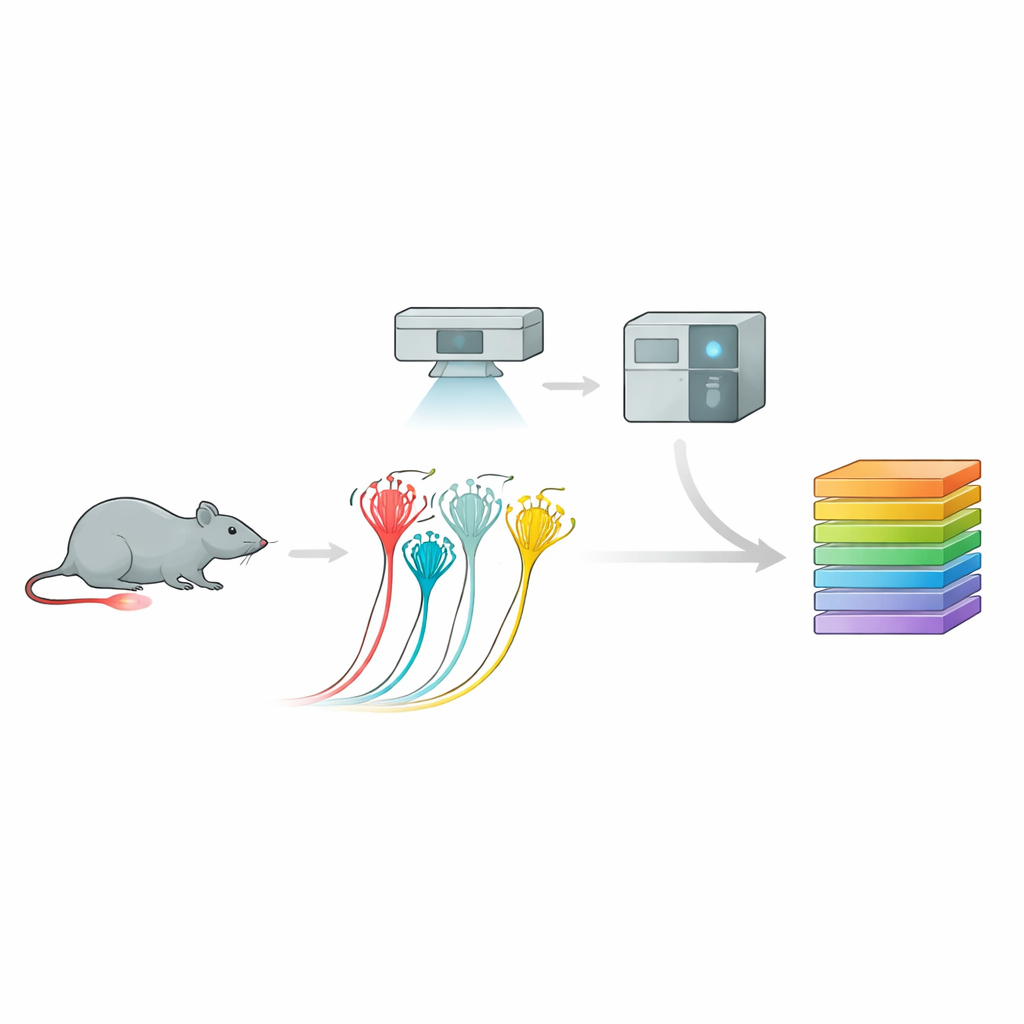
Turning images into protein catalogs
To do this, the team used a powerful pipeline called deep visual proteomics. First, they grew mouse sensory neurons in culture and recorded their electrical responses to pressure, distinguishing peptidergic nociceptors, non-peptidergic nociceptors, and larger touch-sensitive neurons based on their behavior and surface markers. Then, using AI-guided image analysis and laser microdissection, they precisely cut out only the cells of interest. Each tiny sample—down to a fraction of a single neuron—was fed into an ultrasensitive mass spectrometer that identified and quantified thousands of proteins. This approach produced rich protein profiles for each neuron subtype, revealing distinct molecular “fingerprints” that lined up well with, but also extended beyond, previous gene-expression maps.
Pain cell diversity written in proteins
The researchers found that each nociceptor subtype carries a stable core set of proteins, plus its own unique signature. Peptidergic and non-peptidergic nociceptors—two long-recognized pain cell classes—showed clear differences in ion channels, signaling molecules, and metabolic pathways. Many known pain-related proteins appeared, but so did less familiar candidates that strongly marked one subtype over another. Importantly, when the team compared cultured neurons to neurons still embedded in spinal tissue, most proteins overlapped. This suggests that cultured neurons, a mainstay of pain research, largely preserve their in-body identity at the protein level, making them a reliable model for exploring how pain pathways work.
How inflammation “unsilences” pain fibers
The study then zoomed in on a classic pain scenario: inflammatory sensitization. In both animals and people, a growth factor called NGF can make certain nociceptors dramatically more sensitive to mechanical stimuli, a process thought to contribute to the tenderness of inflamed skin or joints. The authors mimicked this by bathing cultured neurons in NGF plus a protein kinase activator, turning previously quiet peptidergic nociceptors into robust responders to mechanical indentation. Comparing the protein profiles of treated and untreated cells, they detected a small set of proteins whose levels rose specifically in sensitized peptidergic neurons. One enzyme, B3GNT2, which helps add sugar chains to other molecules, emerged as a particularly strong candidate.
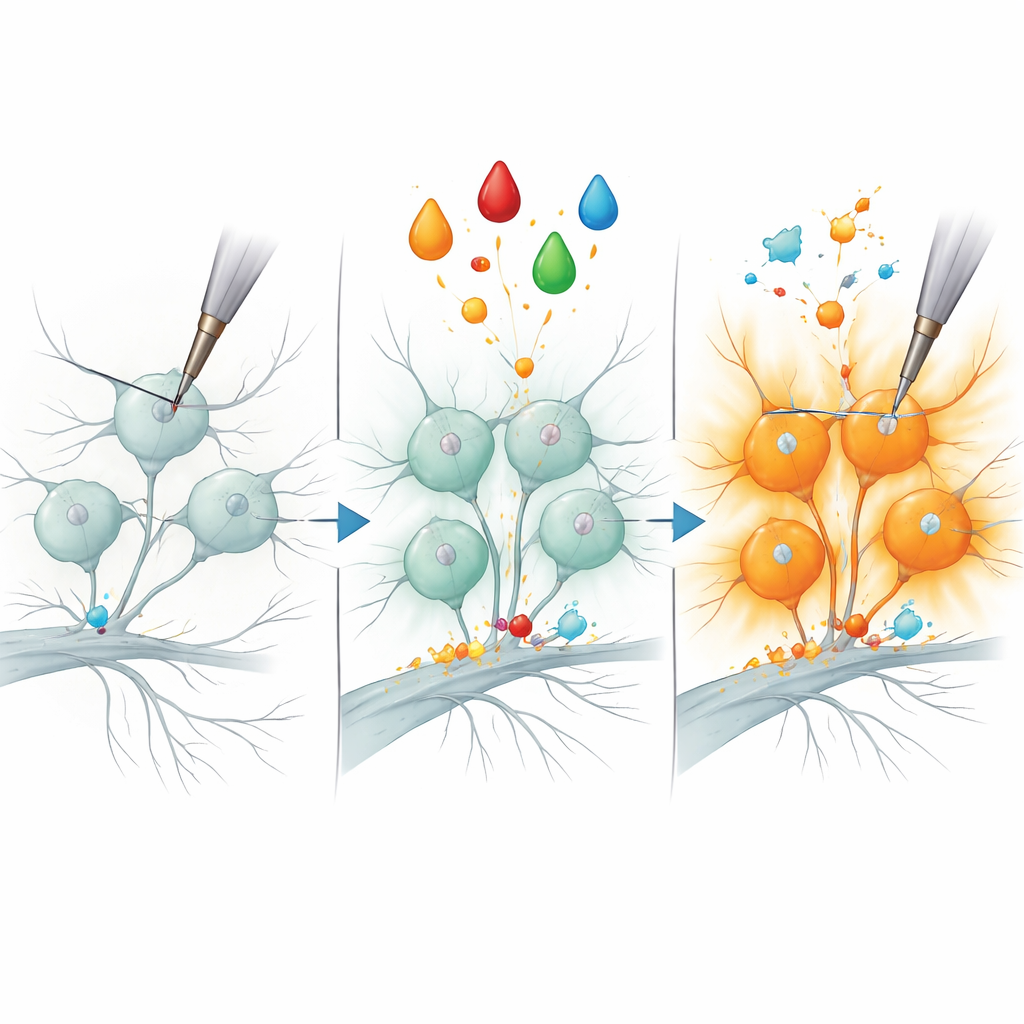
A new molecular handle on pain
To test whether B3GNT2 is more than just a bystander, the team reduced its production in nociceptors using small interfering RNA and then repeated the inflammatory treatment. Strikingly, far fewer neurons became mechanically responsive, suggesting that this enzyme is required for the “unsilencing” of certain pain fibers. While other features of sensitization likely depend on additional proteins, this result links a specific sugar-modifying enzyme to the rapid onset of mechanical hypersensitivity. Because variations in the human B3GNT2 gene have been associated with painful inflammatory conditions, the work positions this protein—and the sugar modifications it performs—as a promising new angle for understanding and eventually treating chronic pain.
Citation: Chakrabarti, S., Makhmut, A., Mohammadi, A. et al. Deep visual proteomics uncovers nociceptor diversity and pain targets. Nat Commun 17, 3437 (2026). https://doi.org/10.1038/s41467-026-71418-8
Keywords: nociceptors, proteomics, chronic pain, sensory neurons, nerve growth factor