Clear Sky Science · en
Co-transcriptional folding orchestrates sequential multi-effector sensing by a glycine tandem riboswitch
How RNA Switches Help Bacteria Make Smart Choices
Bacteria live in constantly changing environments, yet they must decide in real time which genes to turn on or off to save energy and survive. This study uncovers how a tiny RNA “switch” inside the bacterium Bacillus subtilis listens to multiple molecular cues at once—an amino acid, metal ions, and a protein cofactor—and then rapidly chooses whether to keep reading a gene or to shut it down. Understanding this built-in decision circuit not only reveals fundamental rules of gene control, but could also inspire new tools for synthetic biology and antibiotic design.
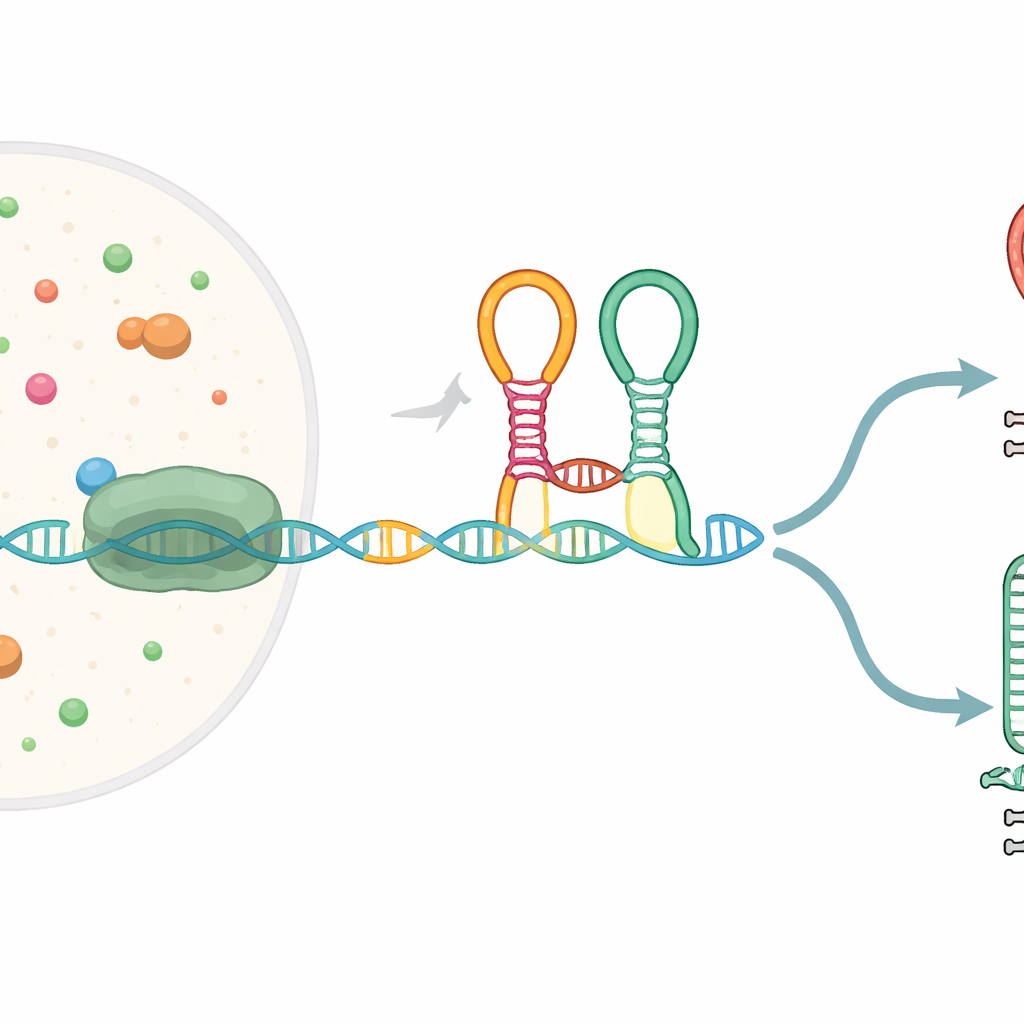
A Tiny Sensor Built Into the Gene Message
Many bacterial genes begin with special RNA segments called riboswitches, which fold into shapes that can sense small molecules and, in response, change gene expression. The riboswitch studied here, known as the glycine tandem riboswitch, sits in front of genes that help break down the amino acid glycine. It contains two adjacent sensing modules (“aptamers”) that both recognize glycine and are linked by a sharp bend called a kink-turn. As the RNA message is being copied from DNA by RNA polymerase, this riboswitch folds on the fly and can steer the process toward either finishing the full-length transcript or cutting it short at a built-in stop signal.
Pauses That Create Decision Checkpoints
The authors found that the RNA-copying enzyme does not move smoothly through this region; instead, it pauses at three specific spots within the riboswitch. These pauses act like traffic lights, giving the emerging RNA time to fold and interact with molecules around it. At the first pause, the upstream aptamer forms and becomes ready to bind glycine. At later pauses, parts of the second aptamer and the final decision helix emerge. A helper protein called NusA lengthens two of these pauses, especially when glycine is present, effectively widening the time window during which the riboswitch can sense its environment and commit to either continuing transcription or terminating it.
Stepwise Sensing of Ions, Glycine, and a Protein Partner
Using single-molecule fluorescence microscopy and high-throughput chemical probing, the team showed that the riboswitch does not act in a single snap. First, potassium ions stabilize the kink-turn at the RNA’s base, orienting the first aptamer toward the path of the growing transcript. Glycine then binds to this first aptamer, tightening its local structure. These events encourage long-range contacts between the two aptamers, which pre-arrange both glycine-binding pockets even before the second glycine arrives. At the final pause near the built-in terminator hairpin, a second glycine can bind to the downstream aptamer in this pre-organized environment, and NusA-enhanced pausing further favors completion of this compact, active configuration.
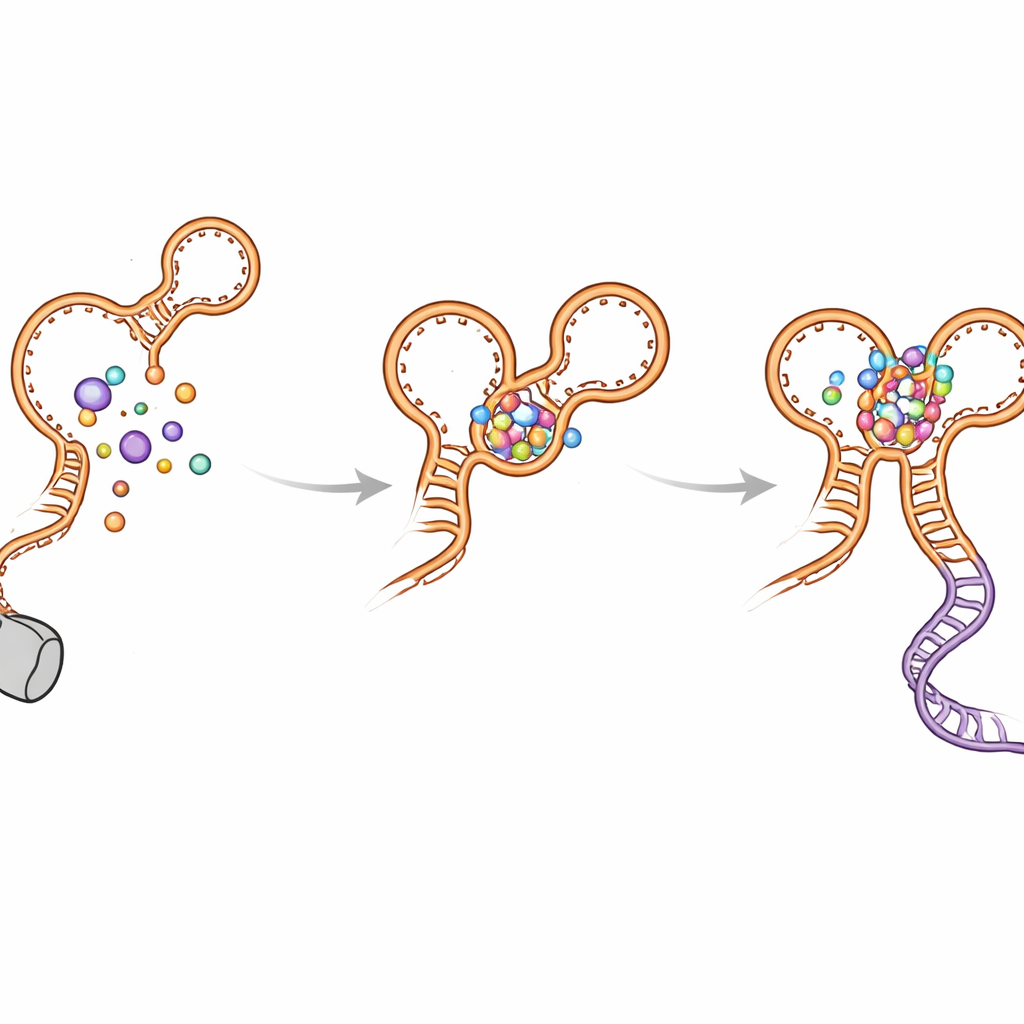
Folding Detours and How the Switch Avoids Getting Stuck
The authors also discovered several “wrong turn” hairpins that form temporarily as the RNA grows. These non-native structures appear in both aptamers and can delay the formation of the final, active shape. The kink-turn helix (called P0) and the inter-aptamer contacts help resolve these intermediates and steer folding toward the functional layout. When P0 or key contact points are disrupted by mutations, the riboswitch loses much of its sensitivity and efficiency: glycine still binds, but the long-range docking is weakened, the decision helix is less stable, and transcription readthrough drops. Computer simulations supported these observations, showing that the riboswitch is much more compact and stable when P0 and both glycine sites are intact.
A Non-Equilibrium Logic Gate in Living Cells
Taken together, the results support a picture in which the glycine tandem riboswitch behaves as a kinetic logic gate rather than a simple equilibrium sensor. Instead of both glycine molecules binding simultaneously to a pre-formed structure, binding and folding occur in a precise 5′-to-3′ sequence, gated by transcription pauses and shaped by potassium ions and the NusA protein. This stepwise, out-of-equilibrium process lets the bacterium integrate several signals—cellular ion balance, glycine levels, and transcription speed—before deciding whether to fully express the downstream glycine-cleaving genes. The work suggests that such kinetically driven cooperativity may be a general strategy by which RNA switches coordinate multiple cellular factors to finely tune gene expression.
Citation: Romero, R.A., Chauvier, A., Teh, S.S. et al. Co-transcriptional folding orchestrates sequential multi-effector sensing by a glycine tandem riboswitch. Nat Commun 17, 2779 (2026). https://doi.org/10.1038/s41467-026-69648-x
Keywords: riboswitch, glycine sensing, RNA folding, transcription pausing, bacterial gene regulation