Clear Sky Science · en
Proteomic characterization of intrahepatic cholangiocarcinoma identifies risk-stratifying subgroups and EIF4A1 as a therapeutic target
Why this liver cancer study matters
Intrahepatic cholangiocarcinoma is a rare but increasingly common form of bile duct cancer that arises inside the liver. It is usually discovered late, often returns even after surgery, and doctors currently have few reliable ways to predict which patients will relapse or benefit from extra treatment. This study uses state-of-the-art protein analysis to sort these tumors into biologically distinct groups and to uncover a shared weak point that might be targeted with a new drug.
Looking closely at proteins in liver tumors
The researchers examined tumor samples and nearby noncancerous liver tissue from two groups of patients treated at major medical centers in the United States and Germany. Instead of focusing on DNA or RNA, they measured thousands of proteins in each sample using advanced mass spectrometry, a technology that can quantify many molecules at once. By comparing tumors to surrounding tissue, they saw broad rewiring of basic cell functions: healthy liver areas were richer in metabolism-related proteins, while tumors were packed with proteins tied to cell division, DNA handling, and the fibrous material that surrounds cells. These differences confirmed that the protein-level view captures features of the disease that cannot be seen from gene data alone.
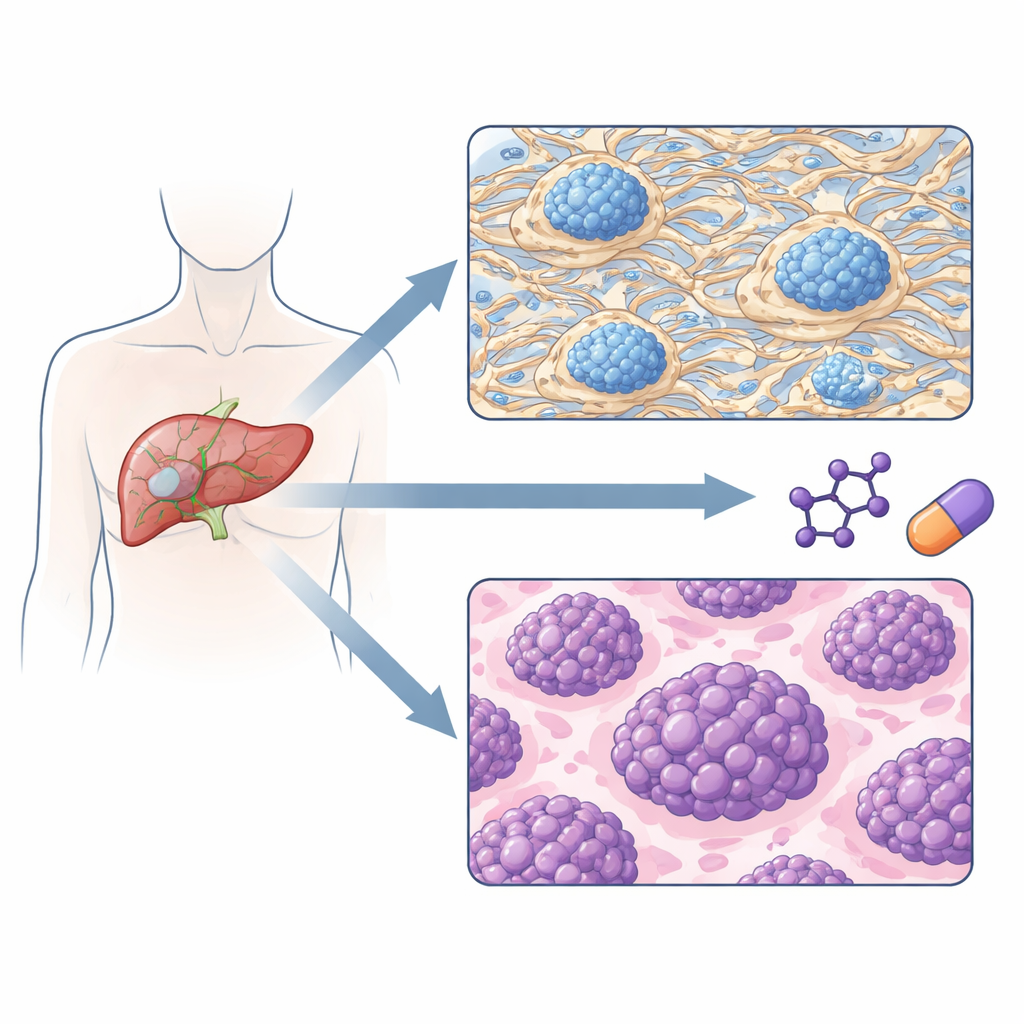
Two tumor types with different risks of coming back
When the team grouped tumors purely by their protein patterns, two main subtypes emerged. One subtype was dominated by proteins that build and remodel the extracellular matrix—the scaffolding between cells—along with components of blood clotting and certain immune signals. The other subtype was dominated by proteins that drive DNA replication, RNA processing, and protein synthesis, all hallmarks of fast-growing cells. Importantly, these two protein-defined groups had very different clinical courses: patients in the "matrix-rich" group tended to have earlier-stage tumors and went much longer before the cancer returned, whereas patients in the "proliferation-heavy" group relapsed sooner and had shorter overall survival.
A simple test that works across patient groups
To turn these insights into something closer to a practical tool, the researchers used machine learning to build a classifier based on just four proteins that best distinguished the two subtypes. They trained and tested many four-protein combinations on their main patient set and found that most could accurately separate the subgroups. Using the best-performing set, they then applied this classifier to two independent collections of intrahepatic cholangiocarcinoma from other hospitals, including a large Chinese cohort previously characterized at the gene and protein levels. In both outside groups, the same two proteomic patterns reappeared, and patients assigned to the matrix-rich group again fared better than those in the proliferation-heavy group. This suggests that a small protein panel could help doctors estimate relapse risk across different populations.
Finding a common drug target in both subtypes
Beyond risk prediction, the scientists looked for proteins that were consistently elevated in tumors from both groups and that might be amenable to therapy. One promising candidate was EIF4A1, a key component of the cellular machinery that starts translating RNA messages into proteins and is linked to cancer-promoting pathways. EIF4A1 was strongly increased in tumors compared with nearby liver tissue, regardless of subtype. The team tested Zotatifin, an experimental drug that blocks EIF4A1, in cell lines derived from this cancer and found that low doses reduced their viability, especially when combined with another pathway inhibitor.
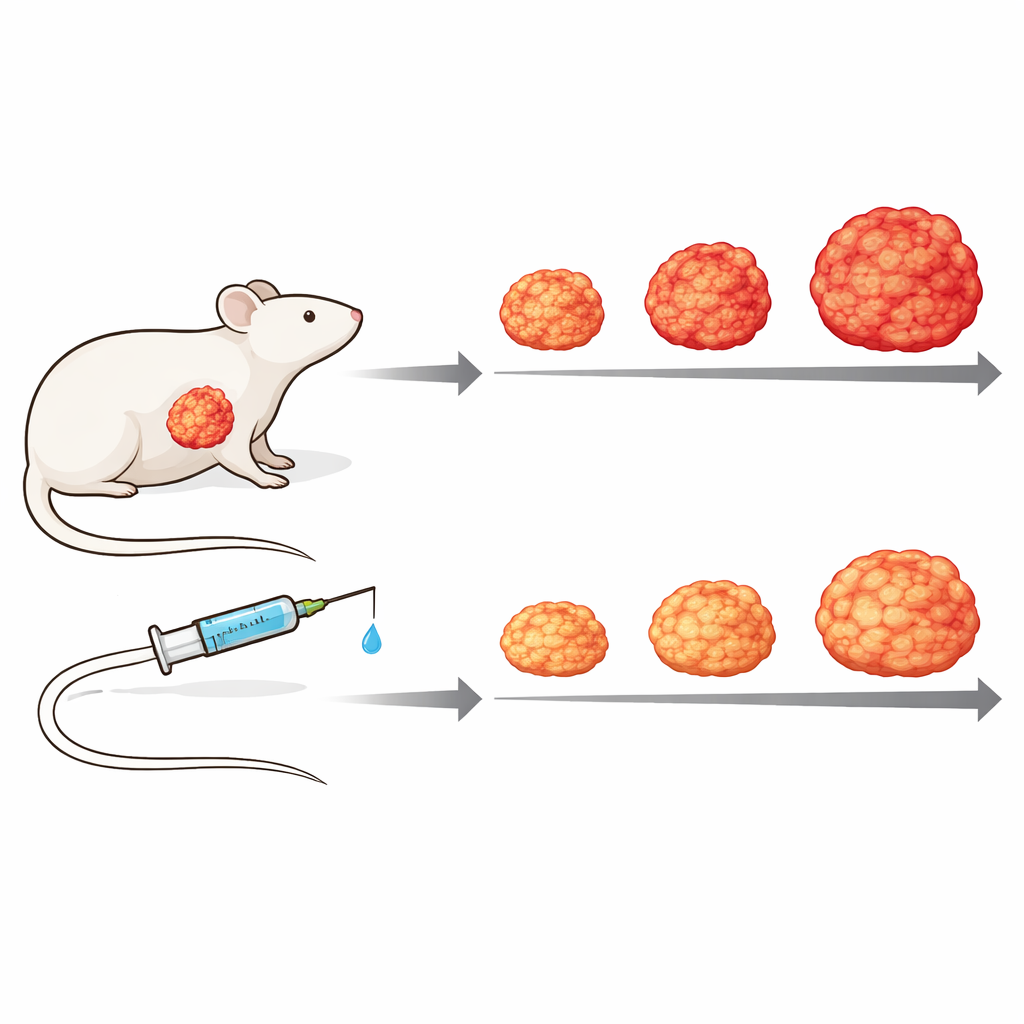
Testing the drug in a living tumor model
To mimic the complexity of human disease more closely, the researchers moved to patient-derived xenograft models, in which pieces of a person’s tumor are implanted into immune-deficient mice. Using protein measurements that could distinguish human tumor proteins from mouse stromal proteins, they showed that the surrounding connective tissue responds differently depending on the tumor’s internal activity, underlining the importance of tumor–stroma crosstalk. In one such model, weekly treatment with Zotatifin slowed tumor growth substantially compared with untreated mice, without obvious toxicity: tumors grew more slowly and stayed smaller, and treated animals maintained their body weight better over time.
What this means for patients
In plain terms, this work shows that intrahepatic cholangiocarcinoma is not a single disease but at least two main protein-defined types with different chances of coming back after surgery. A focused four-protein test could help doctors identify higher-risk patients who might benefit from more intensive follow-up or additional therapy. At the same time, the discovery that both subtypes rely on the translation factor EIF4A1, and that blocking it with Zotatifin can curb tumor growth in realistic models, points to a potential new treatment strategy. While clinical trials will be needed to confirm safety and benefit in people, this study demonstrates how deep protein profiling can both refine prognosis and uncover actionable drug targets in a hard-to-treat cancer.
Citation: Werner, T., Thiery, J., Budau, KL. et al. Proteomic characterization of intrahepatic cholangiocarcinoma identifies risk-stratifying subgroups and EIF4A1 as a therapeutic target. Nat Commun 17, 2741 (2026). https://doi.org/10.1038/s41467-026-70817-1
Keywords: intrahepatic cholangiocarcinoma, cancer proteomics, tumor subtypes, EIF4A1 inhibition, patient-derived xenograft