Clear Sky Science · en
SFPQ directs histone H3.3 deposition to R-loops in DNA repeats to protect genome stability
Guardians of Our DNA Repeats
Much of our DNA is made of long, repeated sequences once dismissed as “junk.” We now know these regions can misbehave, forming unusual three-stranded knots called R-loops that stall copying of DNA and trigger damage. This study uncovers how a protein named SFPQ patrols these repetitive stretches, organizes their packaging, and in doing so quietly protects genome stability and even shapes how certain cancers interact with the immune system.
Hidden Knots in the Genome
When a gene is read, a temporary RNA copy is made from the DNA. In some places, especially in repetitive DNA such as telomeres at chromosome ends, centromeres near the middle, and mobile elements like LINEs and SINEs, the new RNA can fold back and hybridize to the DNA. This creates an R-loop: a short RNA–DNA hybrid paired with a displaced strand of single DNA. While R-loops can be useful in small, controlled doses, persistent ones act like roadblocks for the machinery that copies DNA, leading to broken chromosomes, faulty cell division, and genome instability—key hallmarks of cancer and genetic disease.
The R-Loop Patrol Protein
The researchers show that SFPQ, long known as an RNA-binding and splicing factor, is also a dedicated sensor of R-loops at repetitive regions. In cells where SFPQ was reduced, the team detected more R-loops across telomeres, peri-centromeric satellites, and retroelements using hybrid-specific antibodies and sequencing-based mapping. These hotspots coincided with strong signals of replication stress and DNA damage. Biochemical experiments with purified SFPQ revealed why: SFPQ does not bind ordinary double-stranded DNA or RNA–DNA duplexes well, but it strongly prefers three-stranded structures, especially R-loops that carry exposed RNA tails similar to natural repeat RNAs. This selectivity positions SFPQ as a first responder that recognizes problematic R-loops precisely where the genome is most repetitive.
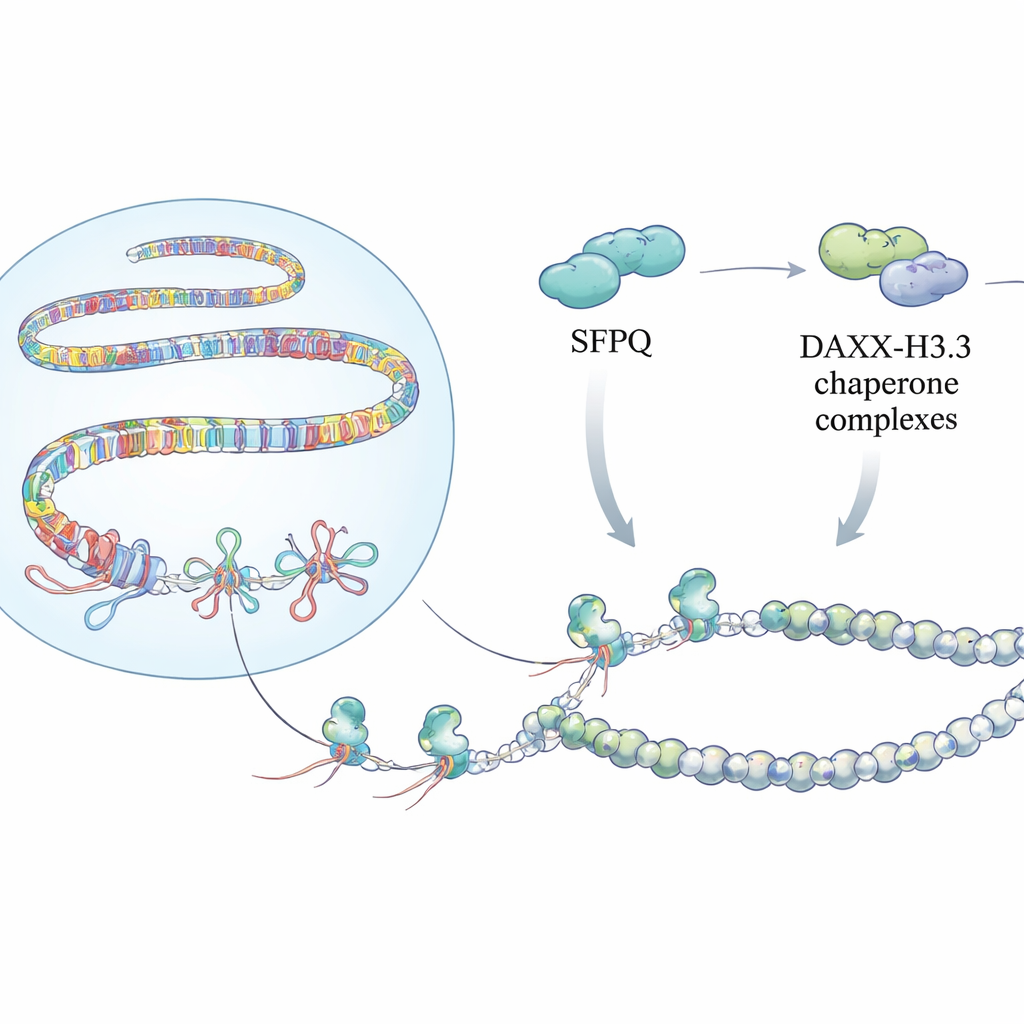
Building Protective Chromatin Armor
Detecting these knots is only part of the story. The team discovered that SFPQ physically interacts with DAXX, a protein that delivers a special histone variant called H3.3 into chromatin. H3.3 helps form stable nucleosomes that silence repetitive DNA and prevent uncontrolled recombination. Mapping experiments across the genome showed that SFPQ and DAXX frequently share binding sites in intergenic and intronic repeats, rather than at classic gene promoters. When SFPQ was lost, DAXX shifted away from repeats toward regulatory regions, and H3.3 was no longer efficiently deposited at repeat-rich stretches, even though overall H3.3 protein levels were unchanged. As a result, the chromatin template at these repeats became poorly organized, allowing R-loops and replication stress to build up.
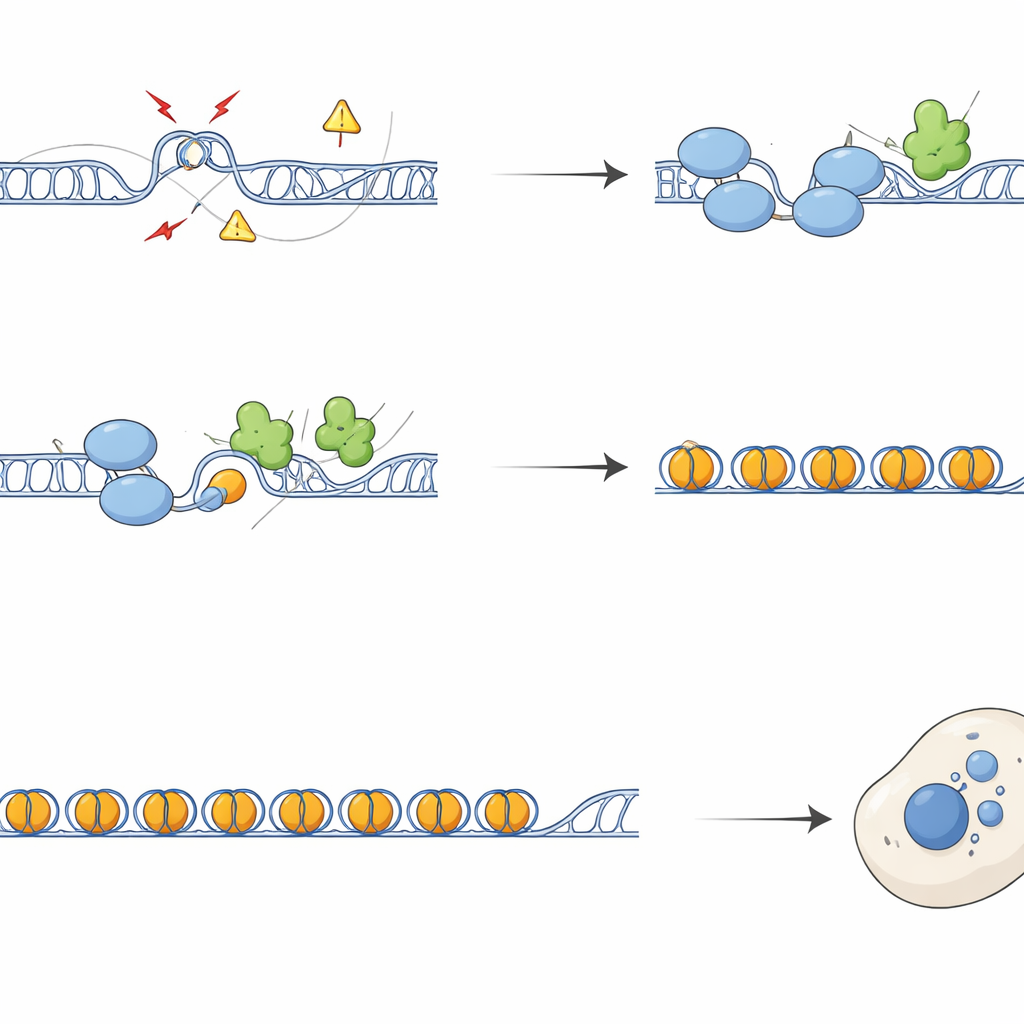
From Broken Chromosomes to Alarmed Immunity
Without SFPQ, repetitive regions accumulated DNA damage markers and repair factors, and chromosomes showed more breaks and exchanges between sister strands. During cell division, this damage translated into defective segregation, forming chromatin bridges and micronuclei—tiny extra DNA-containing bodies outside the main nucleus. These micronuclei frequently bore telomeric and centromeric sequences and were coated with cGAS, a sensor of misplaced DNA that triggers the cGAS–STING innate immune pathway. Cells lacking SFPQ or DAXX activated interferon and inflammatory genes, an effect that could be largely reversed by overexpressing RNaseH1, an enzyme that removes R-loops, or by blocking STING. Thus, excess R-loops at repeats feed directly into immune signaling through the production of cytoplasmic DNA debris.
Implications for Cancer and Patient Outcomes
In sarcoma patient data, high SFPQ expression correlated with poorer survival, whereas a gene signature reflecting strong innate immune activation—similar to that seen when SFPQ is lost—was linked to better outcomes. Patients with low SFPQ but high immune-signature expression fared best, suggesting that, in tumors, disabling this protective axis at repeats may inadvertently expose cancer cells to the immune system. The work outlines a mechanistic chain from R-loop sensing by SFPQ, to DAXX–H3.3–mediated chromatin protection, to prevention of genome instability and dampening of innate immunity. For a lay audience, the core message is that SFPQ acts like a maintenance crew for repetitive DNA: it detects dangerous RNA–DNA knots, calls in packaging helpers to smooth and armor these regions, and in doing so keeps chromosomes intact and immune alarms quiet—a balance that cancers may exploit but that future therapies could potentially tip back in favor of the patient.
Citation: Ferrando, A., Giaquinto, M., Napolitano, L.M.R. et al. SFPQ directs histone H3.3 deposition to R-loops in DNA repeats to protect genome stability. Nat Commun 17, 3151 (2026). https://doi.org/10.1038/s41467-026-69479-w
Keywords: R-loops, genome stability, histone H3.3, innate immunity, sarcoma