Clear Sky Science · en
Recombinant protein platform for high-throughput investigation of peptide-liposome interactions via fluorescence anisotropy depolarization
How Shape-Shifting Protein Pieces Find the Right Membranes
Inside every cell, countless deliveries, signals, and recycling events depend on proteins landing on the right membrane at the right time. Many of these proteins use short, floppy stretches that only fold into tidy helices when they touch a suitable membrane. This paper introduces a new, faster way to study how those shape-shifting pieces recognize specific membranes, using a surprising optical signal as a built-in reporter.
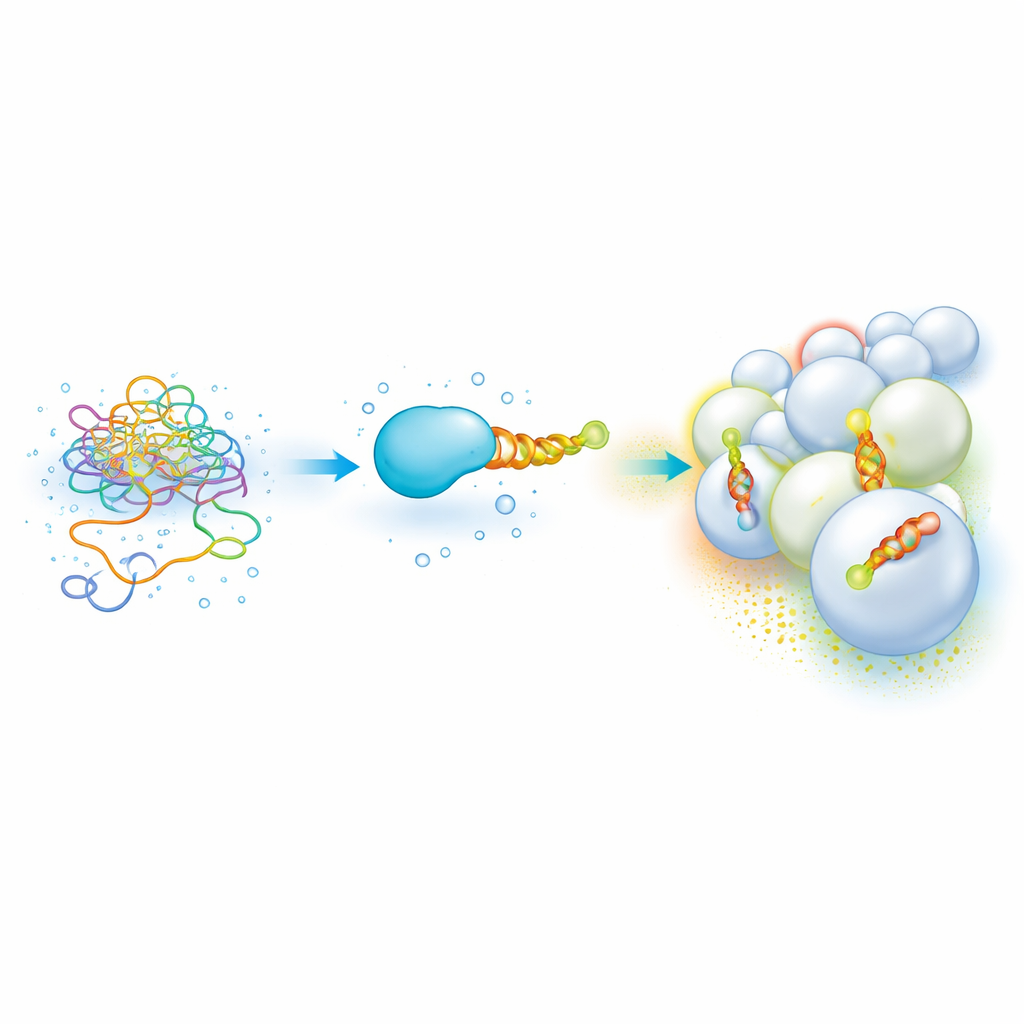
Flexible Protein Ends That Read Membrane "Zip Codes"
Many important cellular proteins carry a short, unstructured segment that becomes an amphipathic helix—a spiral with one oily side and one watery side—once it docks onto a membrane. These helices help proteins tell one organelle from another by sensing lipid makeup and curvature, allowing them to home in on places like endosomes or mitochondria. But taken on their own, these sticky, partly oily pieces are hard to work with: they clump, precipitate, and require slow, sample-hungry methods to measure how tightly they bind to different membranes. As a result, building a clear “code” that links helix sequence to membrane preference has been difficult.
A Modular Protein Platform That Lights Up Binding
The authors built a recombinant platform that tames these unruly helices and makes their membrane choices easy to measure in parallel. Each test helix is fused to a well-behaved carrier protein called SUMO, which boosts solubility and expression, and is tagged at the far end with a fluorescent dye. When the construct is mixed with artificial lipid vesicles and read out in a common microplate reader, the researchers track fluorescence anisotropy—a measure of how much the dye’s light emission “remembers” its polarization, which depends on how freely the dye can rotate. By titrating many lipid mixtures and vesicle sizes in small volumes, they quickly build binding curves for different helices across a wide landscape of membrane compositions and curvatures.
A Counterintuitive Signal That Still Reports Binding
Intuitively, one would expect that when a small protein-dye construct binds a much larger vesicle, its rotation slows and anisotropy goes up. Instead, the authors observe the opposite: anisotropy decreases as binding increases. They tested whether this odd behavior still tracks real membrane association by comparing it to a classic method based on tryptophan fluorescence, which reports how deeply a helix buries itself in the oily core of the membrane. For several biologically relevant helices—from proteins such as Amphiphysin, CHMP4B, Endophilin-B1, and Huntingtin—the new platform and the traditional readout give closely matching binding curves and lipid preferences, even revealing new trends such as stronger Amphiphysin helix binding to cholesterol-rich membranes.
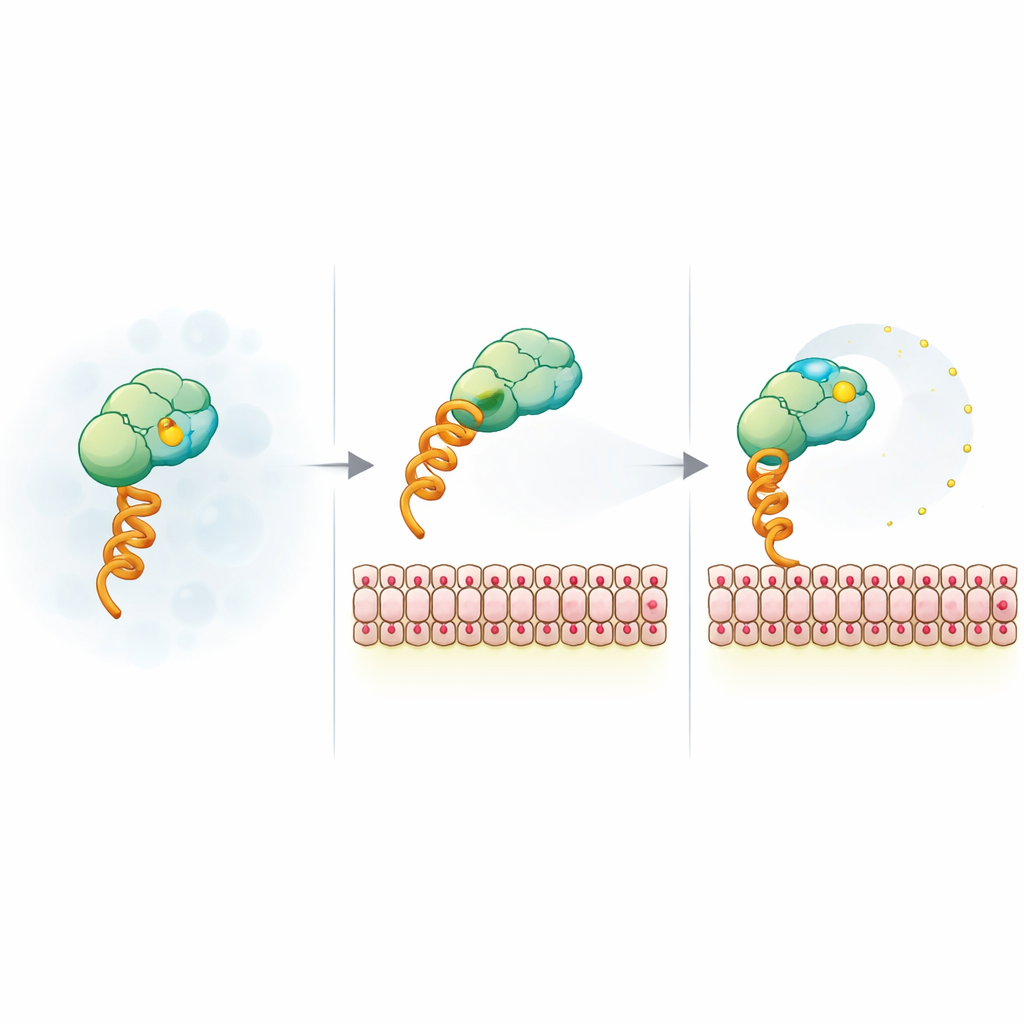
How Membrane Charge Frees the Fluorescent Reporter
To explain the unexpected signal, the team stripped the system down to a model construct that binds membranes via a generic histidine tag and a nickel-chelating lipid. They then varied vesicle size, surface charge, salt level, and dye type. The anisotropy drop turned out to be insensitive to vesicle size and fluorophore identity but highly sensitive to negative membrane charge and screening by salt, pointing to an electrostatic origin. Time-resolved fluorescence measurements showed that, when bound to charged membranes, the dye gains extra local mobility even though the whole complex rotates more slowly. Nuclear magnetic resonance and FRET experiments further suggested that positive patches on the SUMO surface can act as temporary tethers for the negatively charged dye; binding to a charged membrane and mutating these patches both loosen this tether, letting the dye wobble more freely and lowering anisotropy.
Mapping Membrane Preferences at Scale
Because the anisotropy decrease is robust and reproducible, it can be used as a convenient reporter of binding rather than treated as a problematic artifact. Arrays of vesicles differing in lipid type and curvature can be probed quickly, allowing researchers to pinpoint which combinations of lipids and shapes best attract a given helix. The platform works especially well for negatively charged membranes, which are common in organelles involved in trafficking and energy production. By engineering charge-altered variants of the carrier protein, the authors extend the useful range of conditions toward more physiological salt levels and subtly charged membranes, further broadening the method’s reach.
What This Means for Understanding Cellular Targeting
In everyday terms, this work provides a new, high-throughput “reader” for the molecular zip codes that guide proteins to the correct cellular address. Instead of relying on bulky, slow experiments, scientists can now screen many helices and membrane recipes in parallel, using a simple optical signature that flips in an unexpected but well-understood direction. Over time, this should make it possible to decode which amino acid patterns correspond to which lipid environments, clarifying how cells use these small, flexible helices to sort proteins, reshape membranes, and control vital signaling and trafficking pathways.
Citation: Margaritakis, A., Qian, M., Johnson, D.H. et al. Recombinant protein platform for high-throughput investigation of peptide-liposome interactions via fluorescence anisotropy depolarization. Commun Chem 9, 165 (2026). https://doi.org/10.1038/s42004-026-01994-9
Keywords: membrane-binding peptides, fluorescence anisotropy, amphipathic helices, lipid vesicles, protein–membrane interactions