Clear Sky Science · en
Structures of the eIF4G-binding RNA domains among picornaviral IRES types are topologically conserved
How Certain Viruses Hijack Our Cells
Many common viruses, including those that cause colds, heart inflammation, and hepatitis, face the same basic challenge: how to trick our cells into making viral proteins instead of their own. This study uncovers, in atomic detail, how a key piece of viral genetic material folds into a precise three‑dimensional shape that latches onto a host protein and kick‑starts viral protein production. Understanding this shared structural trick across different viruses could point the way to broad‑spectrum antiviral drugs that target them all at once.
A Special Entry Point for the Protein Factory
Most of our cellular messages (mRNAs) begin with a chemical “cap” that helps recruit the cell’s protein‑making machinery. Many RNA viruses in the Picornaviridae family lack this cap, yet they still manage to commandeer the same machinery. They do this using internal ribosome entry sites, or IRES elements—structured RNA segments near the front of the viral genome that serve as alternative landing pads for the ribosome. Within these IRES regions, one particular section, called domain V in several viruses, binds strongly to a host initiation factor named eIF4G, a kind of scaffold that organizes other components needed to begin protein synthesis.
A High-Resolution Look at a Viral RNA Knot
The authors determined a 3‑angstrom crystal structure of domain V from coxsackievirus B3, a virus often used as a model for human enteroviruses. Rather than being a simple hairpin, this RNA folds into an elongated four‑way junction—imagine two stacked rods crossing another short rod at the center, forming an H‑shaped junction held together by an A‑rich loop. This compact knot of helices positions small bulges and loops at just the right angles to form a rigid scaffold. Solution experiments, including small‑angle X‑ray scattering and chemical footprinting, together with modern structure prediction tools, showed that this fold is not an artifact of crystallization but reflects how the RNA behaves in water. 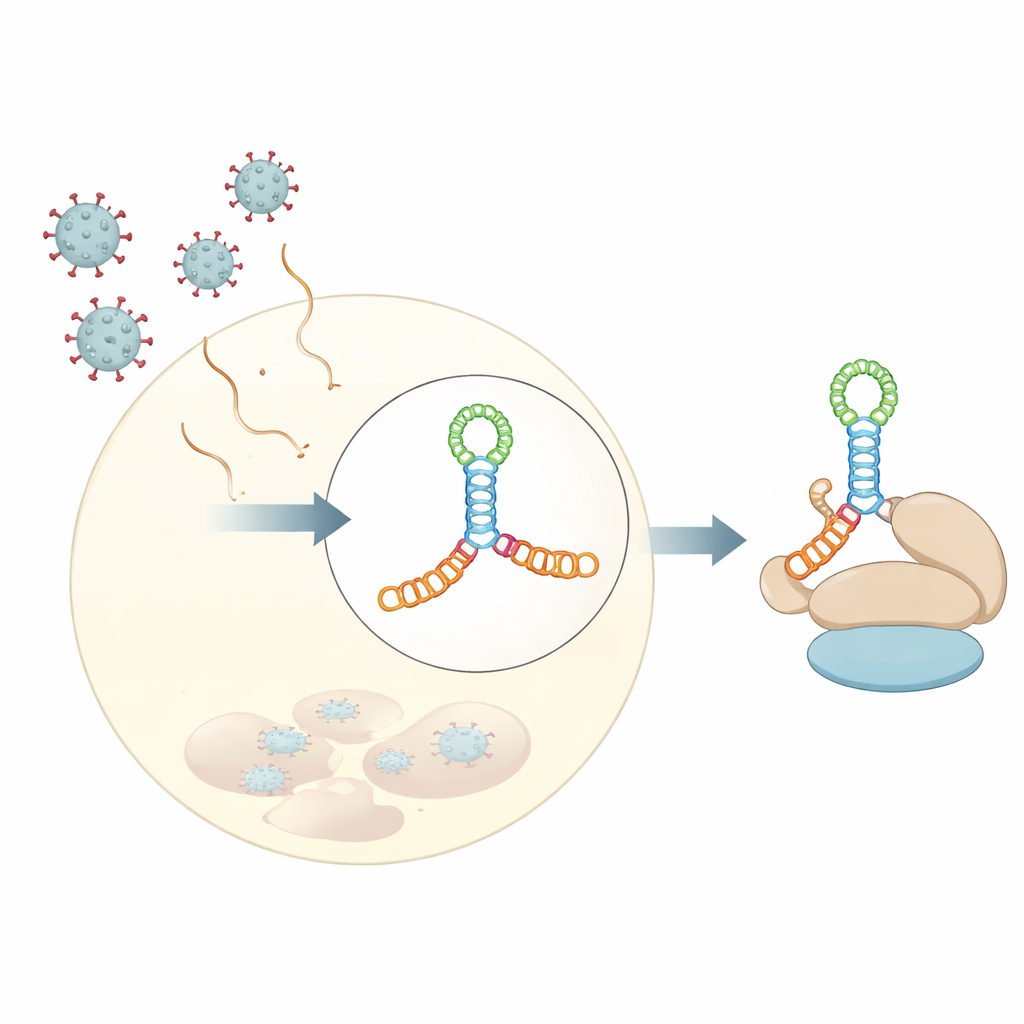
Different Viruses, Same Structural Blueprint
Strikingly, when the team compared this structure to previously solved RNA segments from two other picornaviruses—the JK domain of encephalomyocarditis virus and domain V of hepatitis A virus—they found that the overall three‑dimensional topology is nearly the same, even though the underlying RNA sequences differ considerably. In all three cases, an A‑rich loop docks into the side of a nearby helix and organizes a compact junction where two main stems stack on top of each other. This recurring layout appears to be a structural “motif” that evolution has reused across viral species to present host factors with a familiar binding surface, despite wide variation in sequence and fine details.
How the Viral Fold Grabs a Host Factor
To understand how this shared RNA shape captures eIF4G, the researchers measured binding between the viral RNAs and the HEAT1 region of eIF4G, a curved module known to contact RNA. All three viral domains bound HEAT1 with nanomolar affinity, meaning very tight interactions. By introducing targeted mutations into the RNA junctions and loops, they could weaken or abolish binding and see which features mattered most. Changes that removed small bulges in key helices or disrupted the A‑rich loop greatly reduced binding, even though the mutated nucleotides are not predicted to touch the protein directly. This suggests that the loop and bulges act as internal struts, fixing the relative orientation of the stacked helices so that a continuous lateral surface is perfectly shaped for HEAT1 to dock. 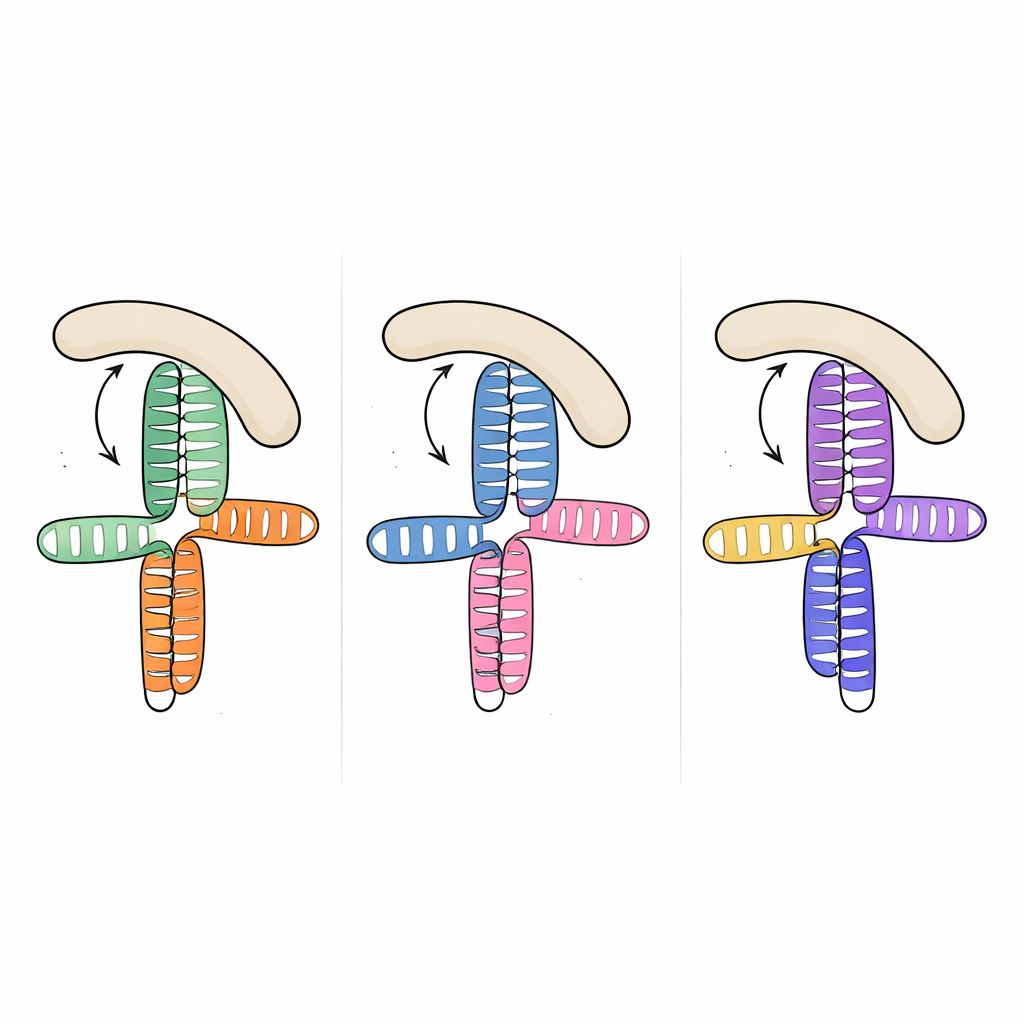
Shared Mechanics and New Drug Possibilities
Chemical “footprinting” experiments, which map which parts of the RNA backbone are protected when the protein is bound, confirmed that HEAT1 primarily contacts one side of the stacked helices opposite the organizing A‑rich loop, in a way that closely mirrors cryo‑electron microscopy observations for one of the other viruses. Taken together, the work shows that several distinct picornaviruses use a common structural strategy: they build similar RNA scaffolds that present conserved three‑dimensional features to the same host factor, eIF4G, even though the sequences themselves vary. For a non‑specialist, the takeaway is that viruses from different branches of the family tree converge on the same folding trick to plug into the cell’s translation hardware. Because this trick depends on shape more than sequence, it may be possible to design drugs or molecules that recognize and disable this conserved RNA architecture, potentially blocking protein production across multiple viral species with a single treatment.
Citation: Banna, H.A., Das, N.K., Kalinina, M. et al. Structures of the eIF4G-binding RNA domains among picornaviral IRES types are topologically conserved. Nat Commun 17, 2758 (2026). https://doi.org/10.1038/s41467-026-69554-2
Keywords: viral RNA structure, internal ribosome entry site, eIF4G binding, picornavirus translation, broad-spectrum antivirals