Clear Sky Science · en
Biomimetic peptide self-assembly: interfacing with biomacromolecules to regulate cellular signaling
Building Tiny Structures to Talk to Cells
Inside every cell, countless molecules bump, bind, and break apart, creating the signals that keep us alive. This review explores how very short pieces of proteins, called peptides, can be designed to spontaneously organize into tiny shapes that latch onto cell components and nudge those signals in useful directions. By learning to mimic nature’s own strategies, scientists hope to build smart materials that can selectively kill cancer cells, correct faulty signaling, or deliver genetic therapies with far greater precision than today’s drugs.
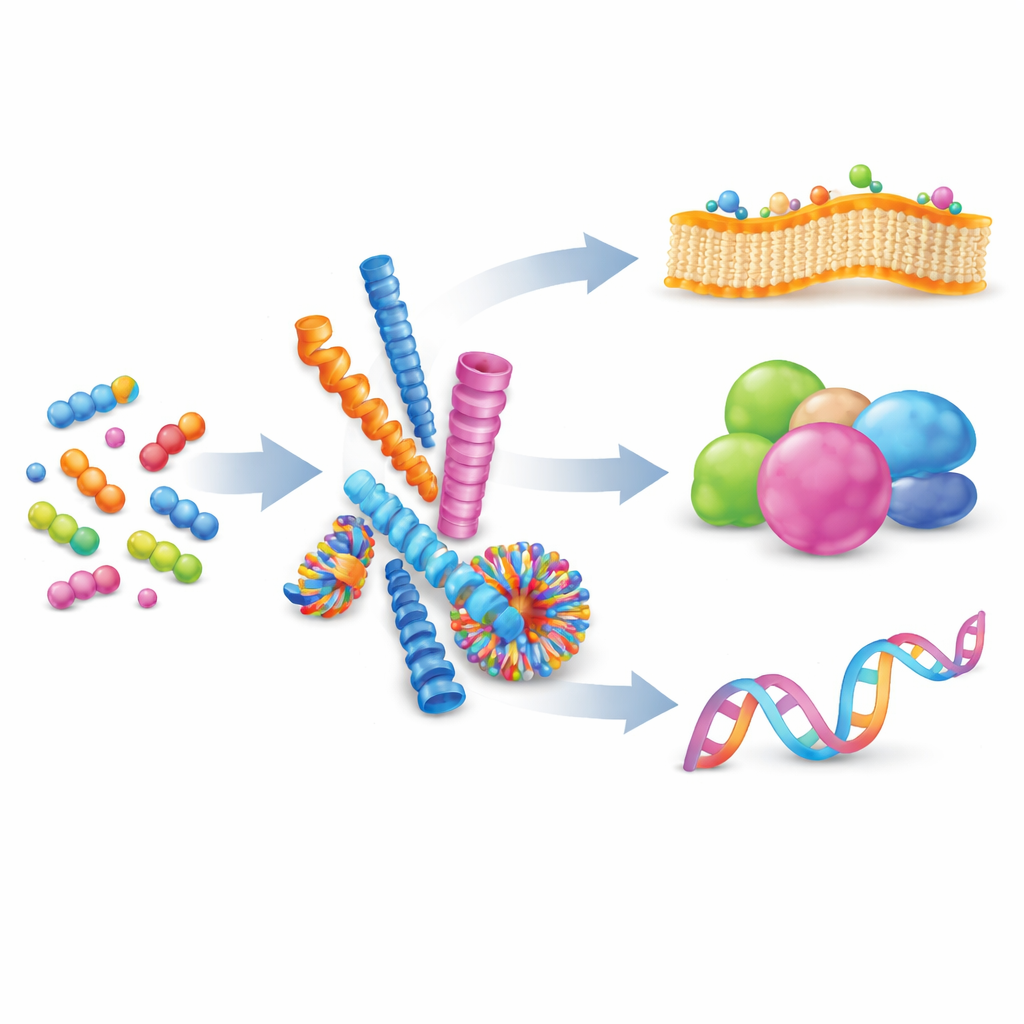
From Loose Molecules to Organized Nanostructures
The authors begin by explaining supramolecular self-assembly: a process where many small molecules gently attract one another and organize into larger structures without forming permanent chemical bonds. By balancing forces such as charge, hydrogen bonding, and stacking between flat rings, chemists can coax peptides to form nanofibers, spheres, tubes, or vesicles. These shapes are not static; their building blocks can exchange in and out, making the assemblies reversible and adaptable. When many weak interactions occur together on a shared surface, they create “multivalency,” a powerful way to strengthen otherwise feeble binding events, much like multiple tiny hooks gripping a surface at once.
Borrowing Nature’s Playbook
Nature already relies on self-assembly to run complex tasks. Proteins fold from simple chains of amino acids into intricate three‑dimensional forms, exposing broad contact areas that bind other proteins, DNA, or membranes. These contacts, often spread over large surfaces, allow cells to switch signaling on and off quickly and reversibly. Inspired by this, researchers now design artificial peptides that imitate such interfaces. Compared with polymers, lipids, or DNA frameworks, peptide systems offer a sweet spot: they are biocompatible, chemically diverse, and precise at the level of individual amino acids, allowing careful tuning of shape, charge, and binding behavior. This review focuses on how these man‑made peptide assemblies are used to engage three main targets: cell membranes, proteins, and nucleic acids.
Rewiring the Cell from the Membrane Inward
A major theme of the article is using peptide self‑assembly to act directly on cellular membranes, especially those of key organelles such as mitochondria, lysosomes, the endoplasmic reticulum, and the Golgi apparatus. Peptides can be decorated with targeting groups that guide them to a specific organelle, where local conditions—such as acidity, electrical potential, or enzyme activity—trigger them to assemble into nanofibers or particles on the membrane surface. In mitochondria, for example, certain designs remain harmless while dispersed in the cell but aggregate into stiff fibers once concentrated by the organelle’s negative voltage. These fibers can puncture or destabilize the double membrane, collapsing the energy gradient, releasing cytochrome c, and activating programmed cell death pathways that are valuable for attacking tumor cells.
Turning Degradative Compartments into Control Hubs
Lysosomes, often thought of simply as cellular recycling centers, are recast here as switchboards for life‑and‑death decisions. Peptides engineered to sense the lysosome’s acidic pH or resident enzymes can assemble only after being swallowed by the cell and trafficked into this compartment. Once assembled, their nanostructures can weaken the lysosomal membrane, allowing digestive enzymes such as cathepsins to leak out and trigger cell death. Other peptide systems are designed to escape from lysosomes at carefully chosen stages, helping carry therapeutic cargo like nucleic acids into the cell’s interior. Similar strategies applied to the endoplasmic reticulum and Golgi apparatus let researchers dial up “stress” responses or disturb protein‑sorting traffic, again steering cells toward self‑destruction in a controlled way.
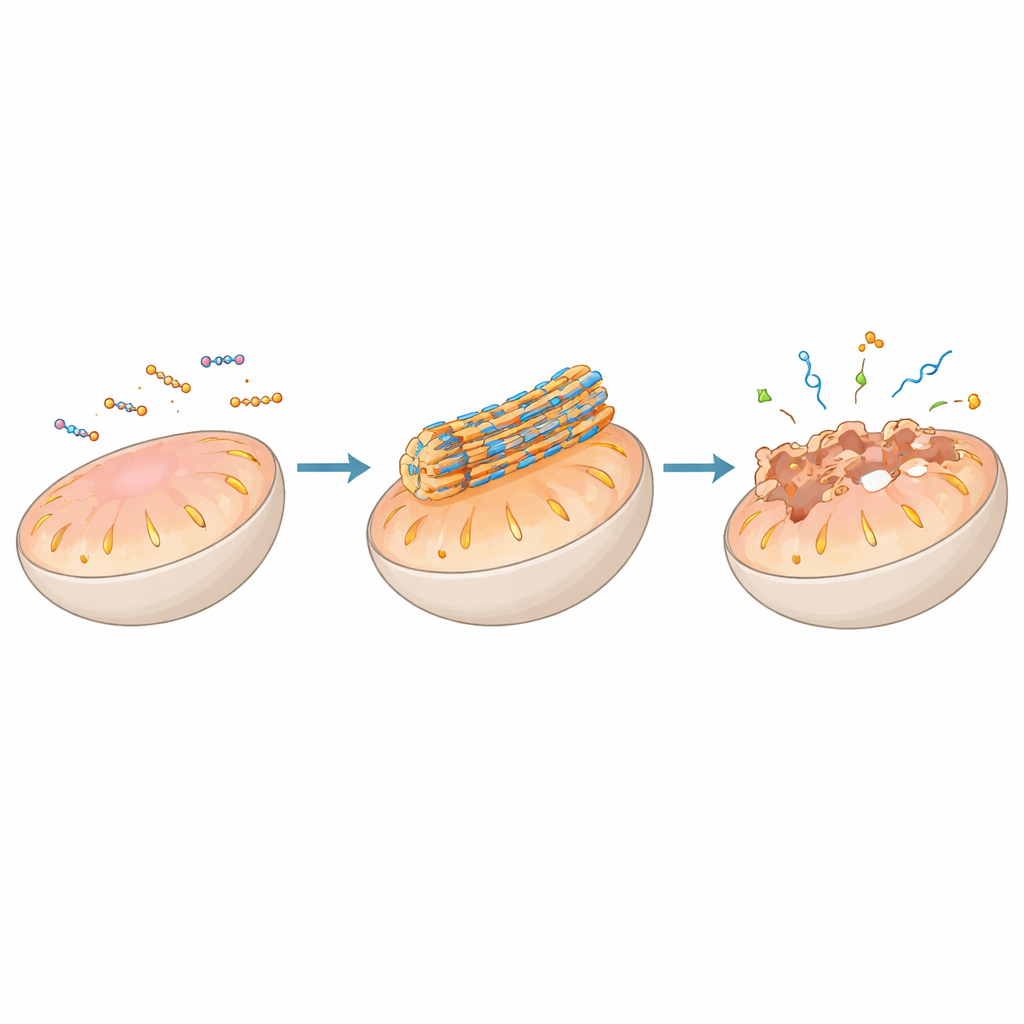
Guiding Proteins and Genes with Peptide Scaffolds
Beyond membranes, peptide assemblies can clamp onto proteins or nucleic acids and reshape entire signaling networks. Some assemblies mimic natural protein‑binding segments so well that they wedge into protein–protein interfaces once considered impossible to drug, freeing pro‑death signals in mitochondria or blocking growth cues on cell surfaces. Others co‑assemble with antibodies or receptors to cluster them, mark them for removal in lysosomes, or boost immune recognition of cancer cells. When combined with DNA or RNA, positively charged peptides condense genetic material into compact particles or liquid‑like droplets that slip into cells and release their cargo at the right time and place, enabling gene silencing or activation. In certain cases, peptide–RNA droplets behave like artificial “condensates,” echoing stress granules that cells naturally form to manage RNA during crises.
Why These Tiny Builders Matter
The review concludes that biomimetic peptide self‑assembly offers a flexible toolkit for reshaping cellular behavior with molecular‑level precision. Because these systems respond to local triggers and can reorganize themselves, they can, in principle, distinguish diseased cells from healthy ones and act only where needed—by punching holes in energy factories, disturbing recycling centers, silencing harmful genes, or reigniting immune attack. The authors argue that as imaging, modeling, and design methods improve, these tiny builders could evolve from lab curiosities into a new generation of adaptive medicines that speak the same structural language as the cell itself.
Citation: Kim, D., Park, G., Seu, MS. et al. Biomimetic peptide self-assembly: interfacing with biomacromolecules to regulate cellular signaling. Exp Mol Med 58, 1038–1052 (2026). https://doi.org/10.1038/s12276-026-01691-6
Keywords: peptide self-assembly, cell signaling, nanomedicine, organelle-targeted therapy, supramolecular biomaterials