Clear Sky Science · en
Decoding the role of chromatin context in the off-target effects of CRISPR gene editing with EGOLD
Why tiny DNA tweaks can have big side effects
CRISPR gene editing has raised hopes of curing many genetic diseases by precisely changing DNA inside our cells. But the same tools that can fix a faulty gene can also accidentally alter other parts of the genome, creating "off-target" changes. This study introduces a new way to map those unintended edits directly in human cells and reveals that how tightly DNA is packed inside the cell plays a major role in where off-target changes occur.
Looking beyond the DNA letters
Most efforts to understand CRISPR safety focus on the DNA sequence itself: which strings of A, C, G, and T are most likely to be cut or changed. Yet DNA in living cells is wrapped around proteins and folded into more open or closed states, a landscape known as chromatin. Open regions are easier for cellular machines to reach, while closed regions are more shielded. The authors suspected that this local DNA packaging could quietly steer where CRISPR tools make their mistakes, but existing tests did not capture enough real genomic sites to study this effect in depth.
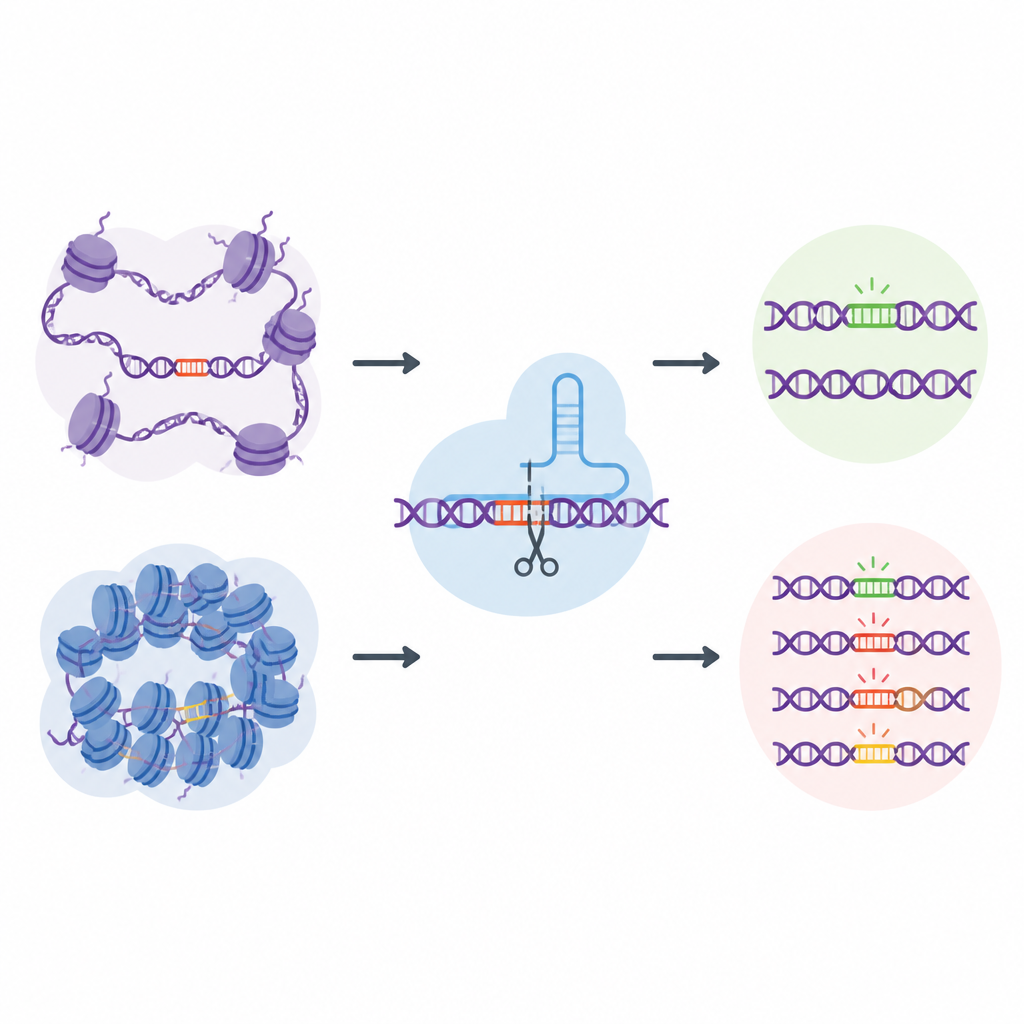
A genome-wide "natural" off-target library
To tackle this challenge, the researchers created EGOLD, short for Endogenous Genome-wide Off-target Library Detection. Instead of building artificial DNA libraries, they took advantage of repeated or very similar sequences already scattered throughout the human genome. They chose special target sites where a single CRISPR guide RNA closely matches thousands of other locations that share the same sequence but sit in very different chromatin environments. By editing one chosen site and then reading the entire genome, EGOLD can compare which of these many look-alike sites are edited and which are left untouched, all while their underlying DNA letters remain identical.
Open DNA is more vulnerable to stray edits
Using EGOLD, the team tested 17 CRISPR-based editors, including standard DNA-cutting Cas9 proteins and newer base editors that change single letters without cutting both DNA strands. Across more than two million detected off-target events, a clear pattern emerged: sites in open chromatin were far more likely to be edited than the same sequence buried in closed, compact regions. Signals of open DNA, such as accessibility to DNA-cutting enzymes and active chemical marks on nearby proteins, were positively linked to off-target activity. In contrast, features associated with tightly packed or chemically silenced DNA tended to protect against unwanted changes.
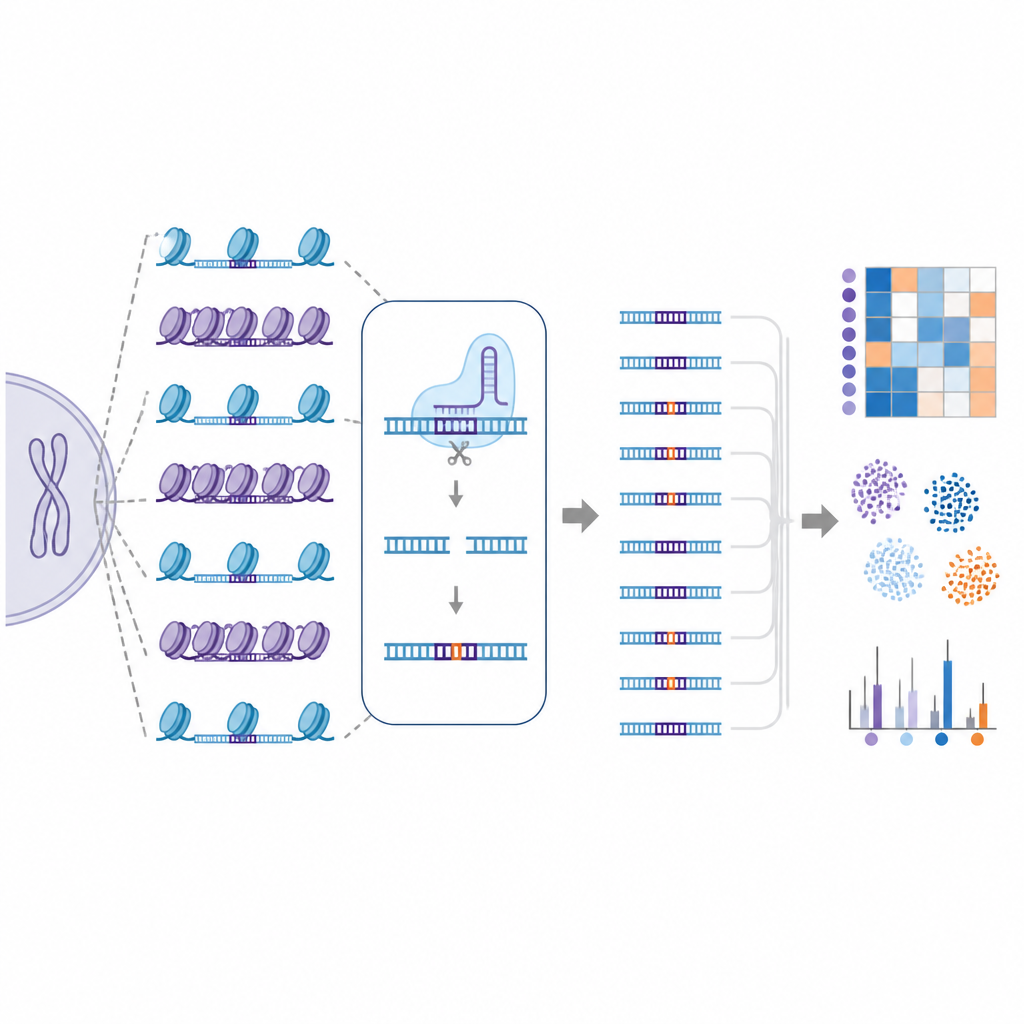
Not all editors and cells behave the same
The study also compared different Cas9 variants engineered to be either more accurate or more flexible in what they recognize. One variant called SuperFiCas9 stood out for keeping good activity at its intended target while producing relatively few off-target changes, especially when built into base editors. Other versions that could tolerate more sequence differences or more relaxed recognition rules showed much higher off-target rates. Even when the same editor and guide RNA were used, different human cell types displayed different off-target patterns, highlighting how cell-specific chromatin landscapes shape editing outcomes.
Teaching computers to predict risky sites
Because EGOLD generates such large, detailed off-target maps, the authors used these data to train machine learning models. These computer models were fed both sequence information and local chromatin features for each potential site and learned to distinguish edited from unedited locations with very high accuracy. When chromatin information was added to sequence-based features, prediction performance improved, underscoring that DNA packaging is not just a side detail but a key part of off-target risk.
What this means for safer gene editing
For people following the progress of gene therapy, this work shows that where CRISPR goes wrong is shaped not only by DNA spelling but also by how that DNA is folded and marked inside cells. EGOLD provides a practical way to chart those risks directly in real genomes and to build smarter prediction tools that account for chromatin context. In the long run, this approach could help researchers choose safer editing strategies, design better guides, and select more suitable editor versions so that life-changing DNA fixes come with fewer unintended changes elsewhere.
Citation: Feng, H., Zheng, J., Li, N. et al. Decoding the role of chromatin context in the off-target effects of CRISPR gene editing with EGOLD. Cell Discov 12, 32 (2026). https://doi.org/10.1038/s41421-026-00889-2
Keywords: CRISPR off-targets, chromatin context, base editing, genome safety, machine learning