Clear Sky Science · en
Molecular mechanism of transition-state inhibitors of bacterial antibiotic efflux pumps
Why stopping bacterial "pumps" could save antibiotics
Antibiotic resistance often arises not because drugs fail to hit their targets, but because bacteria actively pump them back out. This study reveals, in atomic detail, how a newly designed compound jams a major bacterial pump and makes existing antibiotics powerful again. Understanding how this molecular "wedge" works could open the door to combination therapies that revive medicines we are in danger of losing.
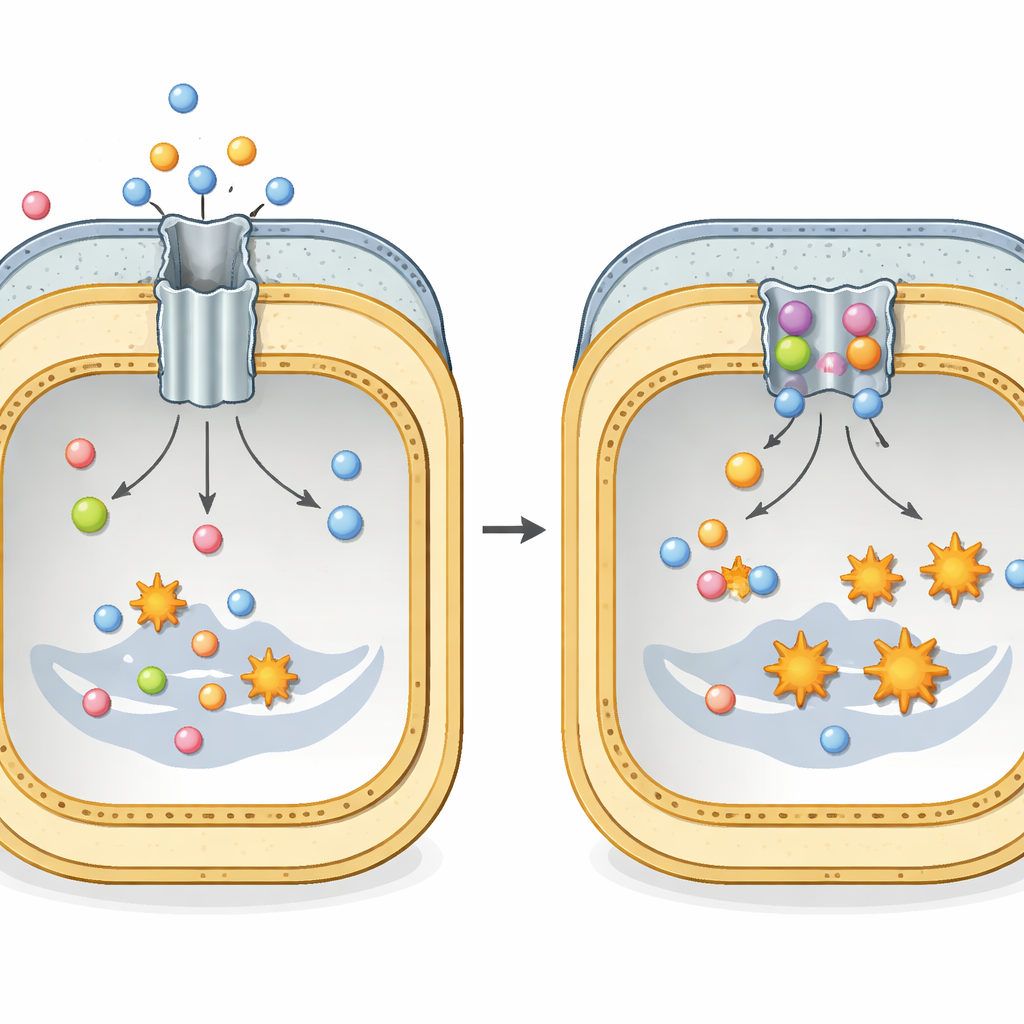
A hidden escape tunnel for antibiotics
Many dangerous Gram-negative bacteria, such as Escherichia coli and Klebsiella pneumoniae, rely on a large protein assembly called AcrAB–TolC to survive antibiotic treatment. This assembly spans the entire cell envelope and works like a tunnel with a motor at its inner end. The inner membrane part, called AcrB, recognizes a wide variety of drugs and other toxic molecules and uses cellular energy to eject them to the outside. Each AcrB unit is built from three repeating parts that continually cycle through three shapes—loosely bound, tightly bound, and open to the outside—creating a peristaltic motion that pushes antibiotics out before they can do lasting damage.
Designing a smarter molecular wedge
Drug developers have long tried to block this pump, but most earlier inhibitors bound in a large greasy pocket that also attracts many other molecules, leading to poor solubility and side effects. Building on previous work, the authors focused instead on a newer family of compounds called pyridylpiperazines that lodge in a more specialized region deep in the membrane-spanning part of AcrB. Guided by high-resolution structural data, they redesigned the chemical scaffold to make stronger contacts with two negatively charged amino acids (D408 and E947) located along the path where the pump normally moves protons. By adding a carefully positioned benzylamine side chain, they created a new compound, BDM91531, that binds about fifty times more tightly than its predecessor and boosts the activity of multiple antibiotics at very low concentrations.
Freezing the pump mid-stroke
X-ray crystallography and single-particle cryo–electron microscopy showed exactly how BDM91531 disables the pump. The compound sits in an elongated cavity between several membrane-spanning helices inside one AcrB subunit. Because BDM91531 carries two positive charges, it forms strong salt bridges with D408 deep in the pocket and E947 near the cytoplasmic rim. These two contact points suspend the inhibitor like a bridge between the inner wall of the pump and the cell interior. Locked in place, the compound traps that subunit in a halfway state between the open and loose conformations—the kind of transient shape the pump normally passes through only briefly. Since the three subunits must rotate through their states in a synchronized way, freezing one of them stalls the entire machine and stops drug efflux.
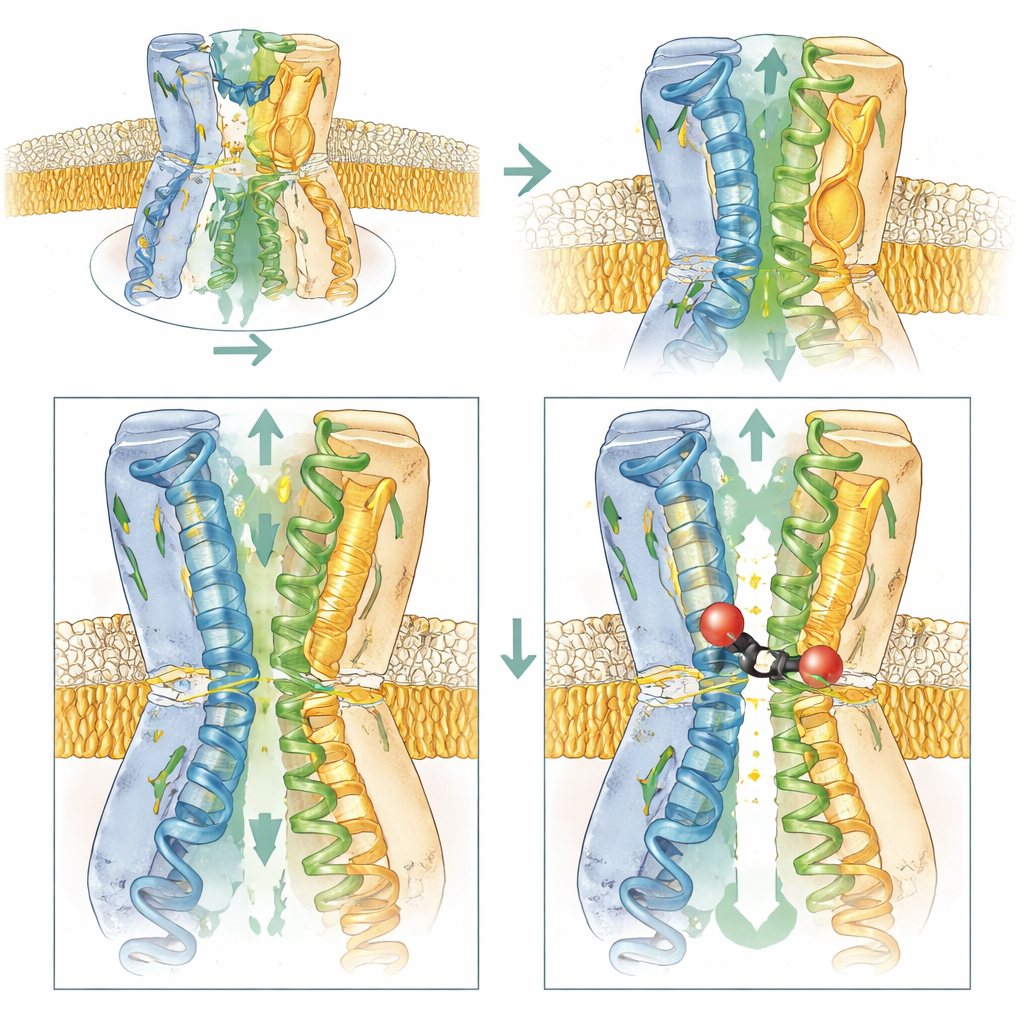
Charging the doorway to pull the inhibitor in
The team combined biophysical measurements, bacterial growth tests, and computer simulations to probe which parts of AcrB are most important for BDM91531’s action. Removing the negative charge at D408 eliminated binding altogether, confirming its central role. Altering E947 or a nearby acidic residue, D951, made bacteria less sensitive to the inhibitor, while reintroducing negative charges at neighboring positions restored or even enhanced sensitivity. This shows that, beyond serving as anchor points, these acidic residues help attract the positively charged inhibitor from the cytoplasm into the entry groove of the binding pocket. In contrast, changes that removed these charges or made the entry zone more neutral reduced both access and binding.
Reviving antibiotics by targeting motion, not the drug
In everyday terms, BDM91531 works by jamming a moving part of a bacterial pump at just the right moment, preventing the machine from completing its cycle and trapping antibiotics inside the cell where they can take effect. Because the inhibitor does not kill bacteria on its own at the doses required to block the pump, it is best viewed as a booster to be combined with existing drugs. The structural and mechanistic insights from this work highlight exactly which features of the pump should be targeted to design even better blockers, raising the prospect of new adjuvant therapies that restore the power of antibiotics against some of our most stubborn infections.
Citation: Börnsen, C., Müller, R.T., Vieira Da Cruz, A. et al. Molecular mechanism of transition-state inhibitors of bacterial antibiotic efflux pumps. npj Antimicrob Resist 4, 35 (2026). https://doi.org/10.1038/s44259-026-00207-6
Keywords: antibiotic resistance, efflux pump inhibitor, AcrB, Gram-negative bacteria, structure-based drug design