Clear Sky Science · en
Dynamic subcellular proteomics identifies regulators of adipocyte insulin action
Why this matters for everyday health
Insulin is a hormone best known for keeping blood sugar in check, and when it fails, conditions like type 2 diabetes can develop. This study looks inside fat cells to see, in unprecedented detail, how thousands of proteins quickly shift their positions when insulin arrives. By tracking this molecular reshuffling, the researchers uncover new players that help fat cells respond properly to insulin, offering fresh clues to why insulin sometimes stops working as it should.
A moving map inside fat cells
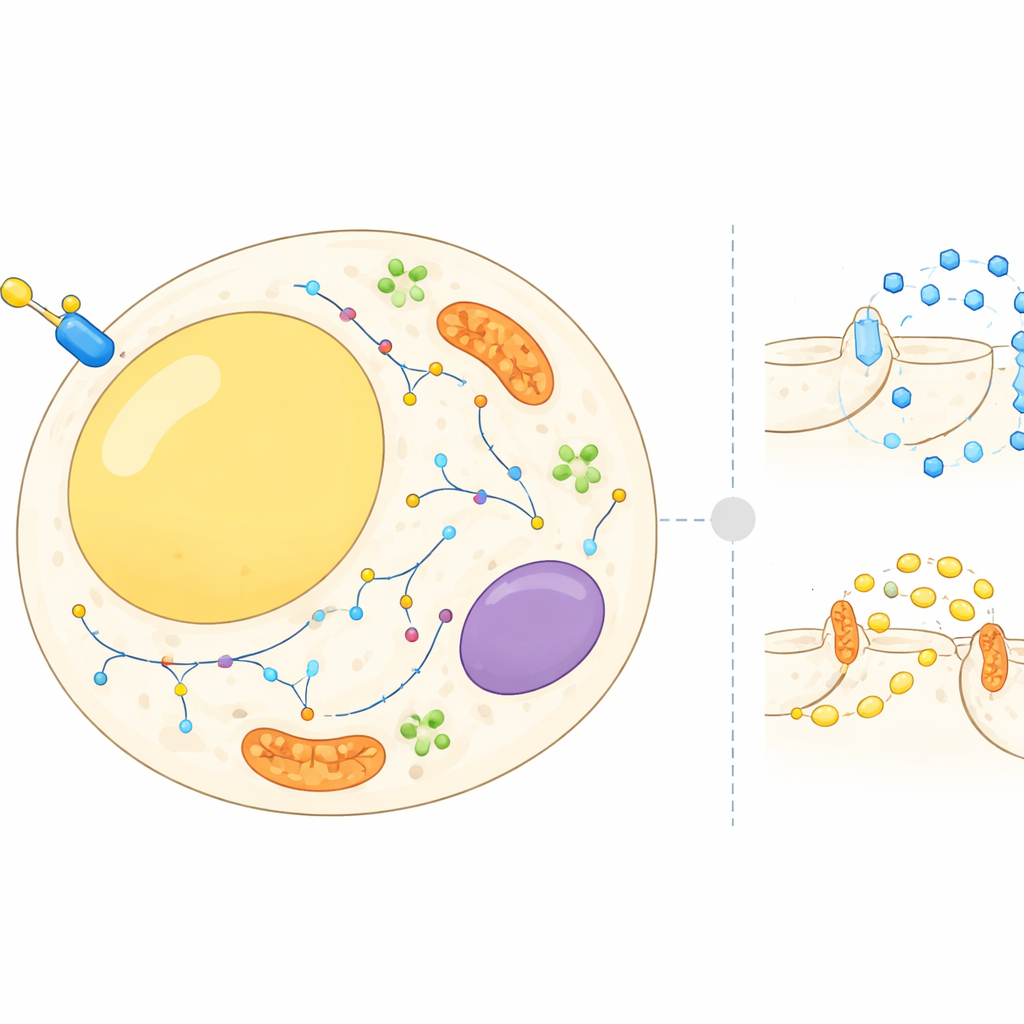
Fat cells are not just passive storage depots; they constantly balance storing and releasing energy in response to hormones and nutrients. Insulin tells these cells to pull sugar out of the bloodstream and to slow the breakdown of stored fat. To do this, many proteins must move between different cellular compartments, such as the cell surface, internal membranes and tiny transport bubbles. Until now, most research focused on a few well-known proteins. In this work, the authors set out to build a cell-wide atlas of where thousands of proteins sit in fat cells before and after an insulin burst, and how their locations change within minutes.
How the protein landscape was charted
The team used an approach called subcellular proteomics, which combines cell fractionation and mass spectrometry to measure proteins in different parts of the cell at once. They worked with 3T3-L1 cells, a standard mouse model of fat cells, and separated each sample into multiple fractions enriched for structures like the cell surface, energy factories, recycling compartments and fat droplets. A statistical framework then compared the distribution of each protein in resting cells versus insulin-stimulated cells. In parallel, the researchers used a separate method that selectively tagged proteins exposed on the outer surface of the cell, letting them quantify how much of each surface protein increased or decreased after insulin treatment.
Insulin sparks widespread protein reshuffling
The combined datasets revealed that insulin triggers a striking reorganization: more than 500 proteins showed high-confidence shifts in their location. Many belonged to known signalling pathways, including components that sense nutrients and regulate growth, as well as protein kinases that switch other proteins on or off. Others were involved in the traffic of cargo-carrying vesicles that shuttle molecules between internal compartments and the cell surface. Some transport complexes moved noticeably in response to insulin, while others stayed put, suggesting that the hormone targets specific branches of the cell’s transport network. The surface-focused experiments showed that over 300 proteins changed in abundance at the outer membrane, confirming that the cell’s “interface” with its environment is reshaped during an insulin response.
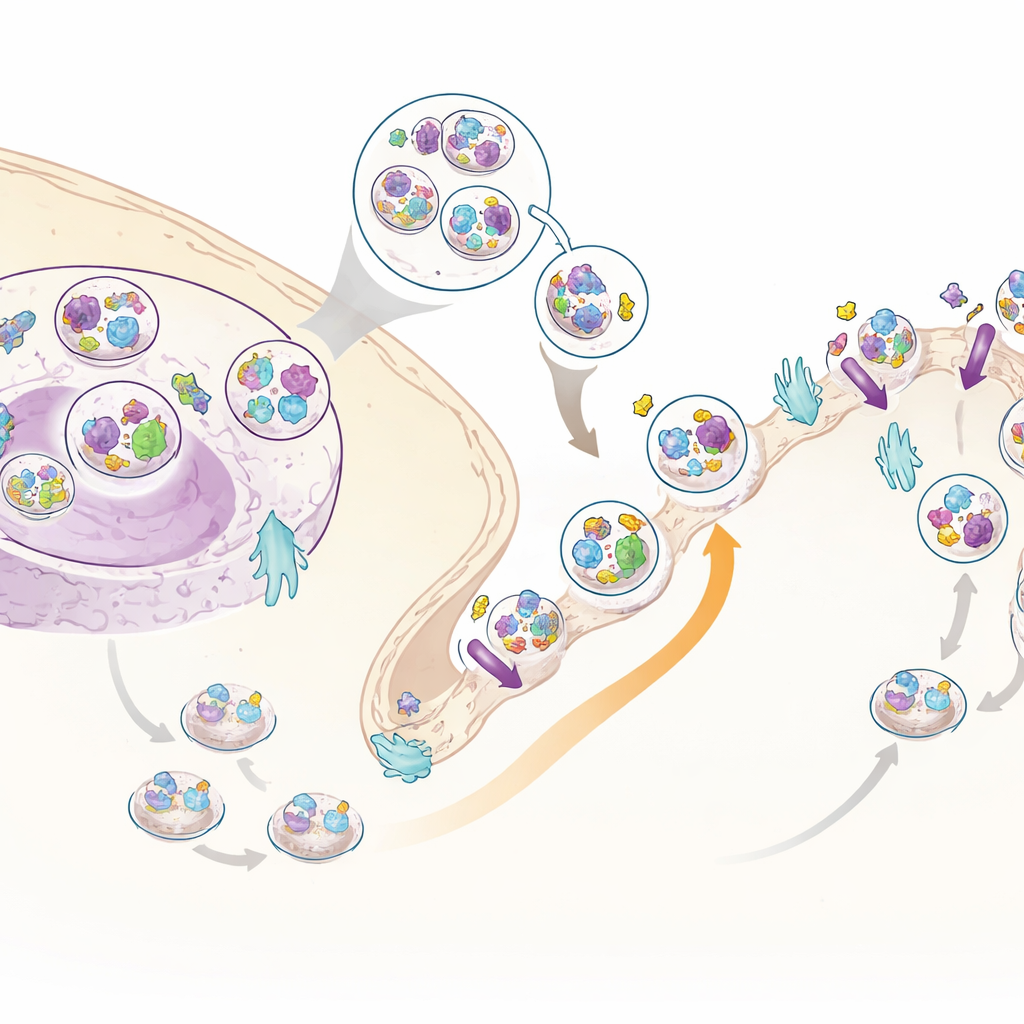
A new insulin helper: C3ORF18
Among the many moving proteins, one stood out: a little-studied protein called C3ORF18. It was strongly pulled toward the cell surface by insulin in both the whole-cell and surface-specific datasets. Microscopy showed that C3ORF18 sits near the cell’s center and in small vesicles, many of which overlap with those carrying the transferrin receptor, a well-known recycling protein, and with GLUT4, a key sugar transporter. When insulin or growth factors were added, C3ORF18 rapidly moved to the cell surface in a way that depended on the same signalling branch used by GLUT4. This behavior suggests that C3ORF18 travels in an insulin-sensitive recycling route that helps deliver important transporters to the membrane.
What happens when the new player is missing
To test whether C3ORF18 does more than just tag along, the researchers selectively reduced its levels in mature mouse and human fat cells. Over several days, cells lacking C3ORF18 became less responsive to insulin: sugar uptake fell, and both GLUT4 and another sugar transporter, GLUT1, were less abundant. Insulin-driven movement of GLUT4 and the transferrin receptor to the surface was also blunted, and core signalling steps downstream of the insulin receptor were weakened. Early after C3ORF18 depletion, problems appeared first in the recycling of transferrin receptors, hinting that disruptions in endosomal recycling may be the starting point that eventually undermines insulin’s broader effects. Data from human fat tissue further supported this view: people with higher blood sugar and long-term glucose markers tended to have lower C3ORF18 expression, and its levels rose with weight loss.
Big-picture takeaways for understanding insulin sensitivity
This study shows that a successful insulin response in fat cells depends not only on which proteins are present, but also on where they are in the cell at any given moment. Insulin rapidly redistributes hundreds of proteins, especially at the cell surface and within recycling compartments, and these location changes work alongside chemical modifications like phosphorylation. By systematically mapping this movement, the authors discovered C3ORF18 as a previously unrecognized helper that supports the recycling routes needed for efficient sugar uptake and healthy insulin signalling. Understanding such spatial dynamics opens new avenues for exploring why insulin sensitivity erodes in metabolic disease and may eventually point toward targets to preserve or restore this crucial hormone response.
Citation: Conway, O.J., Christopher, J.A., Breckels, L.M. et al. Dynamic subcellular proteomics identifies regulators of adipocyte insulin action. Nat Commun 17, 3310 (2026). https://doi.org/10.1038/s41467-026-70116-9
Keywords: insulin resistance, adipocyte, protein trafficking, spatial proteomics, glucose uptake