Clear Sky Science · en
Calcium dependent activation of the TMEM16F scramblase and ion channel
Why this tiny membrane valve matters
Every second, the membranes around our cells quietly shuffle molecules to keep us alive. One unsung player in this balancing act is a protein called TMEM16F. It helps blood clot, repairs torn membranes, guides brain immune cells, and even gets hijacked by viruses like SARS‑CoV‑2 to infect cells. Yet, until now, scientists did not understand how this protein turns on and off, or how it can act both as an ion channel and as a lipid "scrambler" that flips fat molecules between the two sides of the membrane. This study uses high‑resolution imaging and computer simulations to reveal TMEM16F’s working motions in unprecedented detail.
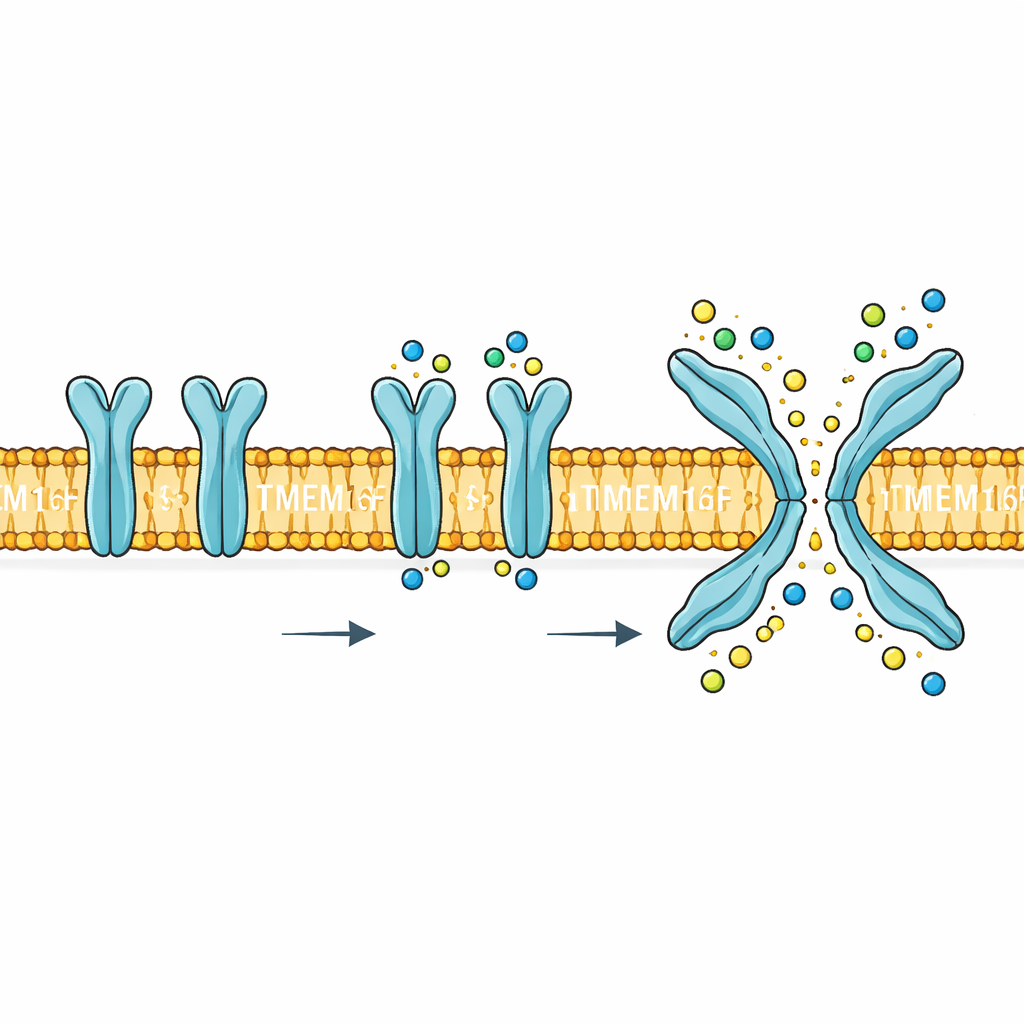
A dual‑purpose gate in the cell boundary
TMEM16F sits in the cell’s outer membrane and responds to rises in calcium inside the cell. When activated, it performs two very different jobs. First, it forms a pathway that lets small charged particles such as sodium, potassium and chloride move across the membrane. Second, it flips certain lipids, especially phosphatidylserine, from the inner face of the membrane to the outer face. This lipid flip is a powerful biological signal: it helps platelets form clots, marks damaged cells for clearance and can be exploited by viruses to enter cells. Mutations or misregulation of TMEM16F are linked to blood, bone and nervous system problems, making its mechanism more than a curiosity—it is a potential drug target.
Seeing the protein in its native habitat
Previous structural studies examined TMEM16F in artificial environments such as detergents or nanodiscs, and consistently saw it in what looked like a closed state, even when calcium was present. This raised a puzzle: functional experiments clearly showed that the protein was active, but its structure did not seem to open a route for ions or lipids. To resolve this, the authors reconstituted mouse TMEM16F into tiny, free‑floating membrane bubbles (liposomes) made of simple, well‑defined lipids—the same type of environment in which they could directly measure ion movement and lipid scrambling. They then used cryo‑electron microscopy to capture millions of snapshots of the protein, and sorted these into distinct shapes representing different activity states.
How calcium reshapes the gate
In the absence of calcium, TMEM16F adopted two similar, inactive shapes in which a key groove through the protein was tightly sealed. When calcium was added, the protein favored a conventional calcium‑bound but still closed form, and a second, rarer but strikingly different form. In this active state, a segment of the protein called TM4 slid against a neighboring segment TM6, creating an X‑shaped groove that crossed the membrane. The central part of this groove formed a continuous, protein‑walled pore wide enough for partially hydrated ions, while the regions closer to the membrane surfaces opened into broader vestibules. The researchers confirmed the importance of this conformation by engineering mutations: those that shifted the protein toward the X‑shaped state made scrambling easier, while those that trapped it in the closed form nearly abolished activity.
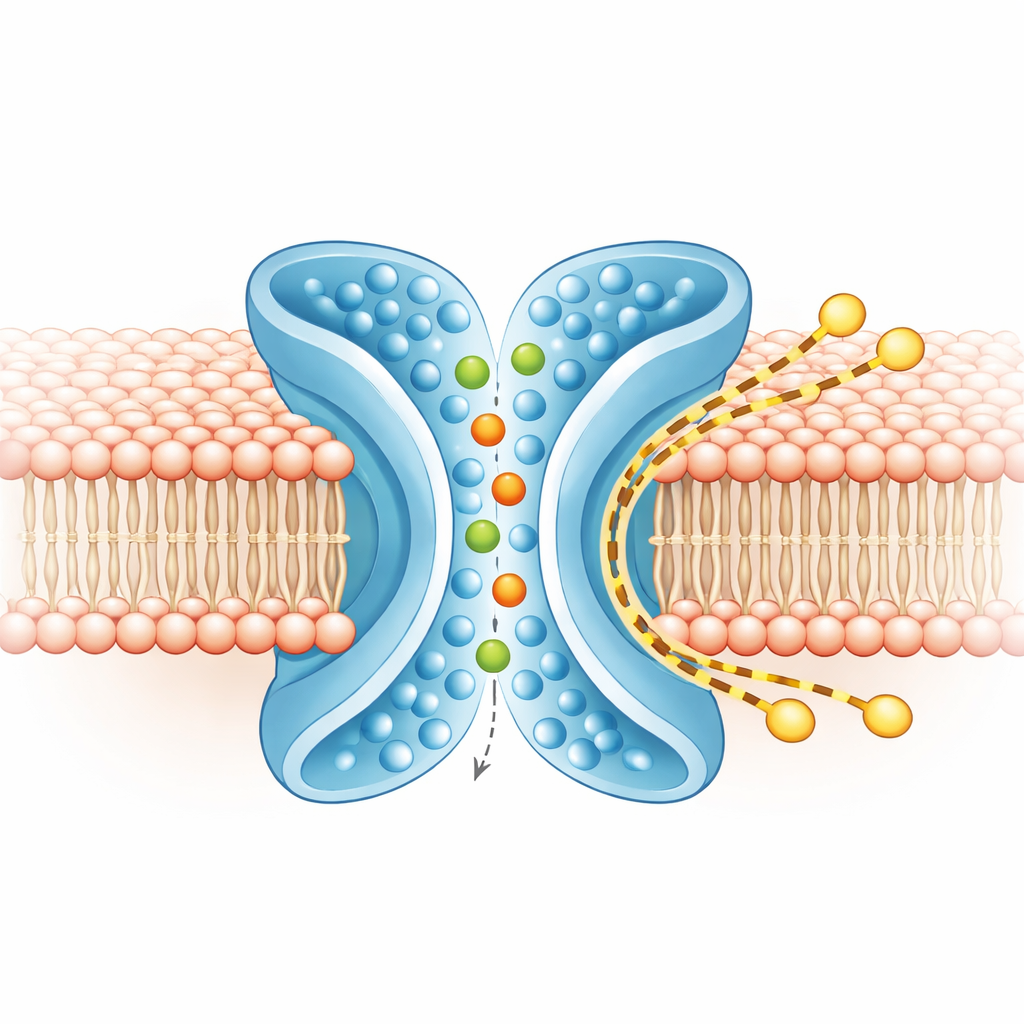
Separate tracks for ions and lipids
To understand what moves where, the team ran extensive computer simulations of TMEM16F in a realistic membrane, both in the closed and active shapes. In the active, X‑shaped form, ions snaked through the central pore, entering from either side and keeping most of their surrounding water shell, consistent with the protein’s loose preference between positive and negative ions. At the same time, the outer layer of the membrane around TM4 and TM6 thinned and bent. Coarse‑grained simulations, run for longer times, showed hundreds of lipid molecules crossing from one leaflet to the other along the outside of the X‑shaped groove, largely skirting the ion pore. The routes of ions and lipids overlapped only minimally, indicating that the protein runs two largely independent tracks side by side: an inner one for ions and an outer one for lipids.
A flexible controller tied to cell signals
The study also uncovers how the protein’s inner, cytosolic domain works like a mechanical linkage that converts calcium binding into these large movements. In the resting state, this domain forms a compact structure held together by a network of electrostatic "locks" between specific amino acids. Calcium binding straightens the TM6 segment and nudges this domain; breaking the salt‑bridge locks lets part of the domain rotate upward and latch onto TM6. That motion is transmitted to the TM4–TM5 region, causing TM4 to slide and form the X‑shaped groove. Mutations that weaken the lock speed up scrambling, while those that favor the closed state slow or block it.
What this means for health and disease
In everyday terms, this work shows that TMEM16F is a tiny, calcium‑controlled machine that can simultaneously open an internal pipe for ions and sculpt the surrounding membrane so that lipids slip across it. Rather than sharing a single mixed pathway, ions and lipids mostly travel on separate tracks shaped by the same protein motion. By revealing how calcium binding, internal rearrangements and membrane bending all cooperate, the study provides a blueprint for understanding diseases linked to TMEM16F and for designing drugs that can fine‑tune its activity in blood clotting, membrane repair and viral infections.
Citation: Feng, Z., Alvarenga, O.E., Di Zanni, E. et al. Calcium dependent activation of the TMEM16F scramblase and ion channel. Nat Struct Mol Biol 33, 664–676 (2026). https://doi.org/10.1038/s41594-026-01789-5
Keywords: TMEM16F, lipid scrambling, ion channels, cell membrane, calcium signaling