Clear Sky Science · en
A nucleotide code governs Lis1’s ability to relieve dynein autoinhibition
How cells keep their tiny cargo movers under control
Inside every cell, miniature machines haul cargo along slender tracks, helping to shape tissues, divide chromosomes and position the nucleus. One of the most important of these haulers is a motor protein called dynein. Because dynein is so powerful and so widely used, cells must keep it safely switched off until and unless it is needed. This paper reveals how another protein, Lis1 (called Pac1 in yeast), reads a subtle “nucleotide code” in dynein to flip it from a folded, locked state into an open, active form that can move cargo.
A molecular engine with a safety lock
Dynein is a large, multi-part motor that walks along microtubules, the cell’s internal rails, carrying vesicles, RNA and even whole nuclei. To avoid misdelivery, dynein normally sits in a self-clamped, autoinhibited conformation nicknamed the “phi” particle, in which its two motor heads fold together and barely interact with microtubules. For long-distance runs, dynein must first unfold and then combine with another complex, dynactin, plus a cargo adaptor. Earlier work hinted that Lis1 helps unlock dynein, but the details were murky, in part because dynein’s motor contains several nucleotide-binding pockets that can each hold different forms of ATP or ADP. The authors set out to decode how these pockets, and Lis1 binding, work together to control dynein’s shape.
Discovering a hidden nucleotide “code”
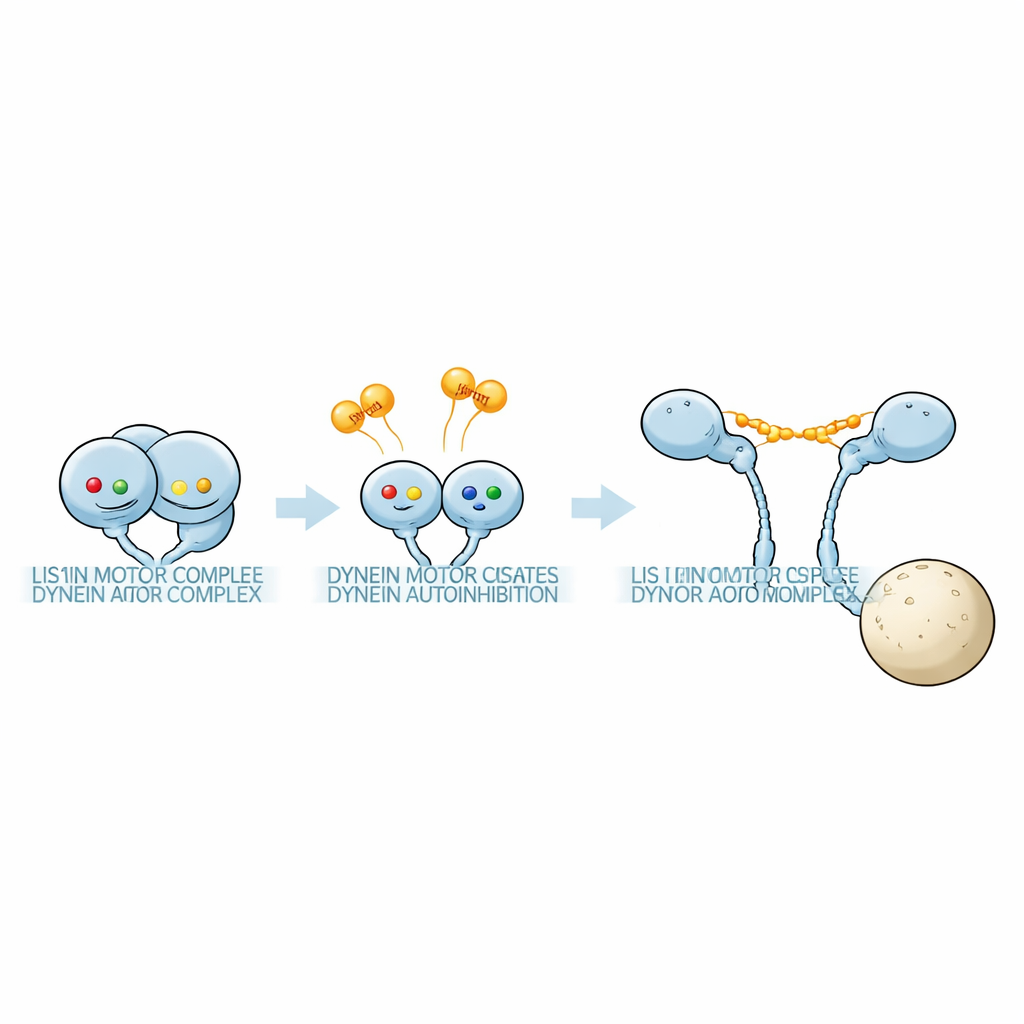
The team used sensitive mass photometry to weigh protein complexes in solutions containing dynein motor fragments and Pac1 under many nucleotide conditions. Dynein’s motor ring carries three active pockets, AAA1, AAA3 and AAA4. By introducing precise mutations that either prevent nucleotide binding or block ATP hydrolysis at each pocket, the researchers could lock individual sites into defined states. They found that the combination of nucleotides at these three sites strongly influenced how tightly Pac1 bound and, crucially, how many Pac1 units attached to each dynein. In some nucleotide combinations, a Pac1 dimer bridged two separate dynein motors; in others, it contacted only one motor; and in still others, it barely bound at all. These patterns revealed that distinct nucleotide “codes” select between a 1:1 dynein:Pac1 complex and a 1:2 complex with extra Pac1 bound.
How Lis1 reshapes the motor
Electron microscopy then linked binding mode to shape. When Pac1 formed a 1:1 complex with full-length dynein, the motors almost completely shifted from the phi conformation into an open form, with the two heads separated at a fixed distance. Negative-stain images showed Pac1 dimers bridging the two motor domains, matching the spacing seen with isolated motor–Pac1 pairs. In contrast, when conditions favored a 1:2 complex, dynein remained in a phi-like arrangement: more flexible and slightly distorted, but still compact rather than fully open. Thus, it is not simply the presence of Pac1 that matters, but exactly how many Pac1 molecules bind and where.
New contact points that unlock the clamp
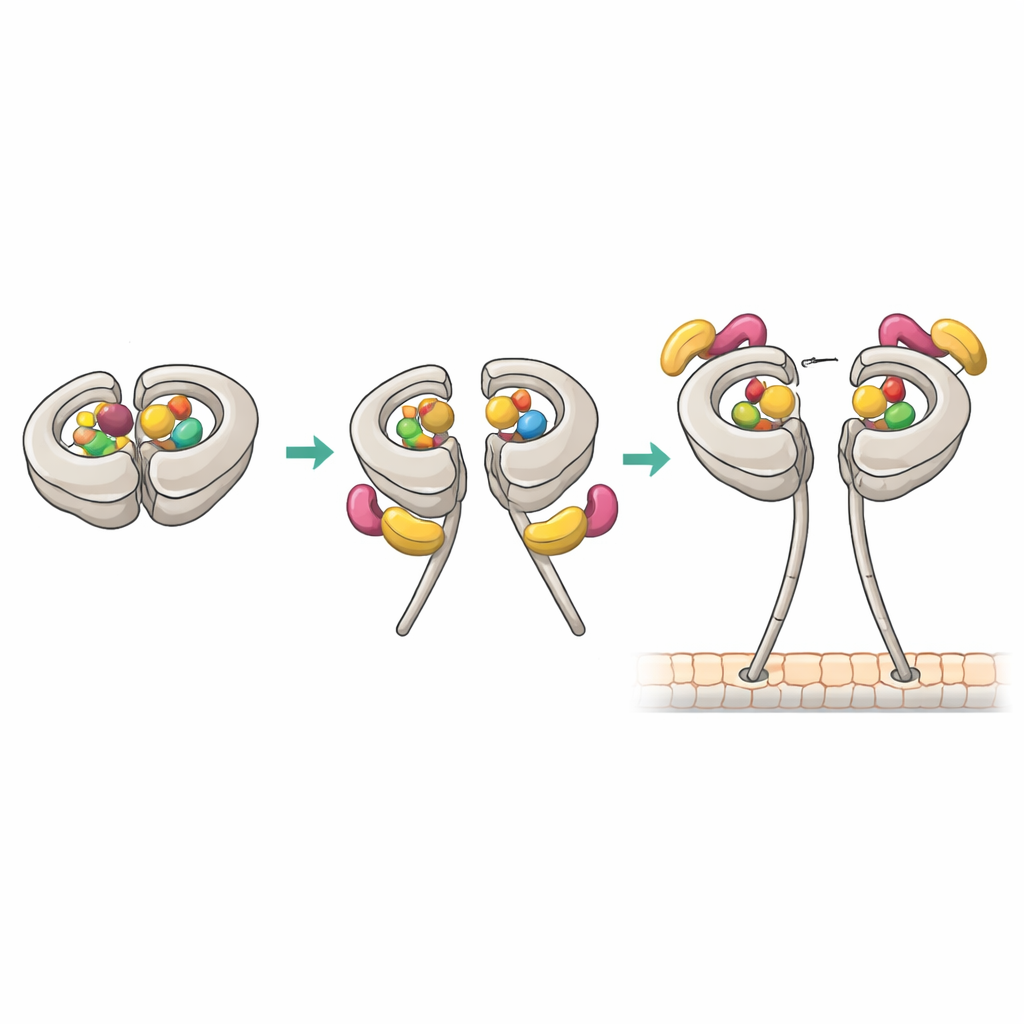
Cryo-electron microscopy at near-atomic resolution revealed how the productive 1:1 complex is built. In that state, dynein’s primary site AAA1 and a regulatory site AAA4 both hold ADP, while AAA3 is empty. This nucleotide pattern correlates with high affinity for Pac1. One WD40 “blade” of Pac1 docks at a previously known site on the motor ring, but the structure also uncovered a second contact on dynein’s flexible linker region. These dual grip points stabilize a modified “post-stroke” position of the linker and push Pac1 slightly toward the stalk, which in turn blocks binding at a second ring-adjacent site that would otherwise host another Pac1 WD40. Mutating the linker contact weakened Pac1’s ability to open dynein, shifted binding stoichiometry, and impaired dynein-driven spindle movements in yeast cells, even though the basic motor activity remained intact.
From biochemical switch to cellular movement
Piecing these results together, the authors propose a stepwise activation pathway. Dynein typically sits in phi with ADP in several pockets, which allows Pac1 to bind in a way that does not open the motor. Occasionally, one regulatory pocket (AAA3) releases ADP, letting the linker swing into a particular position. At that moment, Pac1 can clamp onto both the ring and the linker of each motor head, creating a single Pac1 dimer bridging the pair of heads and stabilizing an open conformation with high affinity for microtubules. This open state is then primed for docking onto dynactin and cargo adaptors to form a fully motile transport machine. Once dynein engages a microtubule and begins walking, Pac1’s grip weakens and it dissociates, handing off control to other regulators.
Why this matters for health and disease
For a non-specialist, the key message is that dynein is not simply turned on or off; it is tuned by a combination of nucleotide states and Lis1 binding that together form a safety logic. Only when the right nucleotide code appears does Lis1 act as a molecular wedge, opening the motor and enabling it to join the long-distance transport team with dynactin. Because human Lis1 is essential for brain development and its malfunction causes lissencephaly and other neurological disorders, understanding this code at atomic detail offers a roadmap for how subtle changes in Lis1 or dynein can derail cellular logistics and, ultimately, organismal development.
Citation: Geohring, I.C., Chai, P., Iyer, B.R. et al. A nucleotide code governs Lis1’s ability to relieve dynein autoinhibition. Nat Chem Biol 22, 649–662 (2026). https://doi.org/10.1038/s41589-025-02096-8
Keywords: dynein motor, Lis1 Pac1, molecular transport, protein autoinhibition, cryo electron microscopy