Clear Sky Science · en
Phage-assisted evolution of allosteric protein switches
Turning Proteins into On–Off Switches
Imagine being able to flick a molecular light switch inside a living cell—turning genes, enzymes, or signaling pathways on and off at will. This paper describes how scientists built an evolutionary “training ground” that teaches proteins to behave like such switches. By harnessing viruses that infect bacteria and a clever selection strategy, the team evolves proteins that respond sharply to light, offering powerful new tools for research, biotechnology, and potentially medicine.
Why Remote Control of Proteins Matters
Proteins are the tiny machines that run our cells. Many of them are naturally allosteric, meaning that a change at one spot in the protein—triggered by a signal such as a sugar, a chemical, or light—causes a functional shift at another, distant spot. Scientists would like to redesign this built-in wiring so that almost any protein can be controlled by a chosen input, like blue light. Yet this has proved difficult: when researchers simply bolt a sensor domain onto a working protein, the chimera often becomes weak, leaky, or barely responsive. The challenge is that the “rules” of allostery are spread across many interacting residues and shapes, making them hard to predict by design alone.
Borrowing Evolution’s Playbook
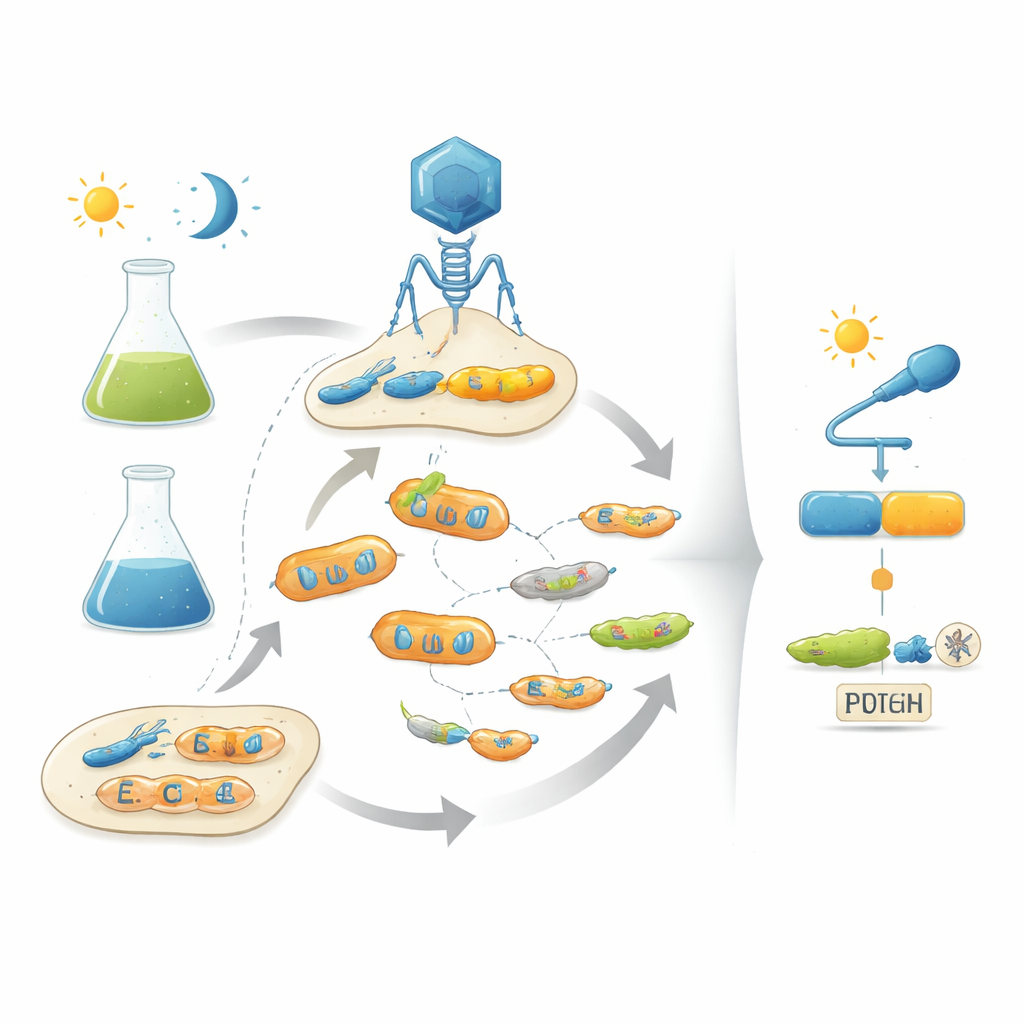
The authors tackle this problem by mimicking natural evolution in the lab using bacteriophages—viruses that infect bacteria. Their platform, called POGO-PANCE, links the success of each phage to how well its encoded protein behaves as a switch. In one phase, if the protein is active under the desired condition (for example, in the dark), the phage prospers. In the opposite phase, if the protein is active when it should be off (for example, under light), a sabotaging version of a viral component is produced, and the offending phage line collapses. By alternating these positive and negative selection steps while constantly mutating the protein, the system nudges viral populations toward variants that are both powerful and tightly controlled by the input signal.
Training a Protein to Respond to Light
To demonstrate their strategy, the researchers focus on AraC, a well-studied bacterial protein that normally responds to the sugar arabinose to control gene expression. First, they evolve AraC itself to become strongly active even without its natural sugar trigger, providing a “high-powered” starting point. Next, they insert a blue-light–sensitive module, known as a LOV domain, into AraC at a site predicted to allow allosteric communication. Initially, this fusion nearly breaks AraC: it barely works and only shows weak light responsiveness when overproduced. Running these crippled hybrids through POGO-PANCE, however, quickly transforms them. After several rounds of alternating selection under light and dark, the team recovers variants that behave almost like digital switches, with gene activity changing by about a thousand-fold between dark and illuminated states.
Peering into the Wiring of a Molecular Switch
Because the evolving proteins are encoded on the phage genome, the researchers can sequence the viral populations after each cycle. This reveals which mutations rise and fall under different selection steps, tracing the pathways by which effective switches emerge. They find that changes appear not only in AraC and the light sensor but also in the short linkers that join them. Using a second tool, RAMPhaGE, based on retron-guided genome editing, they systematically reshape these linkers by adding, removing, or swapping small stretches of amino acids. Evolution repeatedly favors variants where the linker at one side of the LOV domain extends into a smooth helix that physically bridges sensor and effector. This suggests that a continuous, well-aligned helix helps transmit the light-induced shape change through the protein, sharpening the on–off response.
What This Means for Future Bioengineering
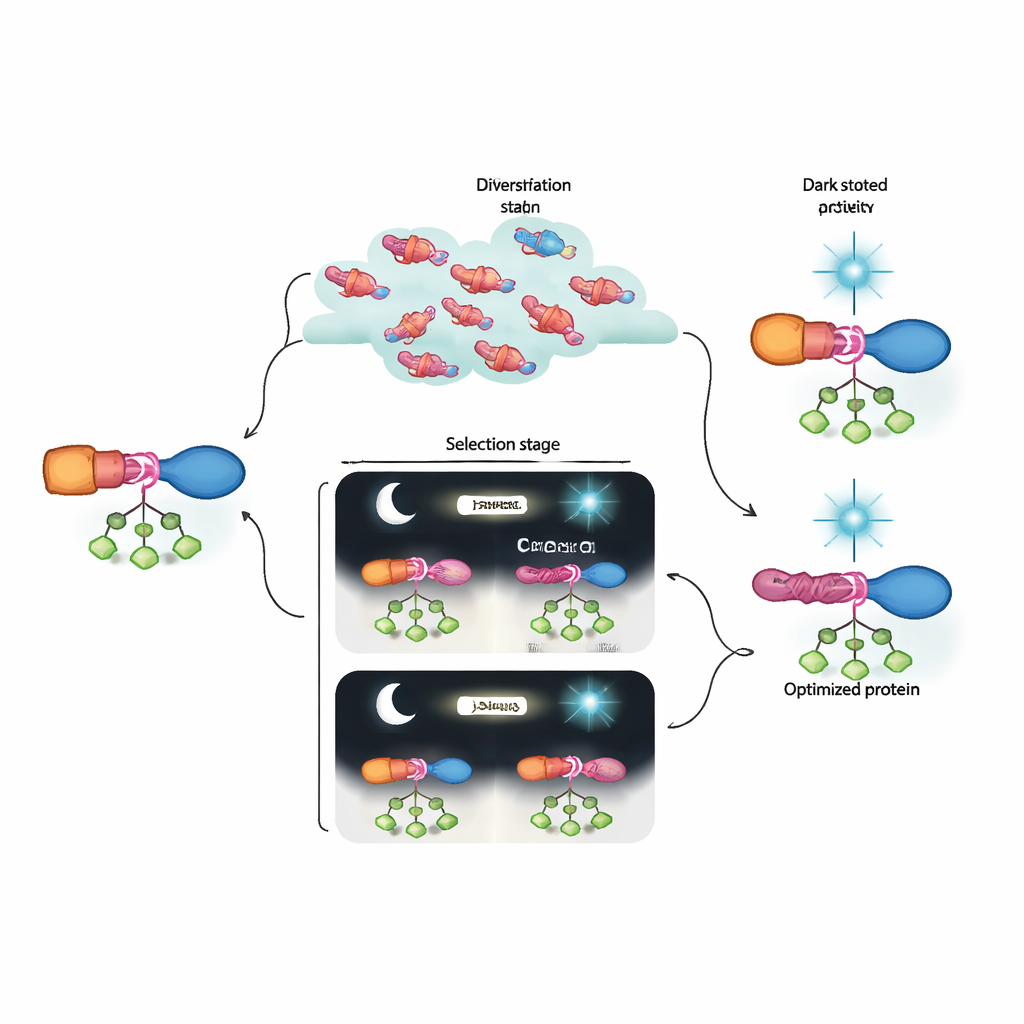
In plain terms, the authors have built a lab-based “evolution machine” that can discover and refine protein switches far beyond what current design tools can easily produce. Their evolved AraC–LOV hybrids show strong activity when needed and shut down almost completely under blue light, outperforming earlier optogenetic versions. Just as important, the combination of dynamic selection and deep sequencing reveals how groups of subtle mutations cooperate to build an allosteric pathway. This framework could be adapted to many other signaling inputs and protein targets, moving us closer to a future where cellular processes can be programmed with the precision of electronic circuits, but using components that evolution itself has helped to design.
Citation: Southern, N.T., von Bachmann, A., Hovsepyan, A. et al. Phage-assisted evolution of allosteric protein switches. Nat Commun 17, 3498 (2026). https://doi.org/10.1038/s41467-026-71717-0
Keywords: optogenetics, directed evolution, allosteric regulation, protein engineering, bacteriophage