Clear Sky Science · en
Toolbox of FRET-based c-di-GMP biosensors and its FRET-To-Sort application for genome-wide mapping of c-di-GMP regulation
How Bacteria Decide When to Stick or Swim
Bacteria constantly choose between roaming freely and settling down in slimy communities called biofilms, a switch that shapes infections, environmental cycles, and even industrial fouling. This paper reveals new tools that let scientists watch a key internal signal in single bacterial cells in real time, and uses them to uncover a surprising link between how hard a bacterium’s tiny propellers spin and its decision to build biofilms.
A Tiny Molecule That Changes Bacterial Lifestyles
Many bacteria rely on a small ring-shaped molecule, c-di-GMP, as an internal switch. Low amounts favor fast swimming, while higher amounts push cells toward settling, sticking to surfaces, and forming biofilms. This signal is made and broken down by families of enzymes that respond to nutrients, oxygen, stress, and more. Until now, however, researchers lacked a single set of sensitive, fast, and broadly tunable sensors to see the full range of c-di-GMP levels inside living cells, especially when the molecule is present at very low concentrations.
Building a Versatile Color-Change Sensor Toolbox
The authors engineered a family of protein sensors based on FRET, a process where one fluorescent protein can transfer energy to another when they sit very close together. They fused pairs of bright fluorescent proteins to a set of bacterial “reader” proteins that change shape when they bind c-di-GMP. Depending on the design, this shape change either brings the fluorescent pair closer or pushes them apart, altering the FRET signal. By screening 90 different reader proteins in Escherichia coli, they identified 16 sensors that respond strongly across a wide span of c-di-GMP concentrations, from about ten billionths of a mole up to a millionth—covering the low levels often found in free-swimming bacteria as well as higher levels in biofilms.
Taking a Closer Look at Single Cells
With this toolbox in hand, the team improved a flow-cytometry method that measures FRET precisely in thousands of cells per second, correcting for optical artifacts so that results can be compared across experiments and sensors. They confirmed the measured binding strengths in two ways: by soaking permeabilized cells in known c-di-GMP solutions and by testing purified sensors in test tubes. They then followed individual bacteria in microfluidic channels, finding that c-di-GMP levels fluctuate strongly from cell to cell and over time, even in genetically identical populations. High-affinity sensors revealed subtle differences in these low concentration ranges that older, weaker sensors largely missed.
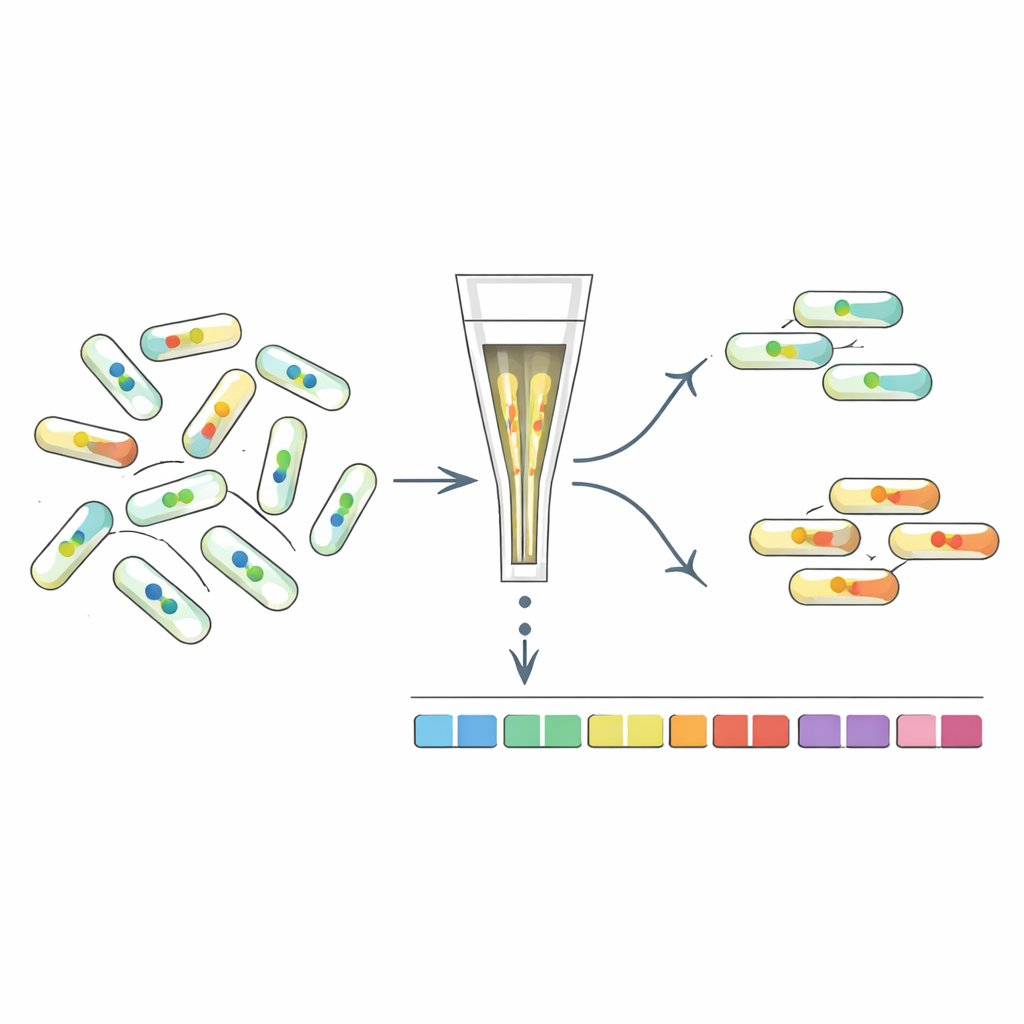
Sorting Mutants by Their Internal Signal
To understand what genes shape c-di-GMP levels, the researchers developed “FRET-To-Sort,” which couples their sensors to high-throughput mutant screening. They introduced selected sensors into a barcoded library of tens of thousands of E. coli mutants, then used a cell sorter to collect cells with unusually high or low FRET signals, indicating altered c-di-GMP. Sequencing the barcodes revealed which genes were disrupted in these outliers. As expected, mutants in known c-di-GMP enzymes were enriched. But the screen also highlighted many other genes, including those involved in the cell envelope, fimbriae (hair-like surface fibers), stress responses, and, strikingly, the rotary flagella that power swimming.
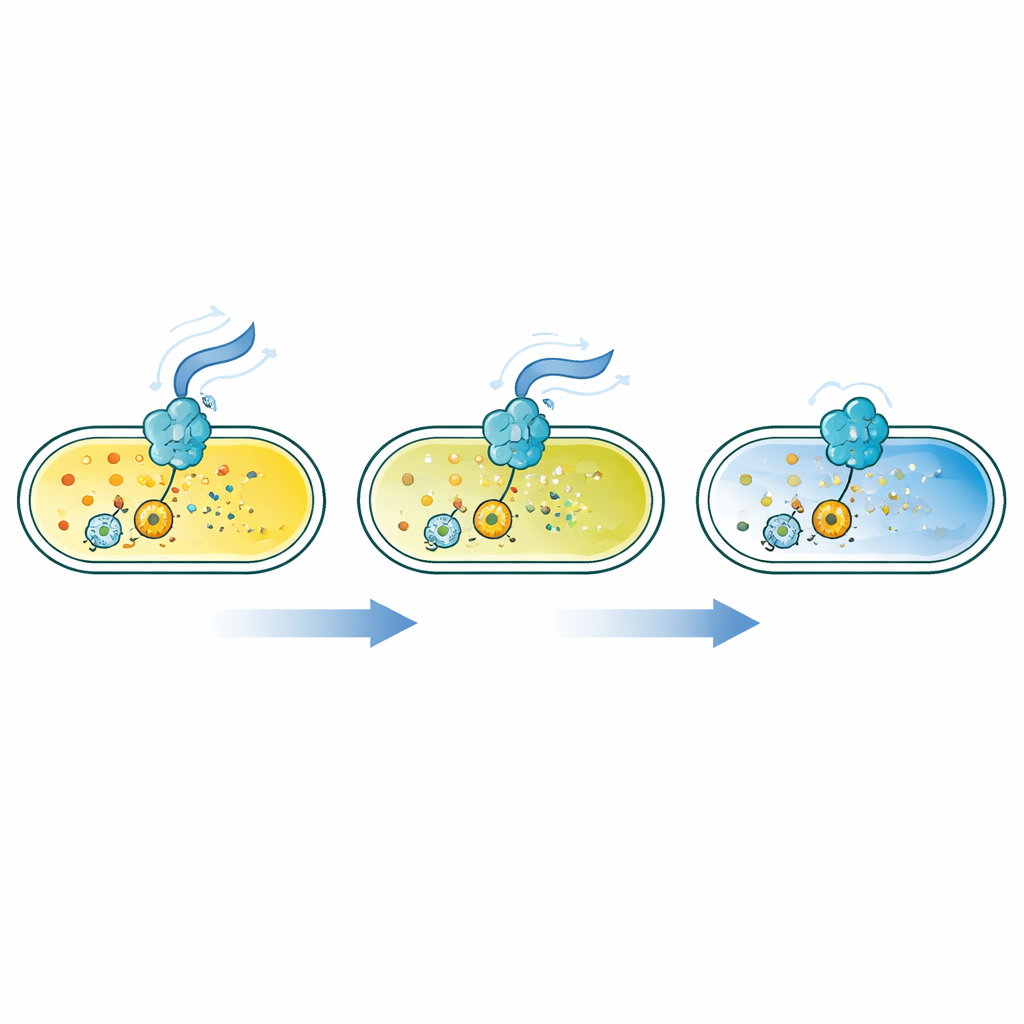
Flagellar Motors as Hidden Signal Knobs
The analysis uncovered a two-faced role for the flagellar system. Mutations that block early steps in building the flagellar base raised c-di-GMP, because they indirectly lowered production of a key enzyme that breaks the molecule down. In contrast, mutations that disabled the spinning parts of the motor or removed the external filament had the opposite effect: they lowered c-di-GMP. Further experiments showed that when flagella cannot rotate, the cell’s membrane potential—the voltage across its outer boundary—rises, and this high voltage dampens the activity of a major c-di-GMP–producing enzyme. Chemically collapsing the membrane potential had the reverse effect, boosting c-di-GMP. Even physically jamming the flagella with specific antibodies reduced c-di-GMP, suggesting that bacteria sense how their motors are loaded and translate this mechanical information into lifestyle decisions.
What This Means for Understanding and Controlling Bacteria
Together, these results provide a standardized set of live-cell c-di-GMP sensors and a powerful strategy to map the genetic networks that feed into this central switch. For non-specialists, the key takeaway is that bacteria do not just respond to chemicals; they also “feel” how easily their tiny propellers turn and adjust their internal signals accordingly. By tying mechanical strain and electrical state at the cell membrane to the molecule that governs sticking versus swimming, this work opens new ways to probe and perhaps disrupt biofilm formation in medically and environmentally important microbes.
Citation: Wang, L., Malengo, G., Sanches-Medeiros, A. et al. Toolbox of FRET-based c-di-GMP biosensors and its FRET-To-Sort application for genome-wide mapping of c-di-GMP regulation. Nat Commun 17, 2955 (2026). https://doi.org/10.1038/s41467-026-71105-8
Keywords: bacterial second messengers, biofilm regulation, FRET biosensors, flagellar motility, genome-wide mutant screening