Clear Sky Science · en
Cryo-EM structure of bixafen-bound S. cerevisiae complex II unravels SDHI specificity against pathogenic fungi
Why blocking fungus power plants matters
Fungal diseases threaten harvests, forests and even people, and many modern fungicides work by cutting off the pathogen’s energy supply. Yet, until now, no one had seen in atomic detail how these chemicals latch onto their fungal targets. This paper reveals the three‑dimensional structure of a crucial energy‑producing enzyme from baker’s yeast and shows exactly how a widely used fungicide, bixafen, jams this molecular machine. Because this enzyme is closely related in many pathogenic fungi, the work opens the door to designing smarter fungicides that are potent against crop and medical threats but safer for humans and the environment.
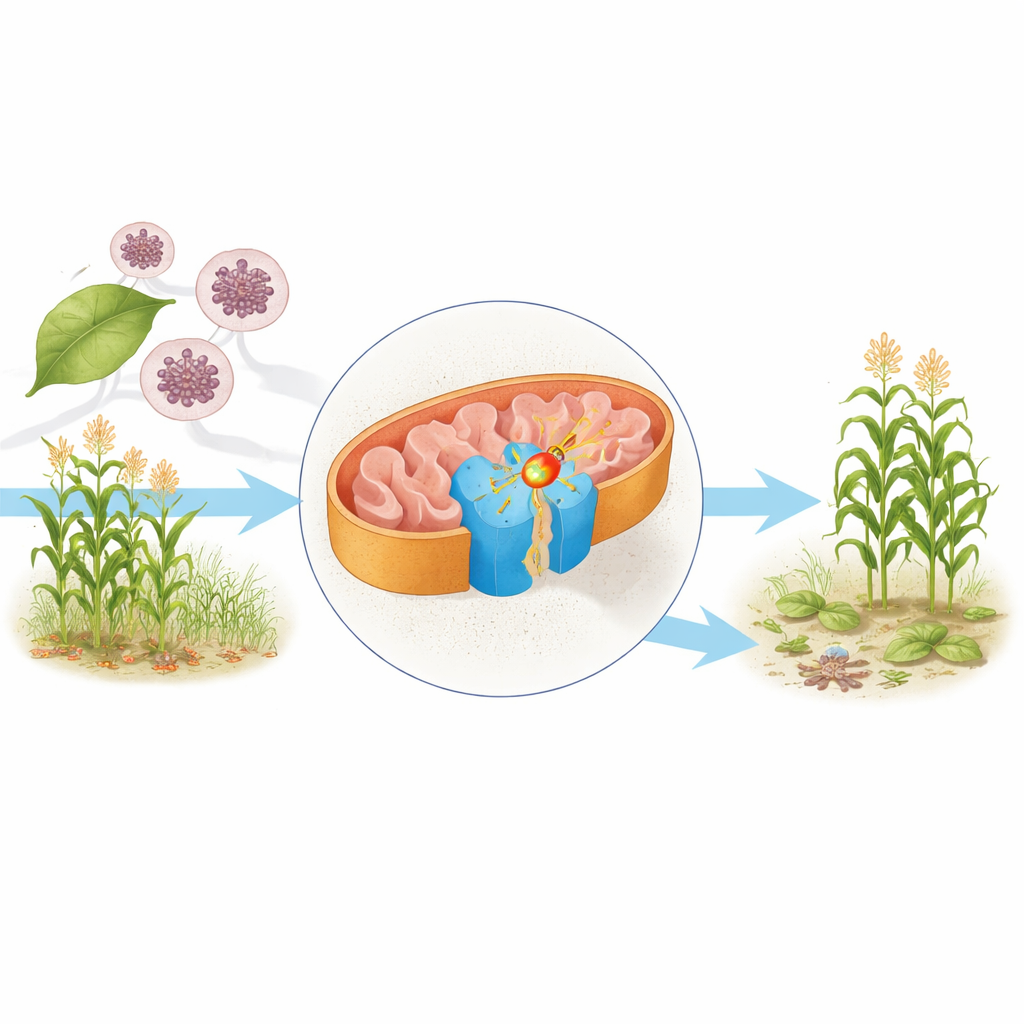
The cell engine under the microscope
Inside the mitochondria—often called the cell’s power plants—food molecules are stripped of electrons that ultimately help make ATP, the cell’s main fuel. One central component of this machinery is complex II, also known as succinate dehydrogenase. It links the Krebs cycle, which breaks down nutrients, to the respiratory chain, which converts that chemical breakdown into usable energy. The authors purified complex II from the yeast Saccharomyces cerevisiae, a workhorse organism in biology, and used cryo‑electron microscopy to determine its structure at near‑atomic resolution in two forms: one carrying its natural electron‑shuttling molecule, ubiquinone, and another bound to the fungicide bixafen.
A familiar engine with a surprising missing part
The yeast enzyme closely resembles complex II from mammals and other organisms, with two large water‑loving subunits handling chemistry and two smaller, membrane‑embedded subunits anchoring the complex in the mitochondrial membrane. Electrons travel from a flavin cofactor through a chain of iron–sulfur clusters toward the membrane, where ubiquinone normally picks them up. However, the structure reveals a major surprise: unlike typical “type C” complex II enzymes, the yeast version completely lacks a heme group—an iron‑containing cofactor previously thought to be a universal, stabilizing feature of this family. Instead, yeast complex II relies on extra hydrogen bonds, more hydrophobic residues, and a deeply buried phospholipid to glue its membrane subunits together and keep the complex intact without the heme.
Fungicide in the docking bay
At the heart of the study is the so‑called Q site, the pocket where ubiquinone normally docks to receive electrons. In the bixafen‑bound structure, this pocket is filled by the fungicide, which mimics many of ubiquinone’s interactions but also makes additional snug contacts. Key amino acids from three subunits cradle the inhibitor through a mix of polar and hydrophobic interactions, while a flexible extension from one membrane subunit (Sdh4) folds into the site and wraps around bixafen’s hydrophobic tail. When the natural substrate is present instead, the very tip of this Sdh4 extension becomes disordered and makes room for ubiquinone’s longer chain, showing that the pocket can adapt in different ways to substrate and inhibitor.
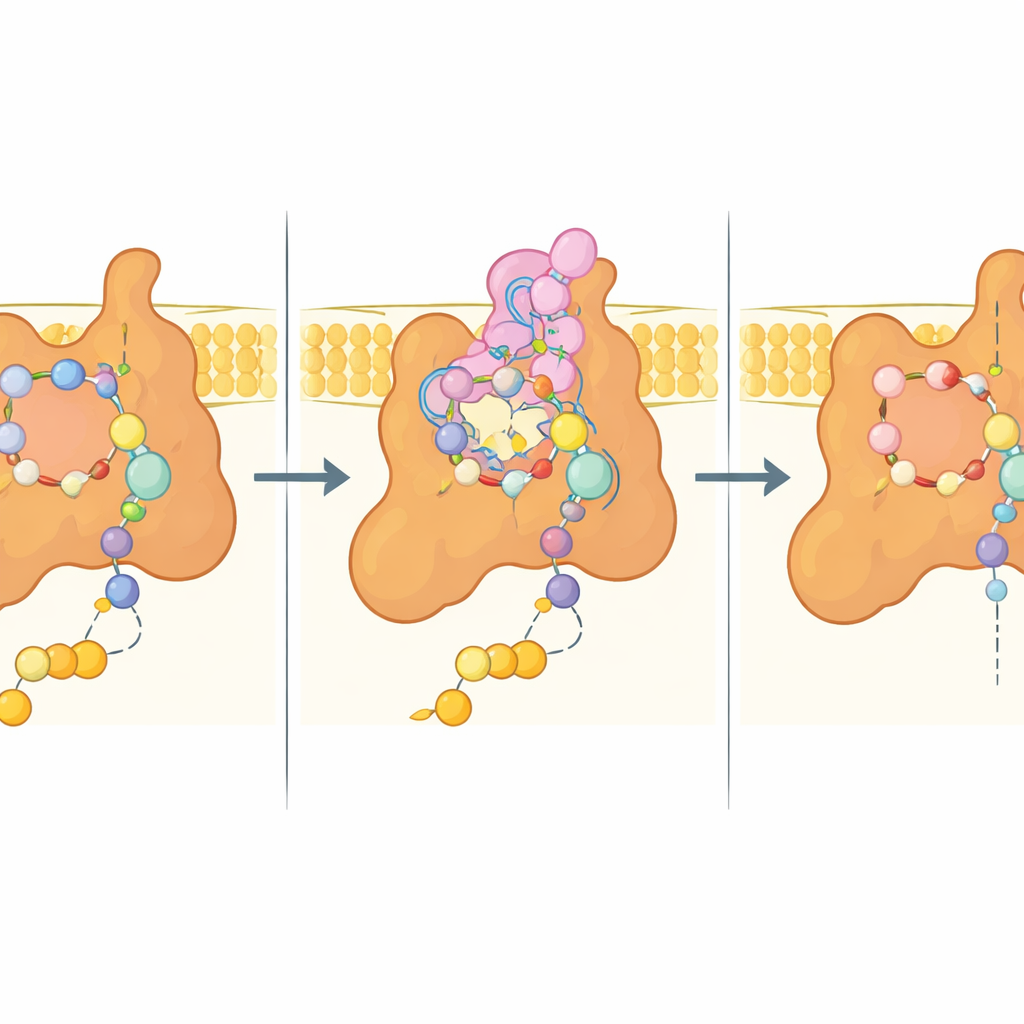
Clues for next‑generation fungicides
Because many agriculturally important and human‑pathogenic fungi share closely related Sdh4 sequences, especially in this N‑terminal extension, the yeast structure likely mirrors their complex II architecture. The authors used their experimental model to computationally dock several commercial succinate dehydrogenase inhibitor (SDHI) fungicides, revealing a common binding logic: a carboxamide “core” that engages conserved amino acids, plus variable hydrophobic decorations that fit into nearby pockets and modulate strength and specificity. Importantly, nearly all residues that contact bixafen in yeast are conserved in multiple crop pathogens, suggesting that subtle sequence differences could explain why some species are more or less sensitive—and why certain mutations confer resistance in the field.
From structural map to practical impact
In plain terms, the study provides a detailed blueprint of how a major class of fungicides wedges into a fungal energy enzyme and shuts it down. It also overturns assumptions about a supposedly universal heme component and shows how yeast has evolved alternative stabilizing features. With this structural map, researchers can now rationally tweak existing fungicides or invent new ones that better exploit fungal‑specific features, reduce the chances of resistance, and minimize off‑target damage to crops, wildlife and people. Baker’s yeast, long beloved for bread and beer, thus becomes a powerful model for building the next generation of targeted antifungal weapons.
Citation: Pinotsis, N., Burn-Leefe, C., Jones, S. et al. Cryo-EM structure of bixafen-bound S. cerevisiae complex II unravels SDHI specificity against pathogenic fungi. Commun Biol 9, 517 (2026). https://doi.org/10.1038/s42003-026-09617-8
Keywords: succinate dehydrogenase, fungicide resistance, cryo electron microscopy, mitochondrial respiration, SDHI fungicides