Clear Sky Science · en
Quantifying phage-host dynamics using droplet microfluidics
Why tiny viruses and droplets matter to us
Bacteria-killing viruses, called bacteriophages, are being revisited as potential allies against drug‑resistant infections. But to turn them into reliable medicines, scientists must measure how efficiently individual viruses infect and destroy bacteria, something classic lab methods struggle to do. This paper introduces a droplet‑based approach that watches thousands of miniature battles between single viruses and bacteria at once, revealing how quickly and how often infections succeed. Such precise insights could guide better phage therapies and deepen our understanding of the hidden viral world that shapes microbial life on Earth.
Limits of traditional petri‑dish tests
For decades, researchers have relied on plaque assays: mix viruses with a lawn of bacteria in soft agar, wait overnight, and then count clear spots where bacteria have been wiped out. These spots estimate how many infectious viruses were present. While robust, this method is slow, inflexible, and offers only a final snapshot. It cannot track how infections unfold over time, and it blends together countless rounds of infection and reinfection. Other approaches, like bulk fluorescence or DNA‑based tests, still average over huge populations, making it hard to see single infection events or to tune conditions such as how many viruses meet each bacterium.
Turning each infection into a tiny test tube
The authors build a high‑throughput platform that packages bacteria and phages into microscopic droplets of water suspended in oil. Two separate liquid streams—one carrying bacteria, the other viruses—meet only at the narrow junction where droplets are formed, so the exposure time starts precisely at encapsulation. Each droplet becomes an isolated reaction chamber with a well‑defined volume and a controlled ratio of phages to bacteria. The team adds a special fluorescent dye that lights up strongly when bacterial DNA is released after the cell bursts, serving as a direct readout of successful lysis rather than just the presence of viral genetic material.
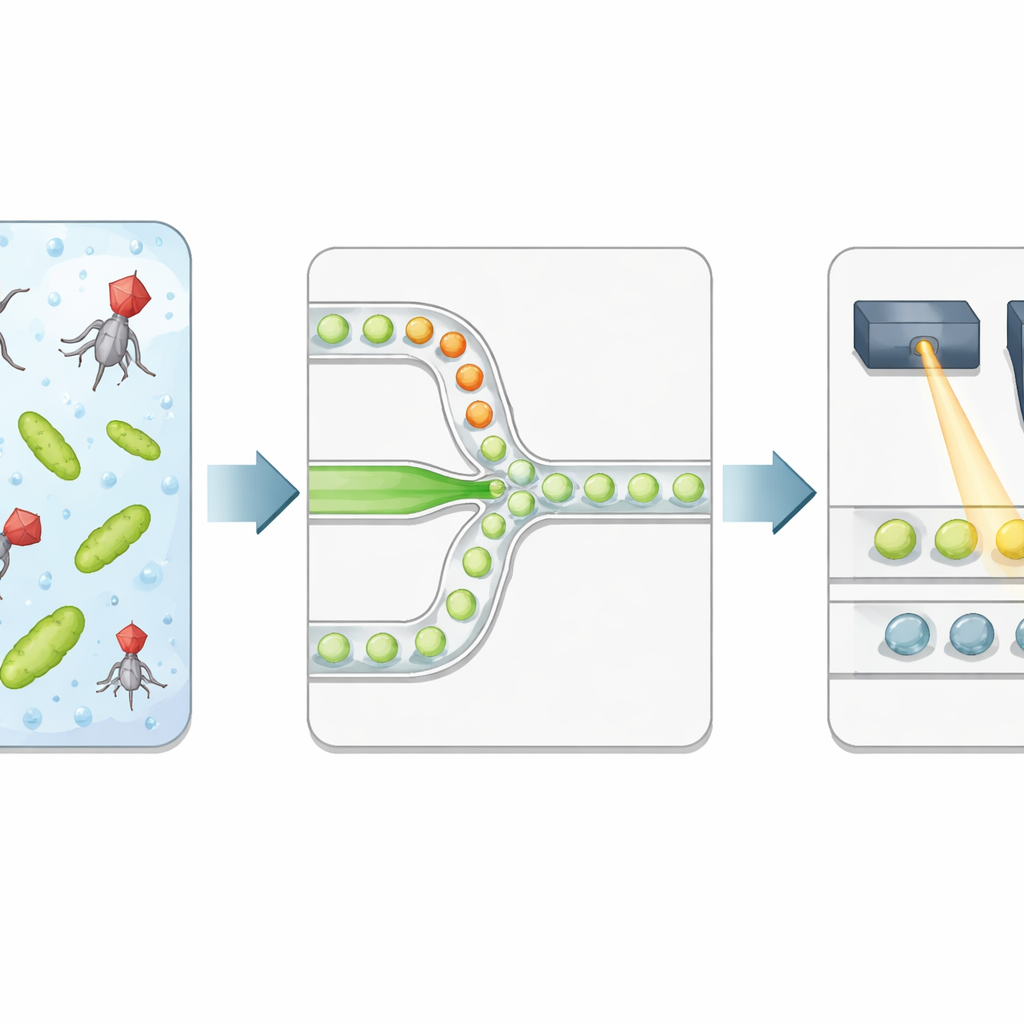
Reading droplet outcomes with light
After a controlled incubation, the emulsion is flowed through a second microfluidic chip that lines up droplets one by one under a focused laser. A fast detector records green fluorescence from the DNA dye and red fluorescence from a reference dye included during droplet production. The red signal reports droplet size and how much of the virus solution each droplet received, while the green signal reports whether lysis occurred. An automated statistical method separates dim droplets (no lysis) from bright ones (at least one infected and lysed bacterium), converting the experiment into a string of digital 0s and 1s. By relating the fraction of bright droplets to the expected numbers of viruses and bacteria per droplet, the researchers recover the virus concentration and compare it to classic plaque counts.
Tuning conditions and probing hidden kinetics
The platform goes beyond simple counting. By deliberately changing the mixing ratio of virus and bacterial solutions, the team generates subpopulations of droplets with different virus‑to‑host ratios, all within a single experiment. This effectively stretches the range of concentrations over which the assay gives precise results and even reveals that not all bacteria seen by bulk optical measurements are actually able to be infected. The authors also vary droplet size and incubation time to watch how the share of lysed droplets grows. Their model of how viruses adsorb to bacteria inside droplets shows that the timing and success of lysis depend on the true number of infection‑capable cells and on how often virus particles collide with them, but not strongly on droplet volume itself.
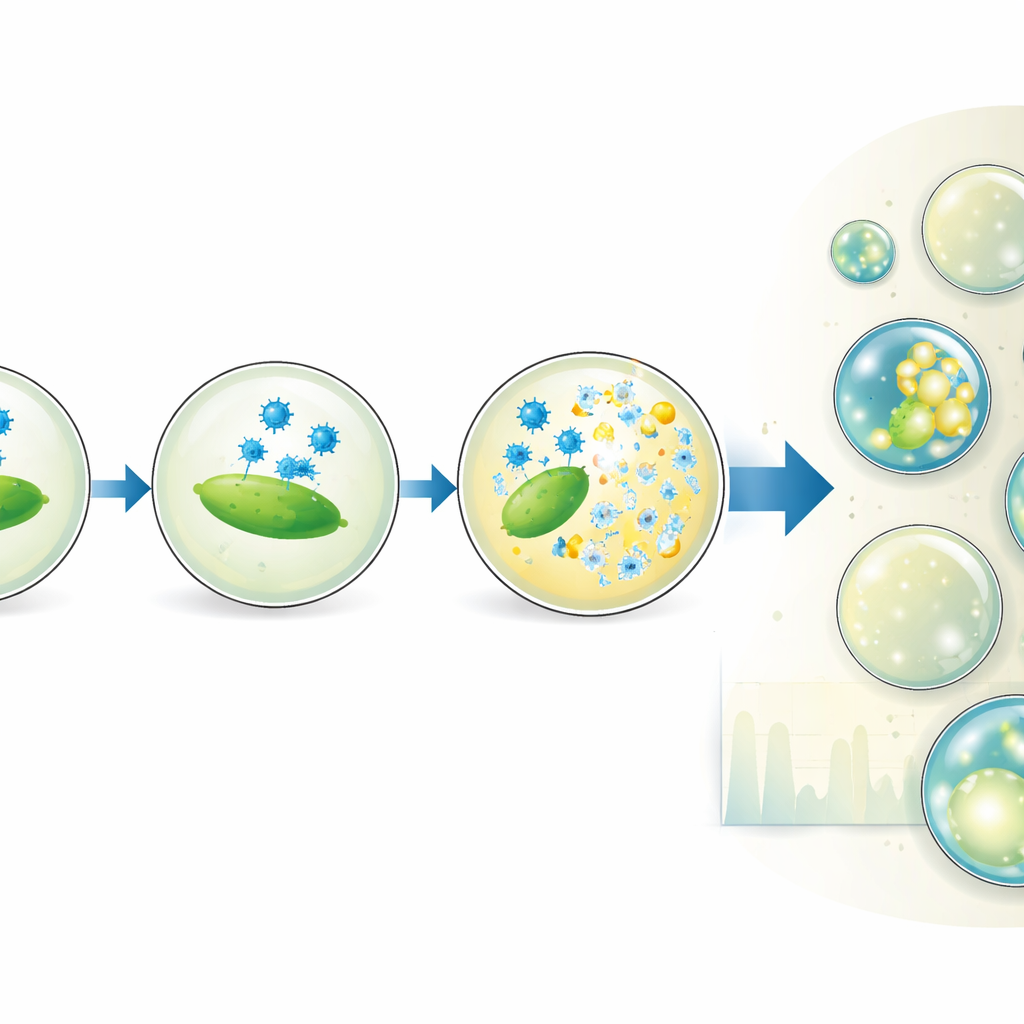
From lab tool to future therapies
By isolating and digitizing individual infection events, this droplet method delivers accurate virus counts that closely match traditional plaque assays, while adding time‑resolved information and fine control over experimental conditions. It can be applied to many different lytic phage–bacteria pairs and scaled to millions of droplets, opening the door to high‑throughput screening of phage variants, media, or temperatures. In practical terms, the work provides a powerful new way to characterize how well therapeutic phages infect their targets, an important step toward using these natural bacterial predators to help combat antibiotic‑resistant infections.
Citation: Givelet, L., von Schönberg, S., Katzmeier, F. et al. Quantifying phage-host dynamics using droplet microfluidics. Nat Commun 17, 3857 (2026). https://doi.org/10.1038/s41467-026-72427-3
Keywords: bacteriophages, microfluidic droplets, phage therapy, single-cell infection, antibiotic resistance