Clear Sky Science · en
The role of hepatocyte epigenetics in the pathogenesis of metabolic dysfunction-associated steatotic liver disease
Why this liver story matters to you
Metabolic dysfunction-associated steatotic liver disease, or MASLD, affects nearly one in three people worldwide and is tightly linked to obesity, type 2 diabetes, and heart disease. Most people never feel clear symptoms until serious damage has already occurred. This review article explores a rapidly developing area of research that asks a simple but powerful question: how do chemical "switches" layered on our genes inside liver cells help drive fatty liver disease, and can those switches be reset to prevent or treat it?
From simple fat buildup to serious liver scarring
MASLD is a broad term that covers conditions ranging from mild fat buildup in the liver to inflammation, scarring, cirrhosis, and even liver cancer. The disease develops through a mix of inherited risk, diet, gut microbes, and metabolic problems such as insulin resistance. Classic models describe an initial "hit" where excess calories and hormones push fat into liver cells, followed by additional "hits" like oxidative stress and chronic inflammation that injure and kill those cells. Newer "multiple strikes" thinking adds more layers, including how liver cells communicate with immune cells and scar-forming cells. Among these influences, epigenetics has emerged as a key bridge that links environment and lifestyle to the behavior of liver cells.
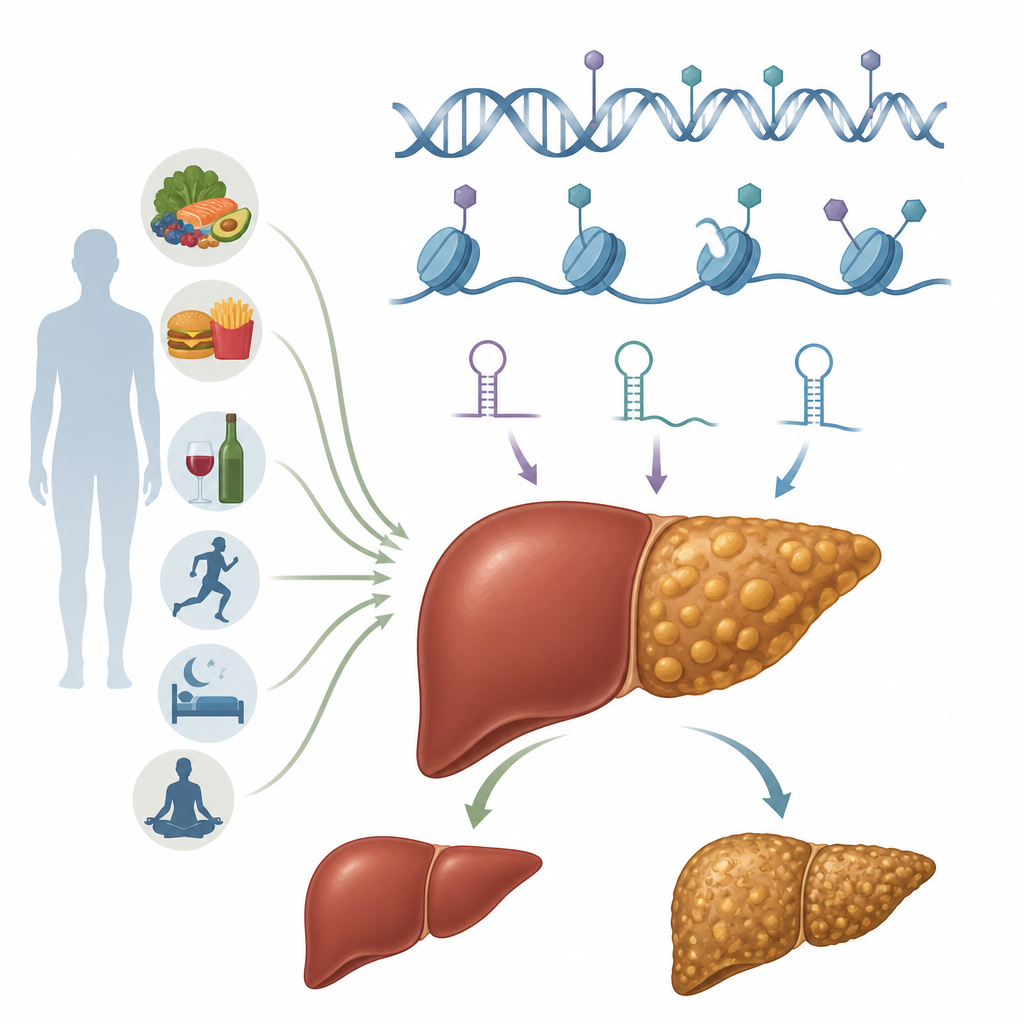
Epigenetic switches in liver cells
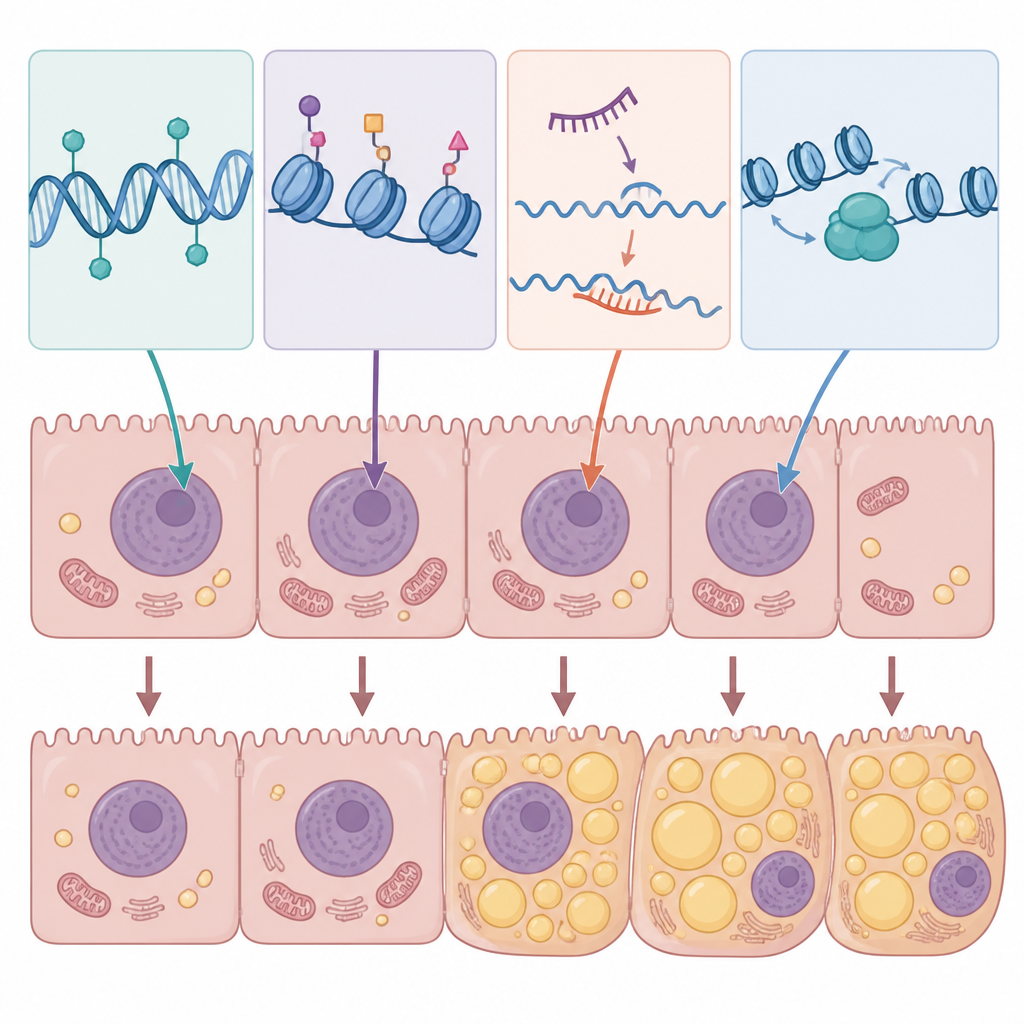
Epigenetics refers to chemical marks and structural changes that adjust how genes are used without altering the DNA code itself. These marks are reversible and can be shaped by diet, metabolic status, and toxins. In hepatocytes, the workhorse cells of the liver, several epigenetic systems work together. DNA methylation can turn genes up or down by placing small chemical tags on specific DNA sites. Histone modification alters the protein spools around which DNA is wrapped, making regions of the genome more open or more closed. Non-coding RNAs, including microRNAs and long RNAs, fine-tune which messages are translated into proteins. Additional layers include chemical changes on RNA itself and large-scale rearrangements of chromatin, the DNA–protein complex that packages genetic material. Together, these systems shape how hepatocytes handle fat, sugar, and inflammatory signals.
How these changes push the liver toward disease
The review gathers evidence that specific epigenetic patterns in hepatocytes favor fat storage, impair fat burning, and fan the flames of inflammation. For example, altered DNA methylation can silence genes that normally support healthy fat breakdown or mitochondrial function, while boosting genes that encourage fat buildup. High sugar and cholesterol levels in experimental models lead to changes in DNA methylation across thousands of genes, shifting entire networks involved in lipid metabolism. Histone-modifying enzymes can open up stretches of DNA that drive fat synthesis or inflammatory pathways, whereas others promote protective programs such as fasting-induced fat burning and cellular cleanup. Non-coding RNAs act as quick-response regulators: certain microRNAs and long RNAs promote fat accumulation and insulin resistance, while others appear to protect against steatosis by restraining lipogenic enzymes or dampening inflammatory signaling.
From mechanisms to tests and treatments
Because epigenetic marks are both stable and reversible, they are attractive as future diagnostic tools and drug targets. Some DNA methylation patterns and microRNA signatures can already be detected in blood and seem to track with the severity of MASLD, raising the possibility of simple blood tests that might spare some patients from liver biopsy. On the treatment side, several medicines and natural compounds tested in cells and animal models, such as resveratrol, betaine, berberine, and certain diabetes drugs, appear to partially improve fatty liver by adjusting epigenetic regulators in hepatocytes. At the same time, the authors caution that these switches control many genes across multiple organs, so broad-acting drugs could cause unwanted side effects if not carefully targeted.
Where this research could lead
The article concludes that epigenetic regulation in hepatocytes is central to how MASLD starts, progresses, and possibly can be reversed. By mapping cell-specific epigenetic changes with modern single-cell and "omics" tools, scientists hope to pinpoint which switches best predict disease risk, which ones mark dangerous progression, and which can be safely manipulated. For non-specialists, the key message is that liver health is not fixed by genes alone. Everyday factors such as diet and metabolic status leave chemical fingerprints on liver cell DNA and RNA that can push the organ toward health or disease. Understanding and eventually guiding these fingerprints may open the door to earlier diagnosis and more precise, less invasive treatments for fatty liver disease.
Citation: Zhu, G., Zhao, Q., Ran, J. et al. The role of hepatocyte epigenetics in the pathogenesis of metabolic dysfunction-associated steatotic liver disease. Commun Med 6, 276 (2026). https://doi.org/10.1038/s43856-026-01651-1
Keywords: fatty liver disease, epigenetics, hepatocytes, DNA methylation, non-coding RNA