Clear Sky Science · en
Mining antibody functionality via AI-guided structural landscape profiling
Why this research matters
Vaccines and antibody drugs have been central weapons against COVID-19, but our immune systems generate an almost unimaginably large variety of antibodies, most of which we never study in detail. This paper shows how artificial intelligence can read antibody genetic sequences and infer where and how they latch onto the coronavirus spike, helping scientists rapidly pick out the rare molecules that both neutralize the virus and stay active as it mutates. That same strategy could speed the search for antibody medicines against many future pathogens.
Turning antibody sequences into a map
The authors focus on antibodies that recognize the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein, a key handle the virus uses to grab human cells. They assembled a large training set from publicly available structures of antibody–RBD complexes and from deep mutational scanning experiments that chart how changes in the virus affect antibody binding. By clustering these data, they defined 12 distinct “epitope classes” – recurring regions on the RBD that different antibodies target, each associated with characteristic strength and breadth of virus neutralization.
Building an AI guide to antibody function
On top of this structural map, the team built an AI tool called RBD-AIM. First, a language model trained specifically on antibody sequences analyzes the genetic code of an antibody’s heavy and light chains and predicts which epitope class it belongs to. Then a structure-prediction step uses known 3D templates and modern folding algorithms to refine how that antibody is likely to sit on the RBD. A confidence score derived from the predicted 3D alignment helps decide when to trust the structural model versus the simpler sequence-based classifier. Overall, this combined approach outperforms traditional homology searches and several state-of-the-art structure predictors at guessing where an antibody binds and how well it is likely to block the virus.
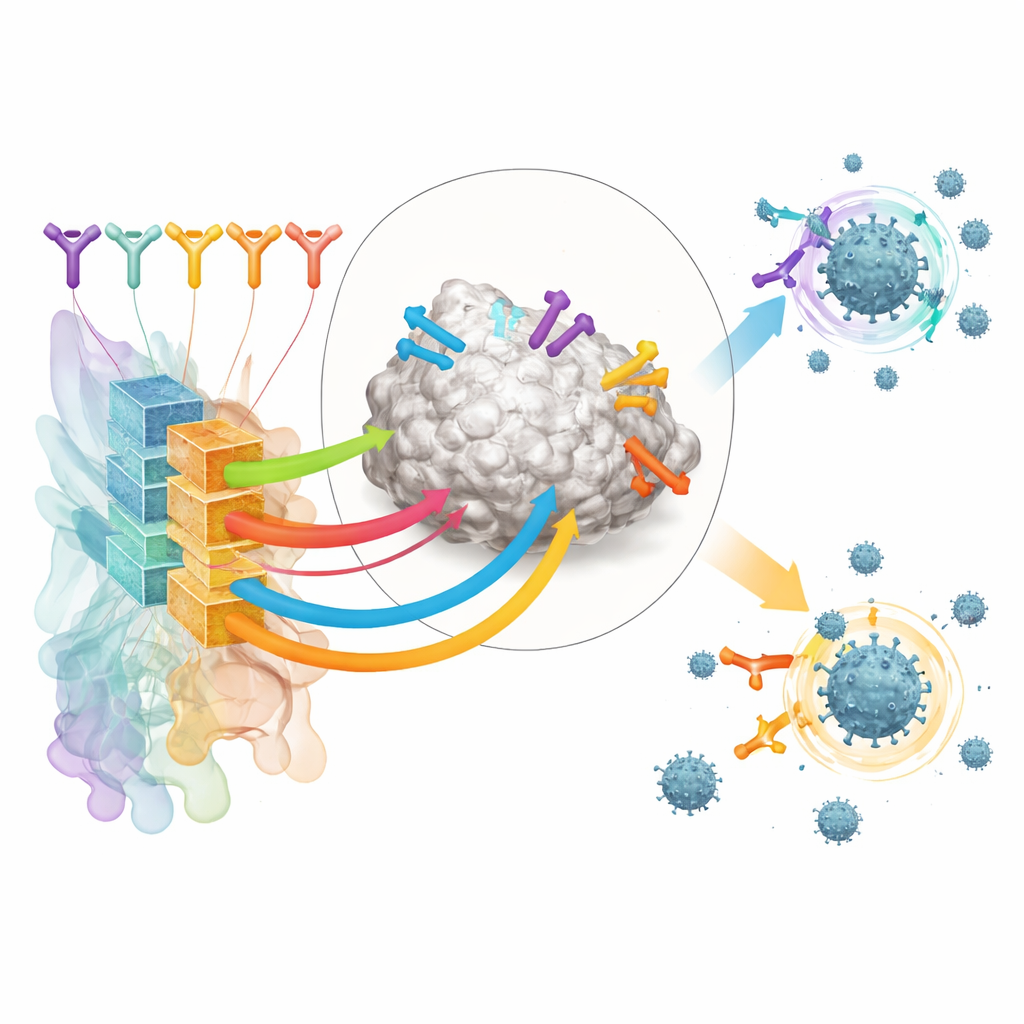
Recreating real immune responses in the lab
To test RBD-AIM on realistic antibody collections, the researchers reconstructed native B cell repertoires from people vaccinated with either an mRNA vaccine (Pfizer–BioNTech) or an adenoviral vector vaccine (Sputnik V). Using droplet microfluidics, they captured the original pairings of heavy and light antibody chains from single B cells and displayed the resulting fragments on the surface of yeast cells. This preserved the natural diversity of antibodies far better than conventional methods that randomly mix chains. Fluorescently labeled RBD allowed them to sort out yeast cells bearing high-affinity RBD binders, including those that compete directly with the human ACE2 receptor for the same patch on the spike.
Predicting and validating which antibodies really work
Dozens of strongly enriched RBD-binding antibodies from these vaccinated donors were then run through RBD-AIM. The tool predicted which epitope class each one recognized, whether it likely blocked ACE2, and how robust it might be to spike mutations found in variants such as Delta and Omicron. As expected, antibodies aiming at the ACE2 contact area tended to be potent but narrow in scope, losing activity against Omicron. In contrast, a subset of antibodies from a more distantly located epitope class (dubbed E221 by the authors) was forecast to retain neutralizing power across variants. Laboratory binding and virus neutralization tests confirmed these patterns, and crystal structures showed that RBD-AIM’s structural guesses were often accurate down to the detailed contact surfaces.
From prediction to protection in animals
To see whether these AI-prioritized antibodies could act as real medicines, the team selected representative molecules: one classic ACE2-blocking antibody and one E221-class antibody that neutralizes by a more indirect mechanism. In a lethal SARS-CoV-2 challenge model using mice engineered to express human ACE2, a single low dose of either antibody protected all treated animals, aligning with their predicted neutralizing profiles. Further structural and simulation work suggested that the broadly active E221-class antibody works by “freezing” flexible parts of the RBD that are needed for efficient receptor engagement, a mechanism less vulnerable to the exact shape of the ACE2-contact patch.
What this means for future pandemics
In practical terms, this study delivers a working blueprint for combining high-throughput display technologies with AI to turn raw antibody sequence data into functional insight. Instead of painstakingly characterizing one antibody at a time, researchers can now quickly scan entire repertoires, rank promising candidates by potency and breadth, and understand which regions of a viral protein are most important to target. While RBD-AIM was trained on SARS-CoV-2, the authors show that with sufficient structural and mutational data the same strategy can extend to other viruses, pointing toward a future in which AI-guided “structural landscape profiling” helps us mine our own immune responses for next-generation antibody therapies.
Citation: Terekhov, S.S., Ivanisenko, N.V., Zhang, N. et al. Mining antibody functionality via AI-guided structural landscape profiling. Nat Commun 17, 4009 (2026). https://doi.org/10.1038/s41467-026-70553-6
Keywords: antibody discovery, artificial intelligence, SARS-CoV-2 spike, epitope mapping, therapeutic antibodies