Clear Sky Science · en
Deep learning-driven integrated pipeline for de novo design and synthesis of antimicrobial peptides
Why new germ-fighting molecules matter
As common antibiotics lose their punch against ever-tougher bacteria, doctors are running out of options to treat everyday infections. One promising alternative is antimicrobial peptides—short protein fragments that punch holes in bacterial membranes instead of targeting a single enzyme. This article describes how researchers used artificial intelligence (AI) to automatically design and test new antimicrobial peptides in a faster, cheaper, and more targeted way than traditional trial‑and‑error experiments.

A new way to search the vast space of molecules
Designing useful antimicrobial peptides is like searching for a few winning lottery tickets in a mountain of random numbers: there are astronomically many possible sequences of amino acids, but only a tiny fraction will be safe and effective drugs. Conventional lab screening is slow and expensive, and even computer models often struggle to balance potency with the diversity needed to uncover genuinely new molecules. The authors address this by building a three‑step pipeline—generation, evaluation, and experimental validation—that lets AI explore huge regions of sequence space while still homing in on candidates that look realistic and promising.
Teaching a protein language model to think like a peptide designer
At the heart of the first step is ProtGPT2, a large “protein language model” originally trained to write plausible protein sequences the way text models write sentences. Instead of fully retraining this massive model—an approach that is costly and prone to overfitting to small antimicrobial datasets—the team uses a strategy called soft prompt tuning. They attach a small, trainable input prompt in front of each sequence and update only this prompt while keeping the model’s internal weights fixed. This gentle steering proved enough to shift the model toward making antimicrobial‑like peptides without sacrificing its broad creativity. Analyses of the generated sequences showed that they matched real antimicrobial peptides in key properties such as charge, hydrophobicity, and structural patterns, while remaining more diverse than sequences produced by full retraining.
Letting multiple AI “experts” vote on each candidate
In the second step, the researchers face the opposite challenge: from 30,000 generated sequences, which ones merit the cost of chemical synthesis and lab testing? They build an evaluation model called MCL‑AMP that combines three different neural network “experts”—one that focuses on local patterns, one that tracks long‑range relationships along the sequence, and one that integrates overall features. A separate voting network learns how much to trust each expert for each peptide, and a special loss function discourages the system from being overconfident when it is wrong. Using rich sequence features extracted by another advanced protein model (ESM‑2), this ensemble achieved higher accuracy than existing tools at distinguishing antimicrobial from non‑antimicrobial sequences, and it produced an interpretable confidence score for each candidate.
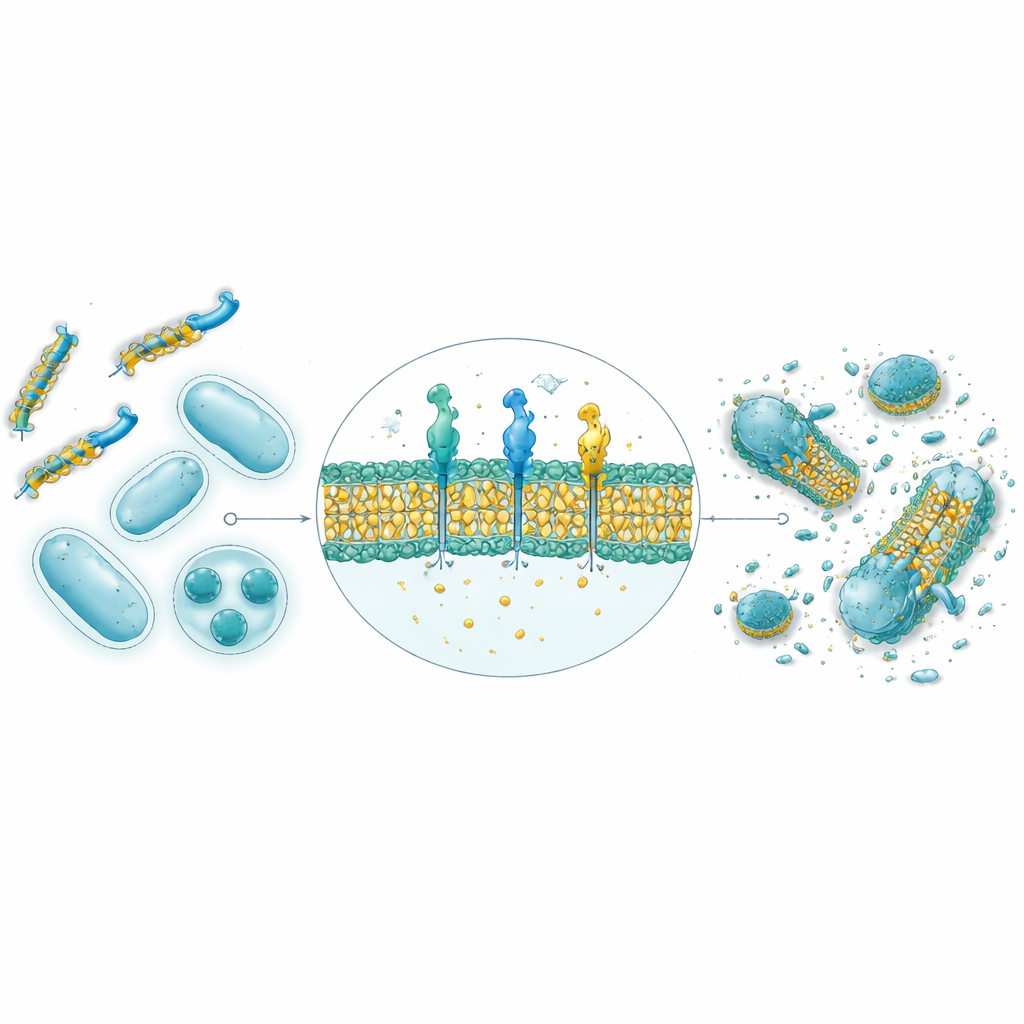
From computer predictions to real bacteria in the lab
From the 30,000 AI‑generated sequences, the nine top‑scoring peptides were synthesized and tested against four bacterial strains, including both Gram‑positive and Gram‑negative species. Structural predictions using AlphaFold suggested that eight of the nine adopt the amphipathic helical shapes typical of natural antimicrobial peptides, with separate water‑loving and fat‑loving faces that favor membrane attack. In petri‑dish tests, four of the nine showed strong antibacterial activity, and two in particular—named GW20 and KI16—approached the performance of established antibiotics in stopping bacterial growth. Microscopy revealed that these peptides visibly damaged bacterial cell surfaces, consistent with a mechanism of membrane disruption. Importantly, both showed minimal damage to red blood cells at therapeutically relevant concentrations and demonstrated powerful ability to prevent and break down sticky bacterial biofilms, often outperforming a standard antibiotic.
What this means for future medicines
For non‑specialists, the key message is that the authors have built a working “design–test” loop in which AI proposes many new germ‑killing peptides, triages them using a committee of models, and then confirms hits in the lab. Even though not every top‑ranked sequence worked, the hit rate—four active peptides out of nine tested—is far higher than traditional blind screening. The two most promising molecules already display a useful mix of potency, safety, and biofilm‑fighting ability, hinting at future therapies for stubborn infections. Just as important, the same modular pipeline could be retuned to design other medical peptides, such as cancer‑targeting agents or antiviral molecules, offering a flexible blueprint for AI‑accelerated drug discovery in the face of rising antimicrobial resistance.
Citation: Liu, J., Chen, Y., Tang, J. et al. Deep learning-driven integrated pipeline for de novo design and synthesis of antimicrobial peptides. npj Drug Discov. 3, 15 (2026). https://doi.org/10.1038/s44386-026-00045-6
Keywords: antimicrobial peptides, artificial intelligence, drug discovery, antibiotic resistance, biofilms