Clear Sky Science · en
Versatile and sensitive detection of mono- and poly(ADP-ribosyl)ation reveals XRCC1-dependent remodelling of PARP1 signalling
Why tiny chemical tags on our DNA repair system matter
Every cell in our body is constantly repairing damage to its DNA. If this repair goes wrong, the result can be cancer, neurodegeneration, or cell death. A key player in this repair network is a protein called PARP1, which decorates itself and nearby proteins with small sugar-like tags made from NAD+. These tags come in short and long forms and act like signals that call in other repair factors. Until now, scientists have lacked precise tools to see and distinguish these different tag types inside cells, especially during disease. This study builds such tools and uses them to reveal an unexpected change in DNA repair signaling when another repair protein, XRCC1, is missing.
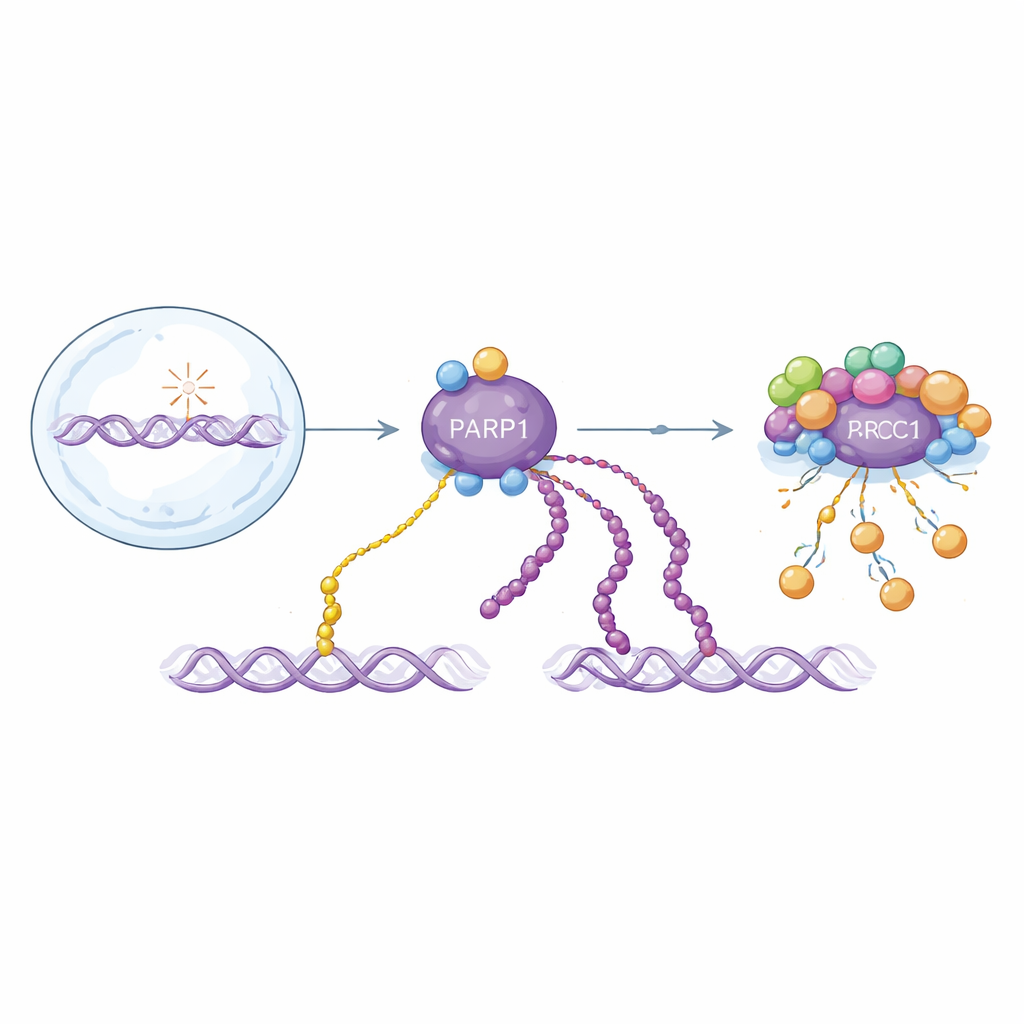
Two flavors of a critical DNA repair signal
PARP1 responds to DNA strand breaks by attaching ADP-ribose units to itself and other proteins. These modifications exist as single units (mono-ADP-ribosylation) or as long chains (poly-ADP-ribosylation). Both forms help organize repair, but they behave differently over time: long chains appear in a rapid initial burst and are quickly removed, while single units accumulate more slowly and persist longer. Because both forms are built from the same building block, most existing detection methods blur them together. That makes it hard to understand how each one contributes to health and disease, or how drugs that target PARP1 really work.
Engineering highly selective molecular "eyes"
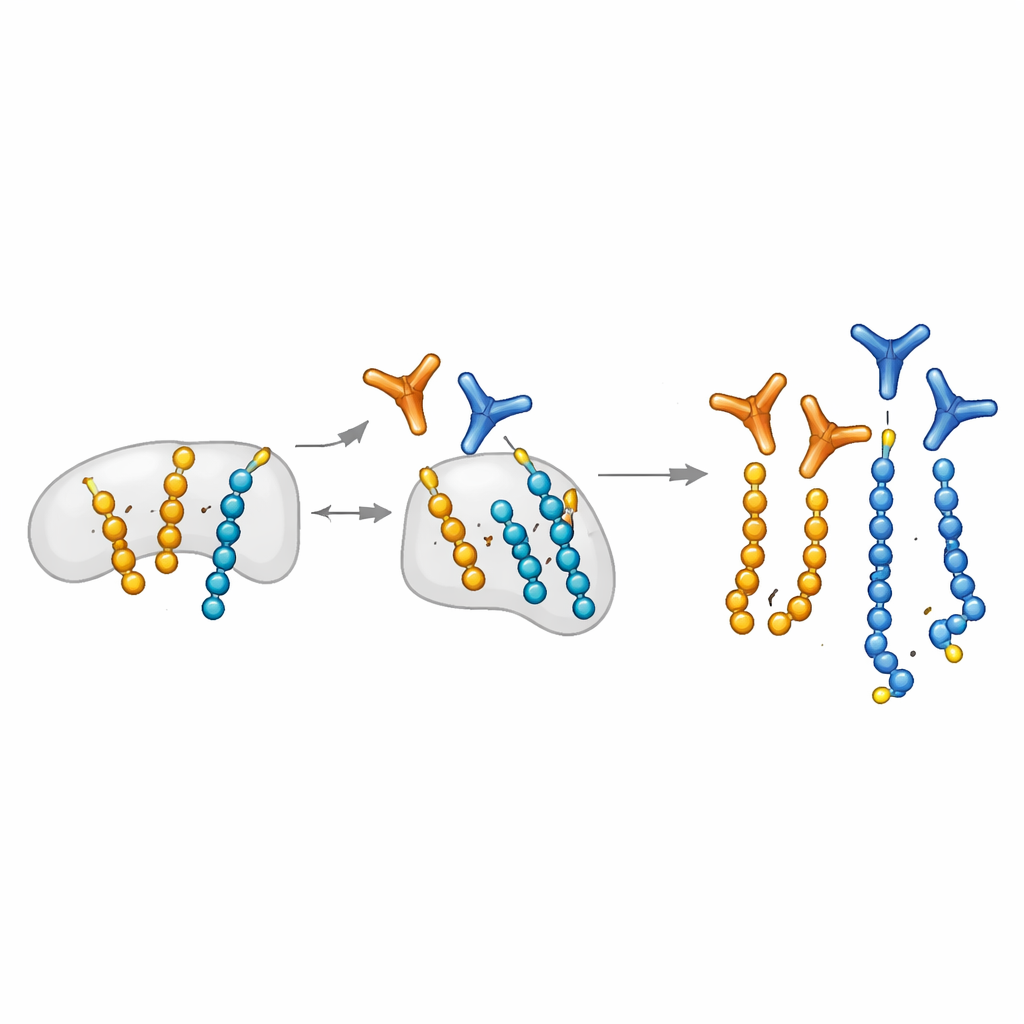
The authors expand a chemical biology platform that can programmatically place ADP-ribose on specific serine residues of short peptides. They use these custom-made peptides as bait to fish out antibody fragments from large display libraries. Crucially, they introduce a "mono-blocking" step: mono-modified peptides are used to soak up antibodies that recognize single units, leaving behind candidates that prefer long chains. This strategy yields modular antibodies that, once assembled using a SpyTag/SpyCatcher system, can be rapidly converted into different formats for blots, microscopy, and pull-downs. The team also discovers a distinctive pattern of fragment ions in mass spectrometry that specifically marks peptides carrying two ADP-ribose units, helping to correctly map where poly-modifications occur on proteins.
Seeing subtle signals where others see nothing
With their new tools, the researchers show that the poly-specific antibodies detect very short chains, long chains, and even free ADP-ribose polymers, and they do so with far greater sensitivity than existing commercial reagents. The antibodies work across different amino-acid linkages, not just serine, and can visualize poly-ADP-ribose in cells under conditions where previous tools saw no signal at all, such as in unstressed cells or cells growing in low oxygen. In parallel, the team affinity-matures earlier mono-specific and site-specific antibodies to boost their sensitivity while keeping their selectivity. Together, this toolbox allows simultaneous imaging of both mono and poly signals in the same cell, revealing that poly-modification appears in localized nuclear foci while mono-modification forms a more diffuse and longer-lasting wave across chromatin.
How losing XRCC1 reshapes the DNA repair waves
XRCC1 normally acts as a scaffold that helps PARP1 engage and then disengage from DNA repair intermediates. When XRCC1 is lost, PARP1 becomes hyperactive and has been known to overproduce long ADP-ribose chains, a state linked to neurodegeneration. Using their refined antibody set, the authors show that XRCC1 loss does more than boost long chains: it dramatically amplifies the mono-ADP-ribosylation wave as well. During ongoing DNA damage from different agents, cells lacking XRCC1 show higher and earlier peaks of both long and short modifications, with damage-type-specific timing. Mono-modification often remains more stable and homogeneously distributed than the rapidly fluctuating poly signal. The work also identifies histone H1 as a prominent mono-modified target whose persistent tagging may keep chromatin more relaxed around lesions.
From altered signals to altered protein traffic
Mono-ADP-ribose on serine can itself be further decorated by an unusual chemical linkage to ubiquitin, forming a hybrid mark called ADP-ribosyl-ubiquitylation. The ubiquitin ligase RNF114 recognizes this composite signal and is drawn to DNA damage sites where it may modify or move other proteins. Using a specialized reader protein (ZUD) alongside their antibodies, the authors show that XRCC1 deficiency increases not just mono-ADP-ribose, but also this hybrid mark and the recruitment of RNF114 to breaks. This suggests that misregulated mono-signaling and its ubiquitin crosstalk may contribute to the cellular stress and disease phenotypes seen when XRCC1 is mutated.
What this means for understanding and treating disease
To a non-specialist, the key message is that what once looked like a single DNA repair signal is really two overlapping waves of small chemical tags, each with its own timing and consequences. By building highly selective molecular tools, this study separates those waves and shows that losing XRCC1 intensifies not only the well-known burst of long chains but also a sustained surge of single-unit tags and their linked ubiquitin marks. These insights refine how we think PARP1 works, help explain why defects in XRCC1 and related enzymes cause brain disease, and provide a toolkit that other researchers can use to dissect ADP-ribosylation in cancer, immunity, and beyond—potentially leading to more precise use of PARP-targeting drugs.
Citation: Dauben, H., Mihaljević, M., Kolvenbach, A. et al. Versatile and sensitive detection of mono- and poly(ADP-ribosyl)ation reveals XRCC1-dependent remodelling of PARP1 signalling. Nat Commun 17, 3216 (2026). https://doi.org/10.1038/s41467-026-71311-4
Keywords: ADP-ribosylation, DNA damage repair, PARP1 signaling, XRCC1 deficiency, ubiquitylation