Clear Sky Science · en
A Perturb-seq map of a differentiation hub reveals synergistic vulnerabilities in KMT2A-rearranged acute myeloid leukemia
Why this leukemia study matters
Acute myeloid leukemia is a fast moving blood cancer that often resists modern targeted drugs. Many new treatments try to shut down single cancer drivers, only to see the disease adapt and return. This study asks a crucial question for patients and families: instead of hitting one switch at a time, can we map the whole control panel that keeps leukemia cells trapped in an immature state, and then find weak spots where combination therapies work far better than any single drug?
The cancer cells that refuse to grow up
In this work, the scientists focus on a high risk form of acute myeloid leukemia marked by changes in a gene called KMT2A. In these cancers, blood stem cells are frozen in an immature phase instead of maturing into infection fighting white blood cells. Earlier drugs that target individual epigenetic regulators, which act like chemical tags on DNA packaging, showed promise but rarely led to lasting remissions. The team reasoned that the leukemia cells survive because several epigenetic systems cooperate, so blocking just one leaves others to pick up the slack. To truly understand this cooperation, they needed a way to watch what happens inside thousands of individual cells when key regulators are turned off one by one.
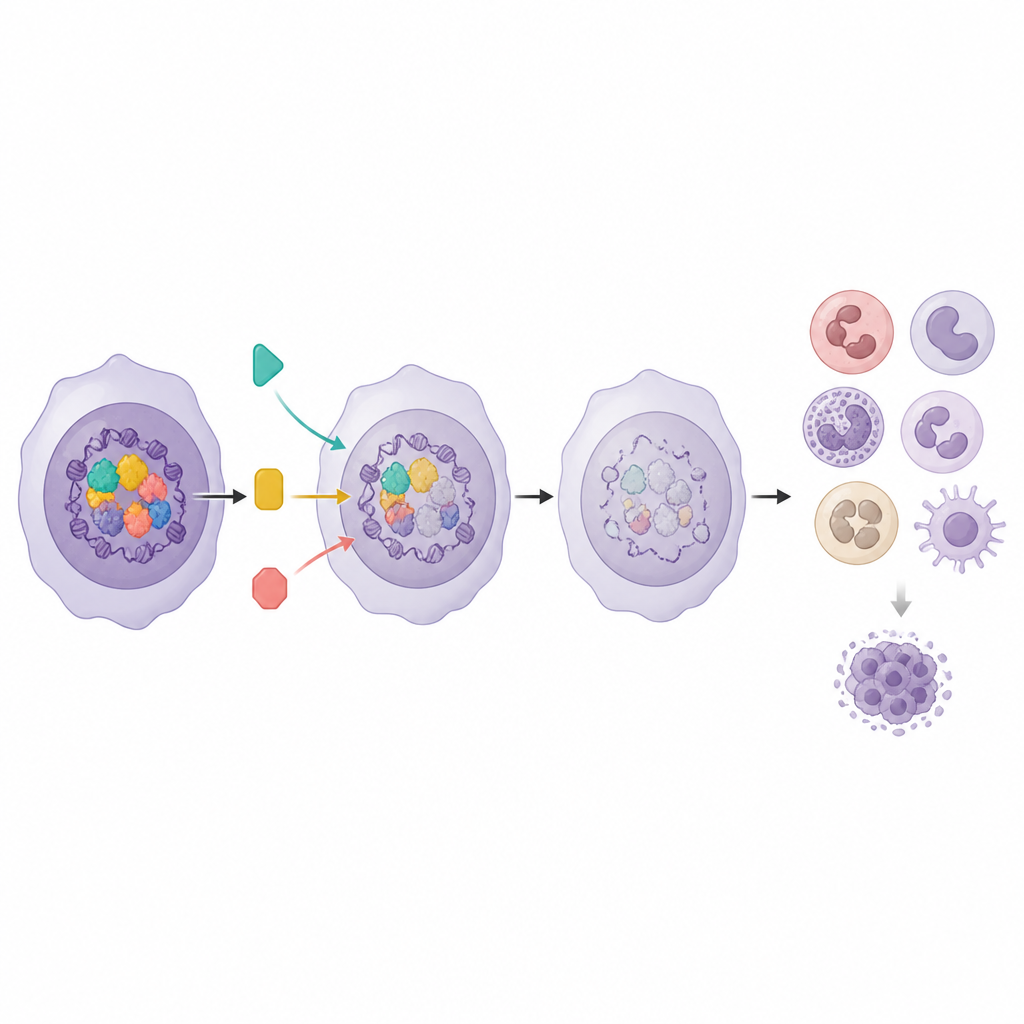
Reading thousands of cells at once
The researchers used a powerful method called Perturb seq, which combines gene editing with single cell RNA sequencing. In a KMT2A rearranged leukemia cell line, they disrupted 16 chosen epigenetic regulators drawn from three major complexes that control which genes are turned on or off. Each edited cell carried a genetic barcode, allowing the team to link a specific edit to that cell’s gene activity profile. By mapping more than 31,000 cells in a reduced two dimensional space, they saw that cells grouped according to which complex was hit, revealing distinct patterns of response. Some factors, especially those tied to the Menin KMT2A fusion and related enzymes, caused sweeping shifts in gene activity, whereas others had subtler effects, hinting at different levels of control within the leukemia network.
A hidden program that favors patient survival
Using computational modeling, the team distilled the complex data into 17 gene programs, each representing a coordinated pattern of gene activity. One program stood out: a set of genes linked to myeloid differentiation, the process by which immature cells become fully functional white blood cells. The authors call this the Myeloid Program. It was most active in normal bone marrow cells that are close to full maturation and showed an inverse link with genes that define leukemia stem cells. When the researchers examined large patient datasets, higher baseline activity of the Myeloid Program was consistently associated with better overall survival, especially in patients whose leukemia tends to respond well to standard treatment. This suggests that even within an aggressive cancer, some cells sit closer to a “ready to mature” state that is linked to better outcomes.
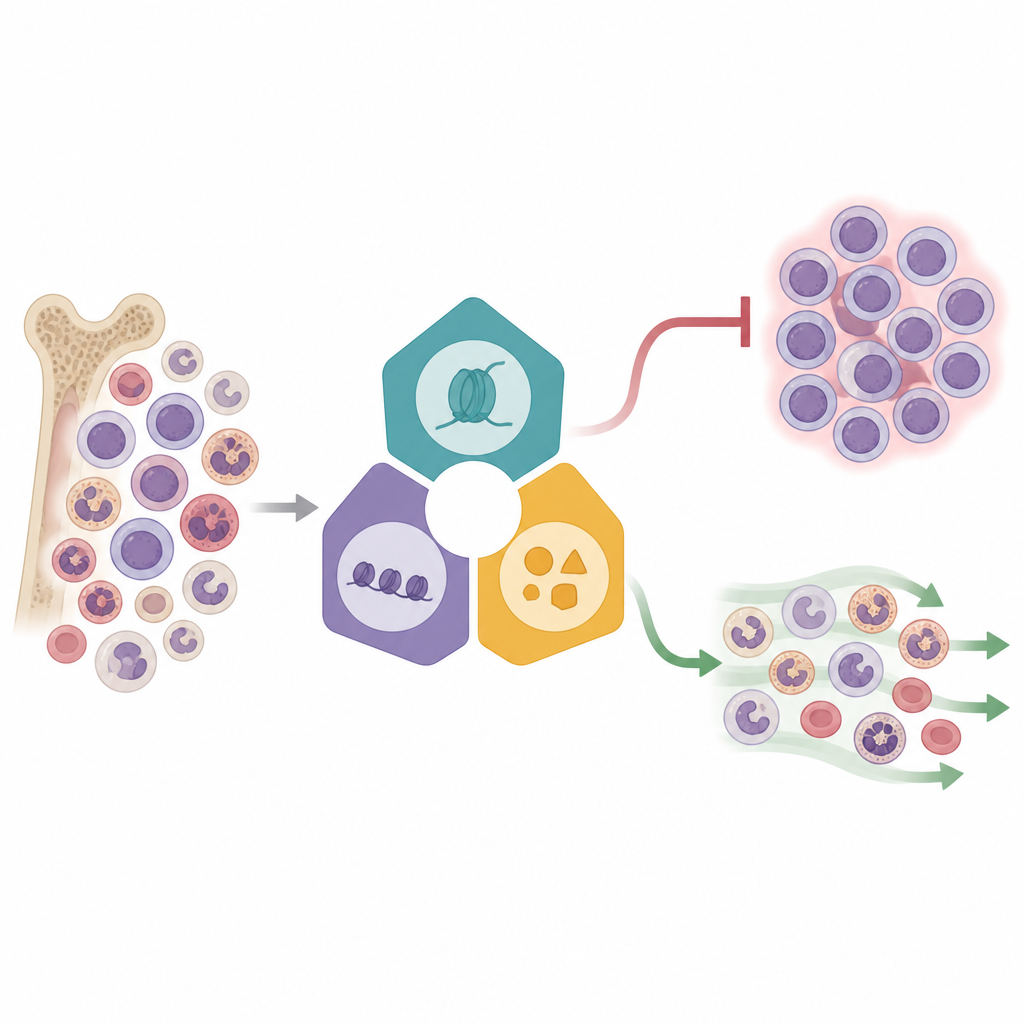
A three part control hub and its weak points
The analysis revealed that three regulators Menin, DOT1L, and KAT6A form a functional hub that jointly keeps the Myeloid Program turned down. When any one of these genes was disrupted, the differentiation program became more active, but only partially. The team then tested drugs that inhibit each partner and discovered striking synergy. Pairing a Menin inhibitor with a KAT6A inhibitor, or a DOT1L inhibitor with a KAT6A inhibitor, sharply reduced leukemia cell growth and strongly boosted expression of Myeloid Program genes compared with single drugs. At the same time, they uncovered an opposing influence from another complex containing PCGF1. Loss of PCGF1 made leukemia cells less sensitive to DOT1L inhibition, highlighting how some epigenetic factors can blunt the effect of targeted drugs and contribute to resistance.
Using gene activity to guide smart combinations
Because the Myeloid Program links to both cell maturation and patient survival, the researchers asked whether its activity could also forecast which drugs would work best. In patient samples, leukemias with higher Myeloid Program scores were more sensitive to several signaling pathway inhibitors, notably those targeting MEK, AKT, and mTOR. When the team combined a Menin inhibitor, which raises the Myeloid Program, with these pathway inhibitors in cell models, the drug pairs showed clear synergy and lowered the doses needed to curb leukemia cell growth. The same gene program also helped distinguish combinations that truly cooperate from those that merely push cells toward a generic differentiated state.
What this means for future leukemia care
For a lay reader, the key message is that this study moves beyond the search for a single “magic bullet” and instead maps how groups of epigenetic regulators jointly trap leukemia cells in an immature, harmful state. By identifying a core Myeloid Program that predicts both better survival and heightened drug sensitivity, and by pinpointing a three part Menin KAT6A DOT1L hub that suppresses this program, the work provides a blueprint for rational combination therapies. In the long term, measuring Myeloid Program activity in patients could help doctors choose drug pairs that both release the maturation block and exploit new weaknesses in the cancer, potentially leading to more durable and personalized treatments for acute myeloid leukemia.
Citation: Aryal, S., Curtiss, B.M., Zhou, X. et al. A Perturb-seq map of a differentiation hub reveals synergistic vulnerabilities in KMT2A-rearranged acute myeloid leukemia. Leukemia 40, 996–1008 (2026). https://doi.org/10.1038/s41375-026-02917-2
Keywords: acute myeloid leukemia, epigenetic therapy, cell differentiation, drug synergy, single cell genomics