Clear Sky Science · en
A critical assessment of aptamer and CRISPR-Cas12a-based biosensors for small molecule detection
Why tiny pollutants and quick tests matter
From drinking water and rivers to the food on our plates and the medicines we take, countless small molecules move through our environment and bodies. Some are life-saving drugs; others are toxic contaminants. Measuring these compounds usually requires bulky and expensive instruments, which are hard to bring into the field or a small clinic. This paper examines a promising alternative: compact biochemical tests that use designer DNA strands and CRISPR enzymes to turn the presence of tiny molecules into a bright light signal. The authors set out to see how well this much-hyped approach actually works in practice.
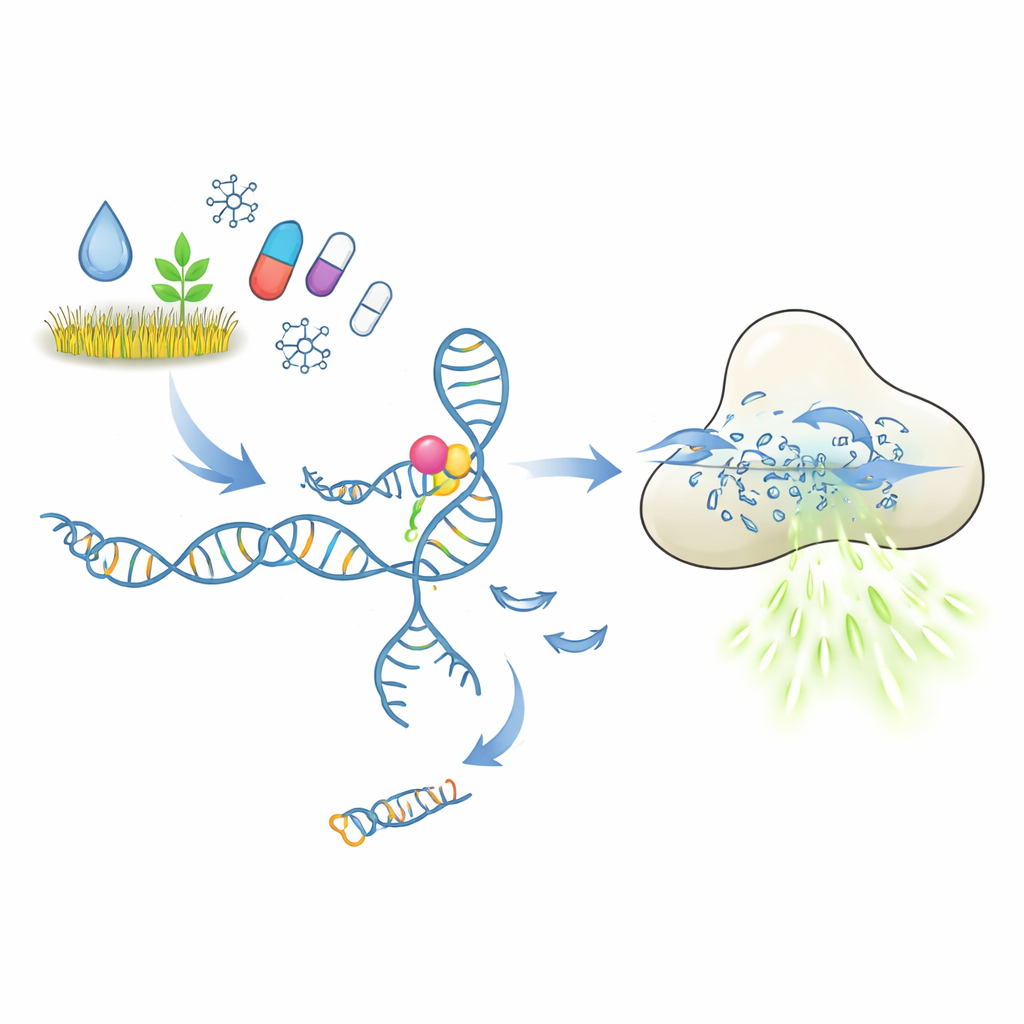
How a molecular lock-and-key test is supposed to work
At the heart of these tests are aptamers, short pieces of DNA or RNA that fold into shapes able to latch onto a chosen target, much like antibodies do. In the setup explored here, an aptamer holds onto a short DNA piece that acts as a trigger. When the target pollutant or drug binds, the aptamer is expected to change shape and release this trigger. The freed DNA then activates the CRISPR-Cas12a enzyme, which behaves like a molecular paper shredder for single-stranded DNA. By providing Cas12a with many short DNA reporters carrying a fluorescent dye, every cut produces more light. In principle, a tiny amount of target molecule should thus be converted into a strong, easy-to-measure fluorescence signal.
Putting the CRISPR-based sensor through its paces
The researchers approached the problem like careful mechanics testing a new engine. First, they tuned the Cas12a reaction itself: choosing the best buffer, metal ions, enzyme amount, and reporter design to get strong and reliable light signals when trigger DNA was present. Under optimized conditions, Cas12a could detect extremely low levels of trigger DNA, down to trillionths of a mole, and did so reproducibly. This confirmed that the signaling machinery—the "amplifier" part of the test—was robust and sensitive on its own.
Where the design starts to fail
The real challenge emerged when they connected this sensitive amplifier to real aptamers from the scientific literature. The team selected nine aptamers that were reported to recognize seven different small molecules, including pesticides, antibiotics, a liver toxin from algae, and antimalarial drugs. In theory, these aptamers should release the trigger DNA when they meet their targets. The authors carefully adjusted how strongly the trigger DNA stuck to each aptamer, tested different temperatures, incubation times, and buffer conditions, and even tried a more complex version where the aptamer complexes were attached to magnetic beads and separated from the reaction afterward. Despite this extensive effort, clear signals appeared for only two of the seven target molecules. For the others, adding the target did not lead to extra trigger DNA in solution, suggesting that either the aptamers did not bind as claimed, or binding did not cause the expected shape change and release.
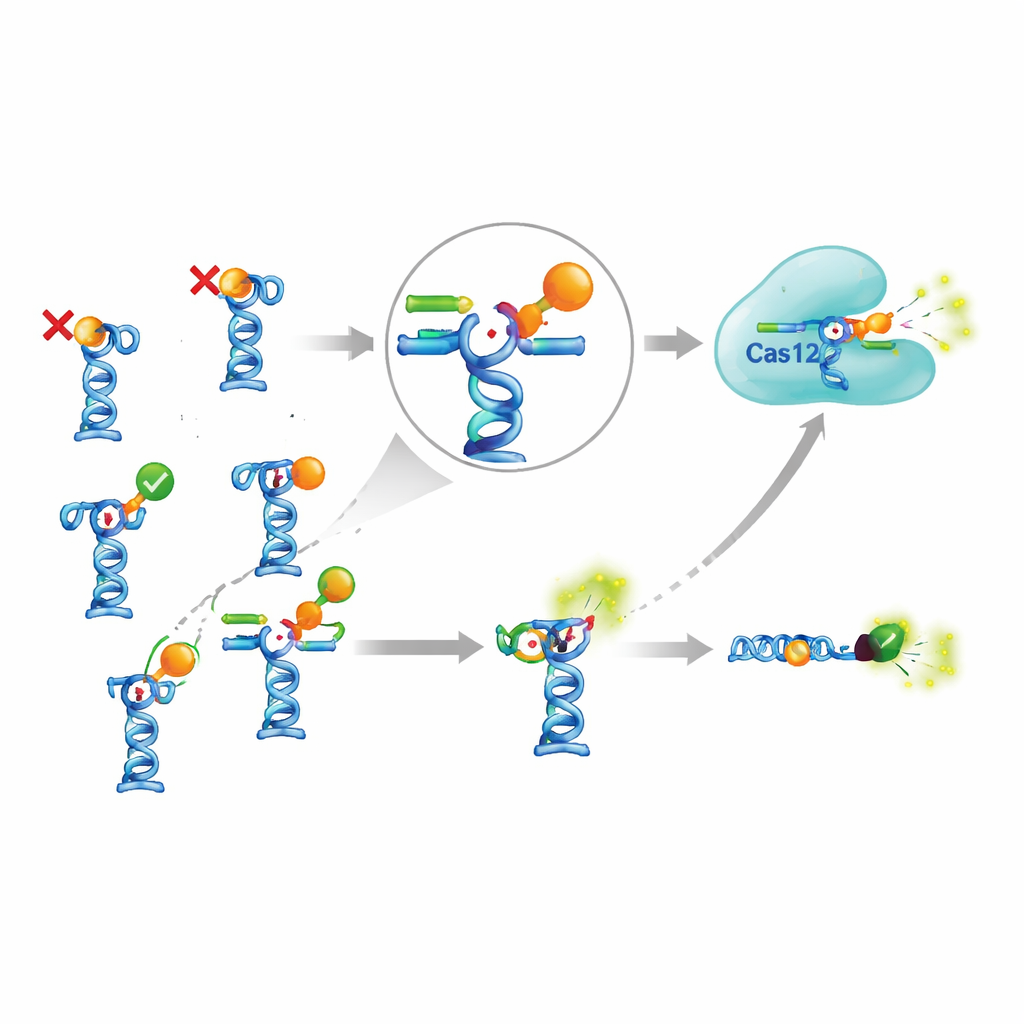
Is the complicated setup worth the trouble?
To judge whether the CRISPR add-on truly pays off, the authors compared it with a simpler aptamer test that skips Cas12a altogether. In this simpler design, the aptamer carries a fluorescent dye and its partner strand carries a light-quenching group; when the partner strand is displaced by the target molecule, the fluorescence increases directly. For one of the few working targets, the antimalarial piperaquine, the CRISPR-based version detected slightly lower concentrations than the simple test, but only by a factor of two to three. Achieving this modest gain required extra enzymes, more reagents, longer protocols, and more optimization. At the same time, some previously published aptamer-based sensors—for example, for a common algal toxin and the insecticide fipronil—could not be reproduced at all, even when the authors followed reported conditions closely and used an independent method to test binding.
What this means for future quick tests
The study concludes that while the Cas12a fluorescence system itself is powerful, the overall aptamer–CRISPR sensor strategy is fragile. Its success hinges on aptamers that not only bind their targets but also reliably let go of the trigger DNA—a demanding requirement that many published aptamers do not actually meet. For now, researchers and application-focused labs should treat plug-and-play claims with caution, invest in thorough validation of aptamer performance, and weigh whether simpler aptamer tests or established methods such as mass spectrometry might serve their needs better. CRISPR-based aptasensors remain an intriguing tool, but not yet the universal solution for small molecule detection they have sometimes been advertised to be.
Citation: Brandenberg, O.F., Janssen, E.ML. & Schubert, O.T. A critical assessment of aptamer and CRISPR-Cas12a-based biosensors for small molecule detection. npj Biosensing 3, 28 (2026). https://doi.org/10.1038/s44328-026-00089-8
Keywords: aptamer biosensors, CRISPR-Cas12a detection, small molecule sensing, environmental contaminants, fluorescence assays