Clear Sky Science · en
RB loss modulates chromatin organization by regulating cohesin-dependent loops and enhancer-promoter interactions
Why the Cell’s Package Manager Matters
Every cell in your body must squeeze about two meters of DNA into a nucleus smaller than a speck of dust. How that DNA is folded—not just what genes it carries—helps decide which genes are turned on or off. This paper explores a famous cancer-fighting protein called RB and reveals that it does much more than halt runaway cell division. RB also helps manage the three-dimensional folding of DNA, fine-tuning which genes can be reached by nearby control switches. Understanding this hidden role offers new clues to how cancers become aggressive when they lose RB.
A Guardian with a Hidden Job
RB (short for retinoblastoma protein) is best known as a brake on the cell cycle. It normally keeps a set of growth-driving genes quiet until the cell is ready to divide. Many tumors disable RB, which lets cells multiply uncontrollably. Yet RB-deficient cancers also show puzzling changes—such as increased invasiveness and the ability to switch cell type—that cannot be explained solely by loss of this cell-cycle brake. Earlier work hinted that RB binds not only to gene starters (promoters) but also to distant control regions (enhancers) and special “insulator” sites that mark the boundaries between neighborhoods of DNA activity. This raised the possibility that RB might also act as an organizer of the genome’s folding.
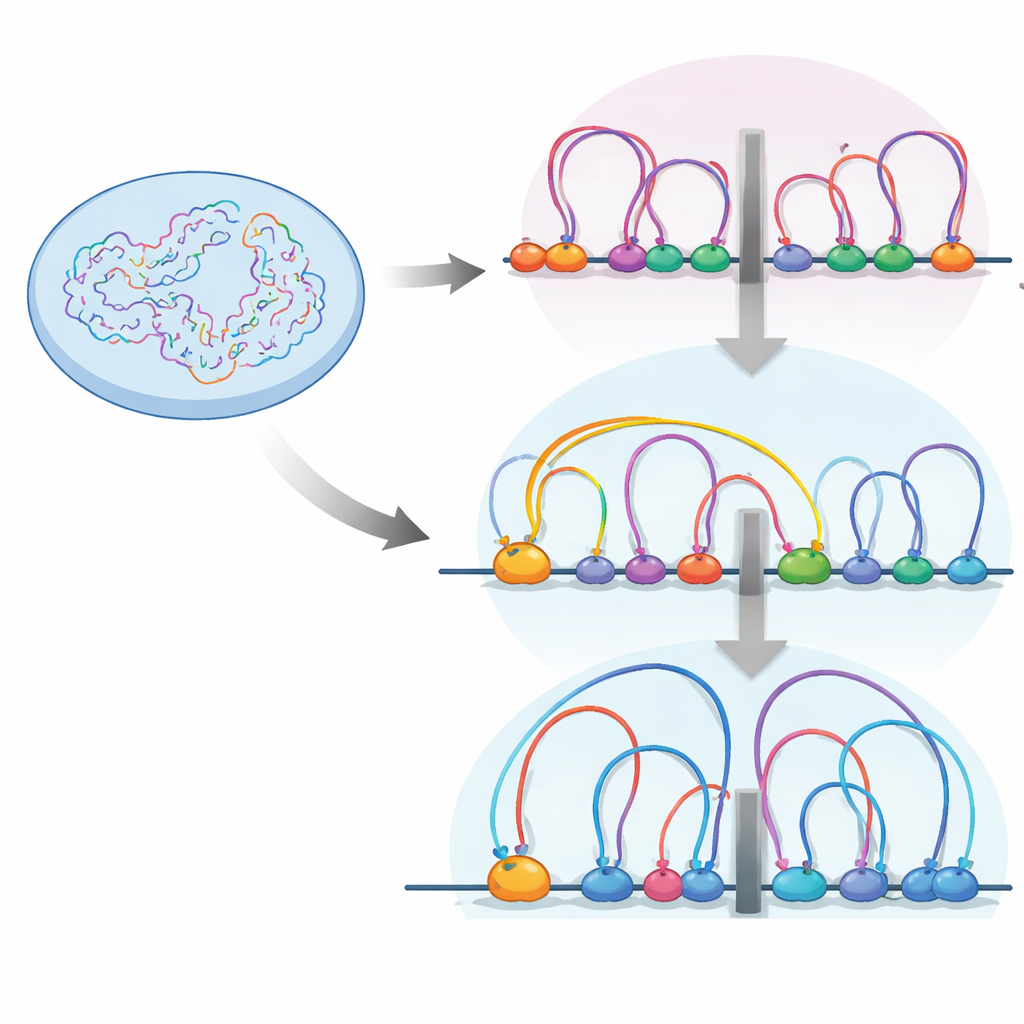
Loops, Boundaries, and Traffic Control
Inside the nucleus, the DNA fiber is folded into loops by a protein machine called cohesin. These loops help group genes and their control elements into units called topologically associating domains, or TADs—essentially neighborhoods within which genes and switches interact more freely. Boundaries between these neighborhoods are often marked by an insulator protein called CTCF, where cohesin tends to pause. The authors used high-resolution chromosome contact mapping (Micro-C) and a suite of other genomic tools to see how turning RB on or off reshapes this looping landscape in human retinal cells.
They found that when RB is missing, the number and average size of DNA loops increase, and TAD boundaries become stronger. At the same time, RB and cohesin sit together at many accessible regions across the genome, especially at insulators. Detailed mapping of cohesin’s location during different cell-cycle stages showed that RB is needed to help remove cohesin from these insulator sites during mitosis, the phase when chromosomes condense and then re-open. Without RB, cohesin piles up at insulators, strengthening boundaries and changing how DNA neighborhoods are insulated from one another.
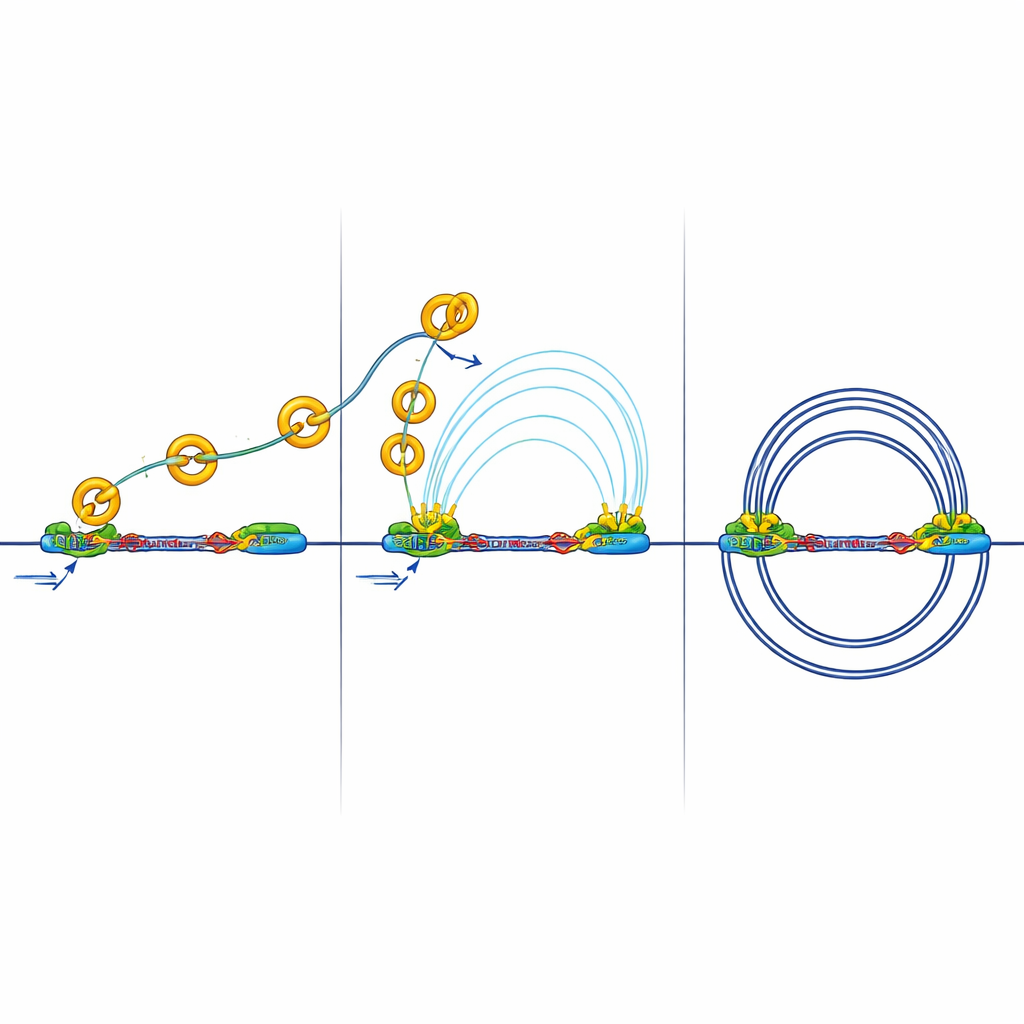
When Insulation Blocks Gene Conversations
Stronger boundaries have consequences. Insulators can block the “long-distance calls” between enhancers and promoters that are needed to activate genes. The team measured physical contacts between active regions of DNA marked by a chemical tag called H3K27ac. In cells lacking RB, they saw fewer and weaker contacts between enhancers and promoters, even though many activation marks were still present. In other words, the switches were on, but their signals had trouble crossing the reinforced boundaries. Additional experiments showed reduced recruitment of a key mediator complex that normally helps enhancers talk to promoters, supporting the idea that the 3D wiring, rather than the basic on/off marks, was being disrupted.
From DNA Folding to Cell Behavior
To link these architectural changes to real genes and traits, the authors focused on genes whose promoters are not directly bound by RB or its usual partner factors, but that sit just beyond an insulator where cohesin levels change when RB is lost. For genes next to insulators that gained cohesin (and thus stronger insulation), expression tended to drop in RB-deficient cells, even though RB often globally raises many growth-related genes when it is removed. These affected genes were enriched for roles in maintaining epithelial features and organizing the cell’s surrounding matrix—properties tied to how tightly cells stick together and how easily they move.
Consistent with these gene changes, RB-lacking cells detached more readily from the culture surface and closed artificial “wounds” faster, both signs of greater mobility. When the authors grouped RB-controlled genes by behavior across the cell cycle, they identified a subset that RB actually activates, rather than represses, in a way that depends on its ability to loosen insulation at nearby boundaries. This suggests RB does not just act as a brake on growth; it also serves as a positive regulator for specific sets of genes that help preserve normal tissue architecture.
What This Means for Cancer
For non-specialists, the key message is that RB guards cells not only by stopping them from dividing at the wrong time but also by shaping how their DNA is folded and wired. By promoting the removal of cohesin from boundary elements, RB weakens certain barriers and allows important control switches to reach their target genes. When RB is lost, these boundaries become too strong, some genes that maintain normal cell identity are silenced, and cells become more prone to detach and migrate—traits that can fuel tumor spread and treatment resistance. This expanded view of RB as an architect of the genome’s 3D structure may help explain why its loss so often marks a turning point toward more dangerous, hard-to-treat cancers.
Citation: Lee, H., Gkotinakou, IM., McGrath, C.G. et al. RB loss modulates chromatin organization by regulating cohesin-dependent loops and enhancer-promoter interactions. Nat Commun 17, 3696 (2026). https://doi.org/10.1038/s41467-026-71655-x
Keywords: retinoblastoma protein, chromatin loops, cohesin, 3D genome organization, cancer progression