Clear Sky Science · en
2′-O-methylation-dependent installation of N2-methylguanosine in the U6 internal stem loop facilitates efficient spliceosome assembly
Why tiny RNA tweaks matter
Inside every human cell, an intricate molecular machine called the spliceosome edits raw RNA messages so they can be turned into working proteins. This editing must be both fast and accurate, and it relies on small RNAs that are heavily "fine‑tuned" with chemical tags. This paper uncovers how a specific sequence of chemical tweaks on one such RNA, called U6, helps the spliceosome assemble correctly and handle especially difficult RNA segments, with potential implications for diseases where splicing goes wrong.
A key helper in RNA editing
Most human genes contain non‑coding stretches that must be removed from RNA before proteins are made. The spliceosome carries out this cutting‑and‑pasting using several small RNAs, among them U6, which forms part of the catalytic heart of the machine. U6 does not act alone: it partners with proteins and other RNAs in a stepwise assembly process. Along the way, U6 is chemically modified at many positions. The authors focus on an internal loop region of U6 that carries several sugar methyl groups and a special modification on one guanosine base. Earlier work showed that the base methylation, known as N2‑methylguanosine at position 72, is needed to splice weak sites and that the enzyme THUMPD2 installs this mark. What remained unknown was when this mark is added, how the enzyme recognizes U6, and how these chemical changes affect spliceosome assembly and splicing.
Finishing touch added late in U6’s life
Using UV crosslinking and high‑throughput sequencing, the researchers showed that THUMPD2 binds U6 RNA far more than any other RNA in cells, confirming U6 as its principal target. By analyzing the tail ends of U6 molecules associated with THUMPD2, they found that the enzyme engages U6 only after its 3′ end has been fully processed, meaning methylation at position 72 is a late event in U6 maturation. Protein‑interaction mapping revealed that THUMPD2 associates with U6‑containing complexes and early splicing factors, but not with the very earliest biogenesis factors. Fluorescence microscopy then placed THUMPD2 in Cajal bodies—nuclear substructures where late steps of spliceosome assembly occur—supporting the idea that the guanosine methylation is one of the final quality‑control marks before U6 joins larger spliceosomal assemblies.
Crosstalk between sugar and base modifications
To understand how THUMPD2 recognizes U6, the team combined structural modeling, crosslinking mass spectrometry, and mutational analysis. They built a model in which U6’s internal stem loop nestles against both the THUMP RNA‑binding domain and the catalytic methyltransferase domain of THUMPD2, placing the target base near the enzyme’s active site. They then showed that both domains are required to grip U6 stably, while a small partner protein, TRMT112, is essential for THUMPD2’s catalytic activity. Strikingly, test‑tube methylation assays revealed that THUMPD2 barely modifies an unmodified U6 loop but works over ten times more efficiently when two nearby ribose sugars in the opposite strand are already 2′‑O‑methylated. In cells, depleting LARP7—a factor needed to install these sugar methyl groups—reduced both the sugar marks and the guanosine methylation. Yet binding strength between THUMPD2 and U6 was similar with or without sugar methylation, indicating that these prior marks tune the reaction step itself, not initial recognition.
Impact on splicing patterns and spliceosome assembly
The authors next asked how these chemical marks influence actual gene splicing. By comparing RNA‑sequencing data from cells lacking THUMPD2, cells depleted of LARP7, and cells missing both, they found that each perturbation subtly but significantly altered alternative splicing choices. Loss of the guanosine methylation alone led mainly to increased intron retention, particularly in introns that are already inefficiently spliced. In contrast, loss of the sugar methylations shifted the usage of alternative 3′ splice sites toward more distant options. When both types of modification were impaired, the number of affected splicing events was highest, and the combined effects could largely be explained as an additive blend of the individual defects. Biochemical fractionation of nuclear extracts revealed the underlying assembly defects: without the base methylation, U6 accumulated in intermediate small RNA–protein complexes, suggesting slowed progression into fully formed spliceosomes. Without the sugar methylations, much less U6 entered small nuclear ribonucleoprotein particles at all, and key spliceosomal components were mis‑distributed, pointing to a more fundamental assembly problem.
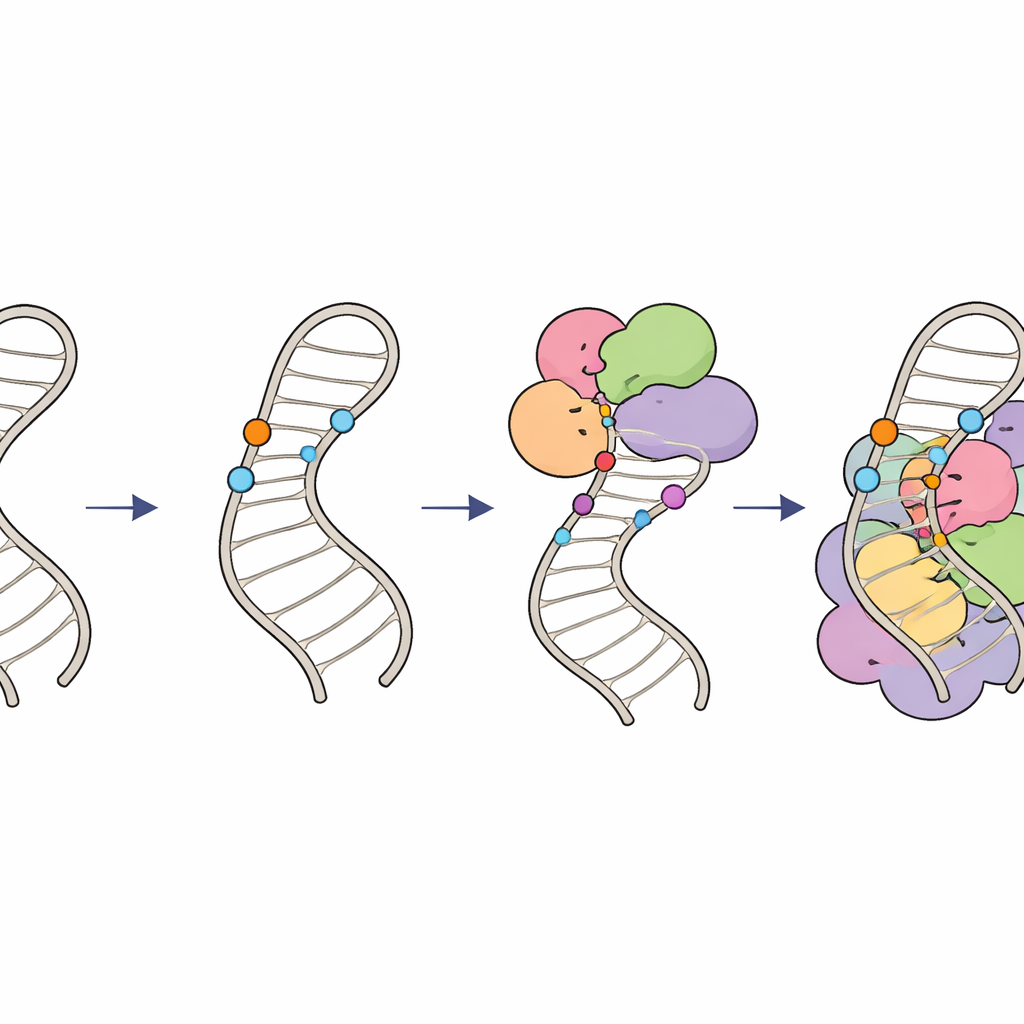
How chemical marks keep RNA editing on track
Together, these findings outline a hierarchical "modification circuit" on U6 RNA. First, specialized guide RNAs and the LARP7 factor place 2′‑O‑methyl groups on sugars within the internal stem loop, shaping its structure and creating an optimal substrate. Then, the THUMPD2–TRMT112 enzyme complex adds a finishing methyl group to a specific base, likely acting as a late‑stage license for U6 to progress into higher‑order spliceosomal complexes. By coordinating when and where these tiny chemical marks appear, cells ensure that U6 is folded correctly and plugged into the spliceosome with just the right timing, maintaining accurate and efficient RNA splicing. Subtle defects in this system can ripple out to change splicing patterns across the genome and may contribute to human diseases linked to splicing errors.
Citation: Kleiber, N., Petrosyan, J., Greve, M. et al. 2′-O-methylation-dependent installation of N2-methylguanosine in the U6 internal stem loop facilitates efficient spliceosome assembly. Nat Commun 17, 3793 (2026). https://doi.org/10.1038/s41467-026-72355-2
Keywords: RNA splicing, U6 snRNA, RNA modifications, spliceosome assembly, THUMPD2