Clear Sky Science · en
Combining xQTL and genome-wide association studies from diverse populations improves druggable gene discovery
Why genes matter for future medicines
Modern genetics has uncovered thousands of DNA markers linked to common illnesses, but turning those clues into real drug targets has been frustratingly slow. This study tackles a central bottleneck: our standard ways of reading large genetic studies often miss the genes that medicines can actually act on. By rethinking how to analyze genetic data across diverse populations and layers of biology, the authors aim to uncover hidden, but druggable, weak spots in diseases such as Alzheimer’s, depression, schizophrenia, and type 2 diabetes.
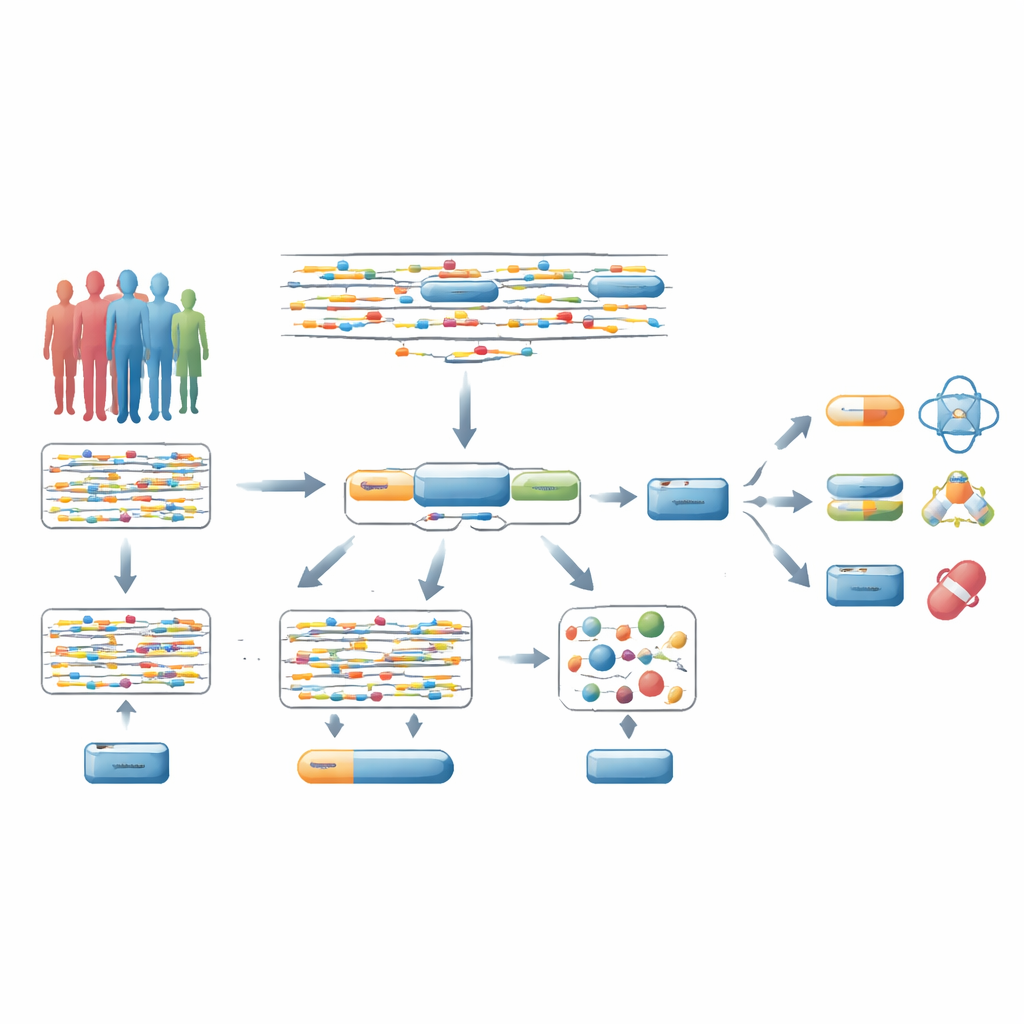
From scattered DNA signals to whole genes
Most genome-wide association studies (GWAS) look at millions of single DNA changes one by one and ask whether each is linked to a disease. Typically, researchers then assume that the most strongly associated DNA change near a gene points to that gene. But this shortcut is often wrong: only a small fraction of these “lead” variants truly cause disease, and they can influence distant genes. Because of this, strict statistical corrections spread across millions of tests, making it hard to find real disease genes, especially those that existing or potential drugs can target.
A new set of tools to read the genetic map
The authors introduce a family of “gene-based” tests, led by a method called GenT. Instead of judging single DNA changes in isolation, GenT groups all relevant variants around each gene and tests them together. Crucially, the team worked out the underlying mathematics so they know exactly how GenT should behave when there is no real signal, avoiding the false alarms that can plague earlier gene-based approaches. Extensions of this framework—MuGenT for multi-ancestry or multi-trait data, and xGenT for integrating molecular readouts like gene expression or protein levels—let researchers combine information across populations and layers of biology while still working from summary statistics that are widely available.
Finding hidden drug targets in brain and body
Applying GenT to large genetic studies of Alzheimer’s disease, amyotrophic lateral sclerosis, major depression, and schizophrenia, the authors analyzed over 18,000 genes. They uncovered 415 genes whose protein products can be targeted by existing or experimental compounds, many of which standard GWAS methods had missed. Dozens of these druggable genes sit in regions of the genome that showed no strong single-variant signal, suggesting that the risk is spread over many modest effects that only become visible when viewed at the gene level. Notable examples include SYK for Alzheimer’s disease and THRB for schizophrenia, both supported by prior biological work yet previously below the radar of conventional analyses.
The power of diversity and molecular layers
MuGenT extends this strategy to combine genetic data from people of European, African, East Asian, South Asian, and Hispanic ancestries with type 2 diabetes. Because many risk variants are shared but vary in frequency and strength across groups, pooling them at the gene level greatly boosts statistical power. MuGenT identified 269 diabetes-associated genes lacking any individually significant DNA variants, 45 of which encode druggable proteins in previously unsuspected regions. To connect genes to actual biology, the xGenT method overlays disease genetics with measurements of how variants affect gene activity and protein abundance in brain tissues. In Alzheimer’s disease, xGenT highlighted 26 druggable genes outside known risk regions, including NTRK1 and RIPK2, and pointed to specific brain regions and cell types where they are active.
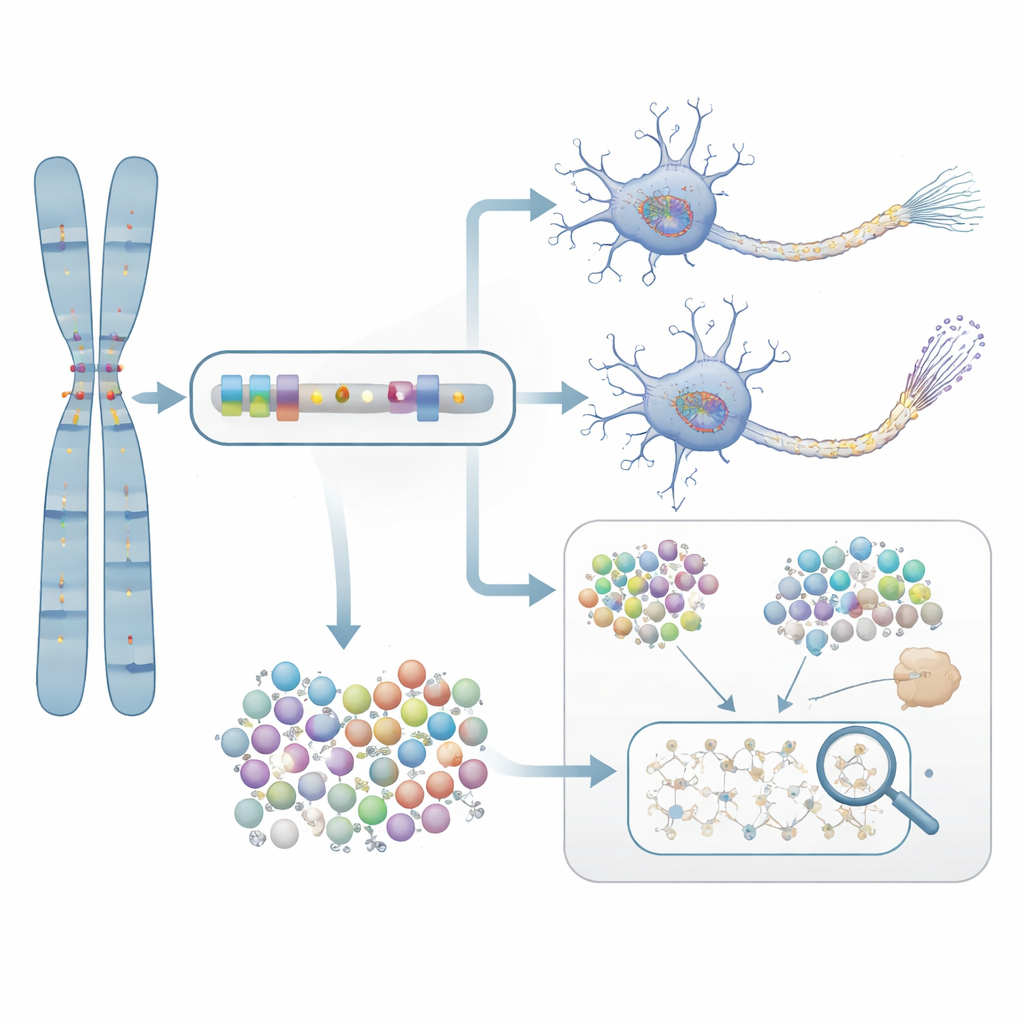
From prediction to lab evidence
To test whether these statistically flagged genes truly matter for disease, the researchers zoomed in on NTRK1, a gene involved in nerve cell survival. They treated neurons grown from the cells of Alzheimer’s patients with a selective NTRK1-blocking compound. In these lab-grown neurons, inhibiting NTRK1 reduced abnormal chemical tags on tau protein—changes closely linked to the tangled protein clumps seen in Alzheimer’s brains. This experiment provides a mechanistic bridge from the gene-based statistics to a biological process that can, in principle, be targeted by drugs.
What this means for future treatments
Viewed together, the work shows that shifting focus from single DNA markers to whole genes, while drawing on data from diverse populations and multiple molecular layers, can reveal many more plausible drug targets for complex diseases. The authors argue that many truly important genes influence disease through networks of modest, distributed effects that are poorly captured by traditional GWAS scans. Their GenT framework, along with its multi-ancestry and molecular extensions, offers a statistically rigorous way to expose these hidden signals and prioritize genes for deeper experimental testing. While further laboratory and clinical studies will be needed before any therapy reaches patients, this approach significantly expands the genetic starting points for drug discovery in conditions that urgently need better treatments.
Citation: Lorincz-Comi, N., Song, W., Chen, X. et al. Combining xQTL and genome-wide association studies from diverse populations improves druggable gene discovery. Nat Commun 17, 2801 (2026). https://doi.org/10.1038/s41467-026-69236-z
Keywords: gene-based association, Alzheimer’s disease, type 2 diabetes, druggable targets, multi-ancestry genetics