Clear Sky Science · en
Structural analyses uncover protease-adhesin interactions and c-di-GMP receptor regulation in sulfate-reducing bacteria
Why sticky bacteria matter
Bacteria often live not as lone cells but as crowded, slimy communities called biofilms. These communities can coat pipes, ship hulls, underground rocks, and even medical devices. In sulfate‑reducing bacteria such as Desulfovibrio vulgaris, biofilms are a double‑edged sword: they help clean up polluted environments but also accelerate the corrosion of metals. This study asks a simple question with big implications: how do these bacteria decide when to cling tightly to a surface and when to let go?
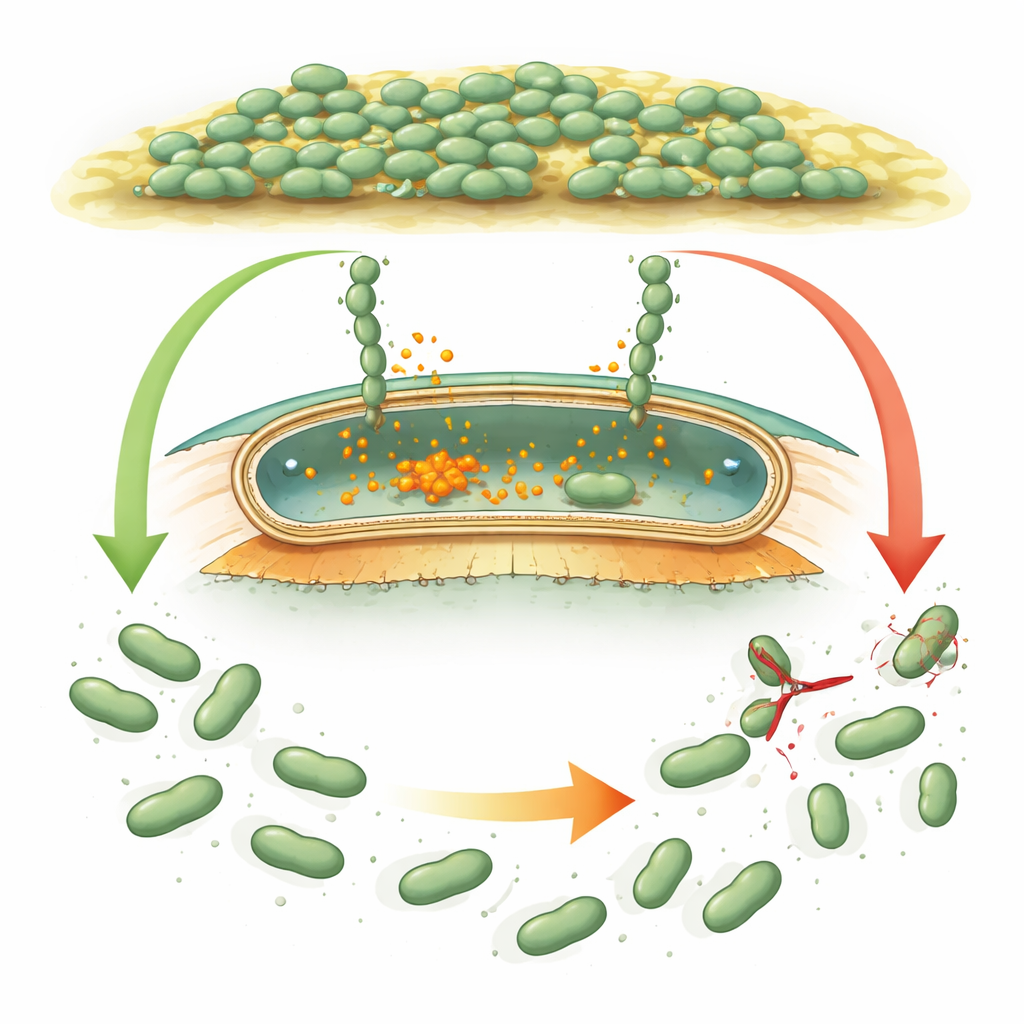
How microbes grip and release
Many bacteria use giant “Velcro‑like” proteins, called adhesins, that stick cells to surfaces and to each other. In well‑studied species like Pseudomonas, these adhesins are controlled by an internal chemical signal, cyclic di‑GMP, which rises when bacteria should form a biofilm and falls when they should disperse. When the signal is high, adhesins stay anchored in the outer shell of the cell. When it drops, a small protein cutter (a protease) snips the adhesin close to the cell surface, and the sticky portion floats away, helping cells detach. Desulfovibrio vulgaris carries what looks like a similar genetic toolkit, but its regulatory proteins differ so much in sequence and shape that scientists could not tell whether it used the same logic or an entirely new strategy.
A specialized molecular scissors
The authors first focused on DvhG, the Desulfovibrio version of the adhesin‑cleaving protease. Using structural prediction and biochemical tests, they showed that DvhG has the same core fold and catalytic machinery as its counterparts in other bacteria and requires calcium to work. It precisely cuts two large adhesins that Desulfovibrio needs for biofilm formation, at nearly identical positions just after a small “retention” segment that keeps these proteins lodged in the outer membrane. Intriguingly, the team found that DvhG does more than simply recognize a short target sequence. The bulky retention domain of each adhesin makes extensive contact with the protease, forming a large docking surface that boosts cutting efficiency. Mutations that disrupt this docking patch make DvhG much less effective, revealing an added layer of specificity compared with related systems.
A signal‑sensing switch inside the cell
The second key player, DvhD, sits in the inner membrane and reaches into both the cell’s interior and the space between the membranes. The researchers solved multiple three‑dimensional structures of DvhD fragments. Outside the cell, its periplasmic region forms a dCache domain, a common sensor architecture that often binds small molecules. Inside the cell, DvhD carries an HD‑GYP domain, a module usually found in enzymes that break down cyclic di‑GMP. Here, however, the crucial metal‑binding residues are missing, and careful assays showed that DvhD does not destroy the signal; instead, it binds cyclic di‑GMP very tightly. Structural snapshots revealed two distinct binding pockets where up to three cyclic di‑GMP molecules can bridge a pair of HD‑GYP domains and force them into a new arrangement, prying apart long helices that connect to the rest of the receptor.
From internal signal to surface behavior
By comparing the “empty” and signal‑bound forms of DvhD, the team uncovered how binding cyclic di‑GMP can reshape the receptor dimer. These changes are expected to travel through the inner membrane and reposition the periplasmic dCache domain, which in turn interacts with DvhG. Modeling suggests that when DvhD engages DvhG, it blocks the protease’s access to adhesins and may even distort its calcium‑binding site, turning the scissors “off.” When cyclic di‑GMP levels fall, the receptor’s shape relaxes, releasing DvhG so it can clamp onto the adhesins’ retention domains and cut them, allowing the outer portions of the adhesins—and thus the biofilm matrix—to slip away from the cell surface. In this way, Desulfovibrio applies an internal chemical readout of its circumstances to a mechanical decision: stay attached or disperse.
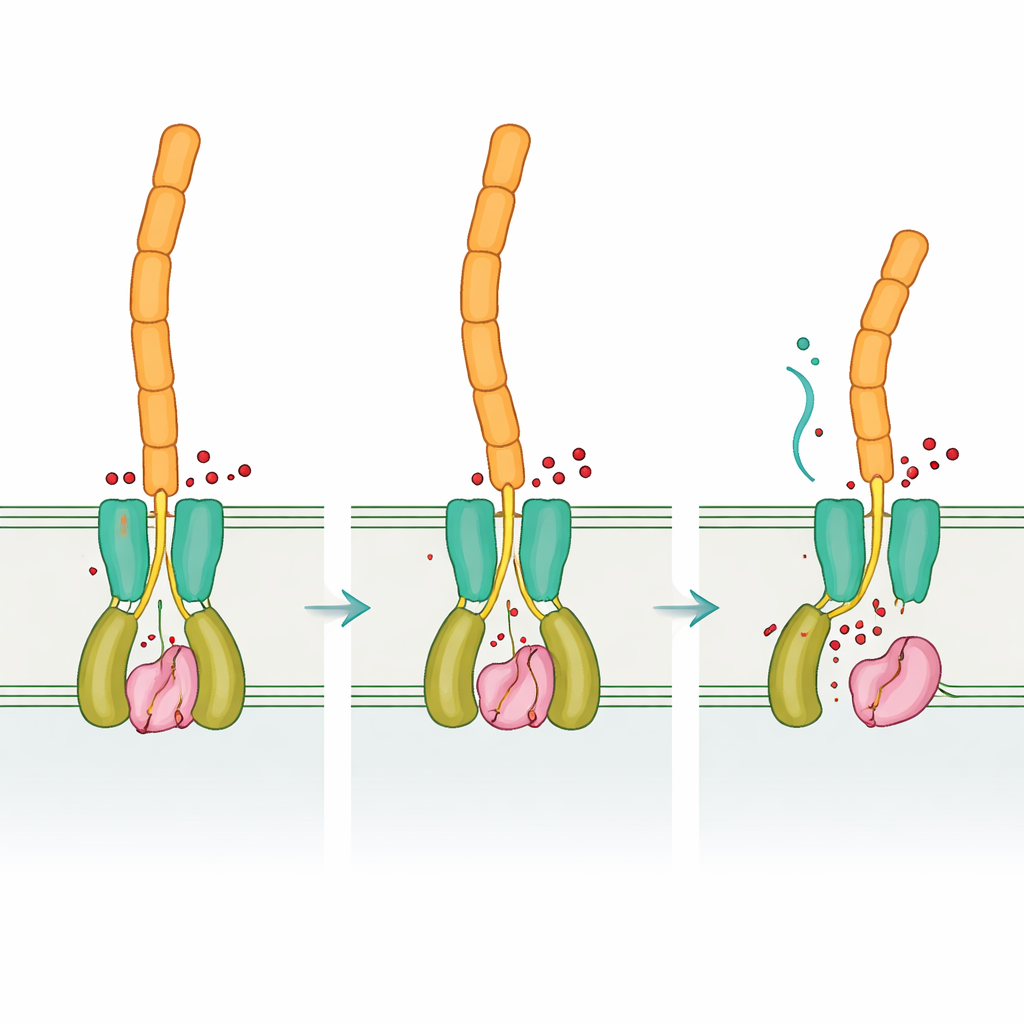
Same logic, different hardware
Taken together, the results show that Desulfovibrio vulgaris uses the same overall regulatory logic as more familiar bacteria—cyclic di‑GMP controls a receptor, which in turn restrains or releases a protease that clips surface adhesins—but implements it with a very different set of protein “parts.” This mix of conservation and reinvention illustrates how evolution can reuse successful signaling ideas while reshaping the molecules that carry them out. Understanding this flexible design not only deepens our grasp of how biofilms form and break apart across many bacterial groups, but may also point the way toward smarter strategies to either promote helpful biofilms in bioremediation or disrupt damaging ones that drive metal corrosion and chronic infections.
Citation: Font, M.E., Karbelkar, A.A., Lormand, J.D. et al. Structural analyses uncover protease-adhesin interactions and c-di-GMP receptor regulation in sulfate-reducing bacteria. Nat Commun 17, 3564 (2026). https://doi.org/10.1038/s41467-026-71936-5
Keywords: bacterial biofilms, Desulfovibrio vulgaris, cyclic di-GMP signaling, adhesin protease, sulfate-reducing bacteria