Clear Sky Science · en
Identification of intracellular target of antimicrobial peptide EWAMP-R in bacteria
Why a Tiny Worm Molecule Matters for Superbugs
As antibiotic resistance rises, doctors are running out of ways to stop dangerous bacterial infections. This study explores a promising alternative: a small natural molecule, an antimicrobial peptide called EWAMP-R, originally found in earthworms. The researchers show that this peptide does more than poke holes in bacterial membranes—it also sabotages a crucial internal machine that bacteria need to survive, pointing toward a new class of treatments that could stay ahead of resistance.
A Two-Front Attack on Bacteria
Most antibiotics work by targeting a single vital process, such as building the cell wall. Bacteria can eventually evolve around these attacks. Antimicrobial peptides, however, often hit bacteria on multiple fronts. EWAMP-R had already been shown to damage bacterial membranes and trigger a type of programmed death in bacteria, somewhat like apoptosis in human cells. In this study, the team set out to understand what happens inside the cell after EWAMP-R gets past the membrane—specifically, which internal molecule it targets and how that interaction leads to bacterial death.
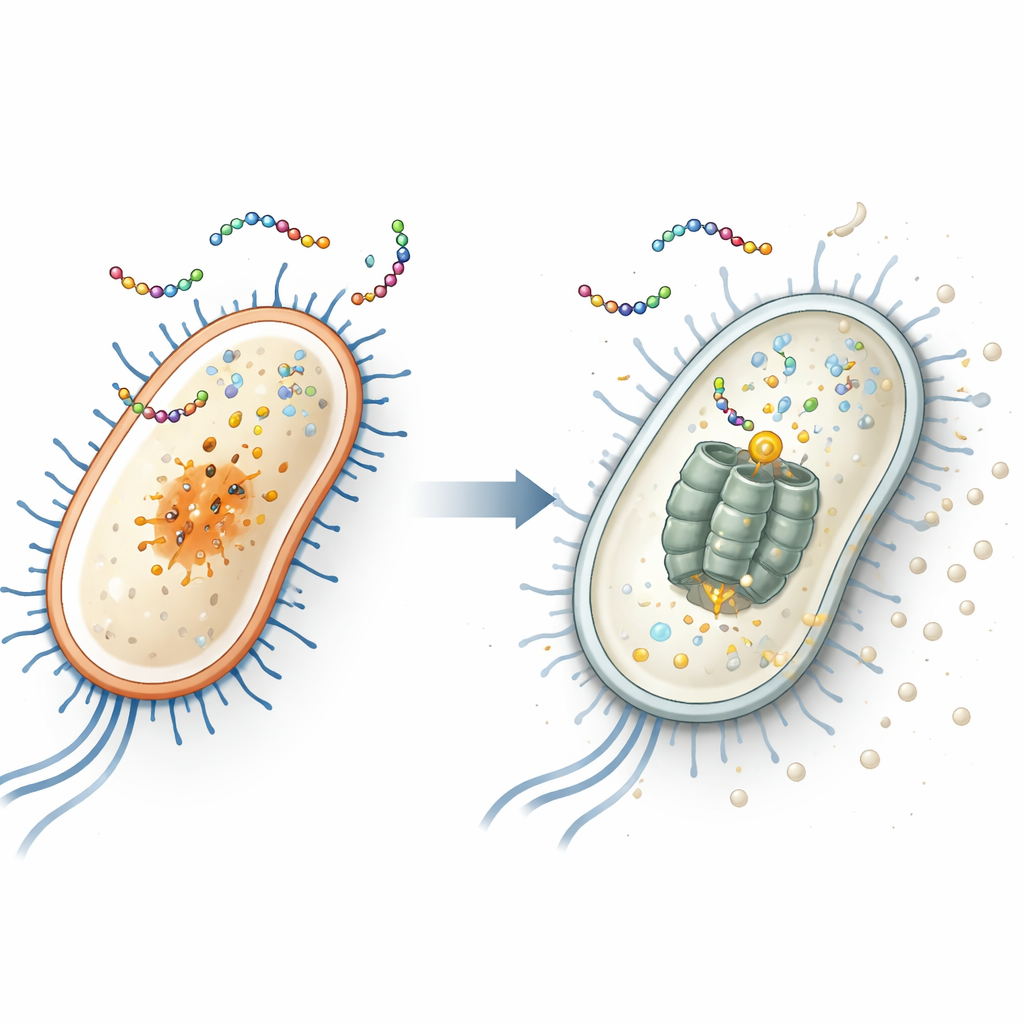
Finding the Hidden Target Inside E. coli
To uncover the peptide’s internal target, the researchers repeatedly exposed two common bacteria—Escherichia coli (a gut bacterium that includes dangerous strains) and Staphylococcus aureus (a frequent cause of skin and hospital infections)—to EWAMP-R until resistant strains emerged. By sequencing the genomes of these survivors, they found that resistant E. coli carried a specific mutation in a gene called clpX. This gene encodes part of a molecular “shredder” that breaks down damaged or regulatory proteins. When they recreated this exact mutation using CRISPR gene editing, E. coli became less sensitive to EWAMP-R, strongly implying that the ClpX protein is the peptide’s key internal target.
Jamming the Bacterial Protein Shredder
Further experiments showed how EWAMP-R interferes with this protein machine. Using binding tests, computer docking, and molecular simulations, the authors found that the peptide nestles into a region of ClpX that normally binds and uses ATP, the cell’s energy currency, to power protein breakdown. When EWAMP-R binds here, it restricts the shape changes ClpX needs to work properly and sharply reduces its ATP-driven activity. One important protein normally processed by the ClpX system is a fragment of a master regulator called LexA, which controls the bacterial SOS response—a stress program that turns on DNA repair genes. When ClpX is blocked, this LexA fragment builds up and continues to clamp down on SOS genes, preventing proper DNA repair and pushing the cell toward an irreversible, apoptosis-like death.
Unexpected Changes in Bacterial Lifestyle
The team also discovered that resistance carries a cost. E. coli that had evolved resistance to EWAMP-R not only carried the clpX mutation but also produced unusually long and numerous flagella—the whip-like tails bacteria use to move. Gene expression patterns confirmed that flagella-related genes were strongly overactive. Previous work suggests that such “hyperflagellated” bacteria may actually become less able to stick to host tissues and more visible to the immune system, potentially reducing their ability to cause disease. In S. aureus, by contrast, resistance did not center on ClpX. Instead, resistant cells slowed down their metabolism, particularly in amino acid and other building-block pathways, growing more slowly and likely becoming less aggressive as pathogens.
Designing Smarter Peptides for Future Medicines
Because EWAMP-R both disturbs bacterial membranes and jams an internal protein machine, it forces bacteria to adapt on multiple levels at once—an energetically costly challenge. Few existing drugs target the ClpX system directly, and this work identifies it as a fresh weak spot in bacteria that affect humans. The authors outline ways to further refine EWAMP-R, such as tweaking specific amino acids to bind ClpX more tightly or help the peptide slip more easily through different bacterial membranes, and using non-natural building blocks to make it more stable in the body. 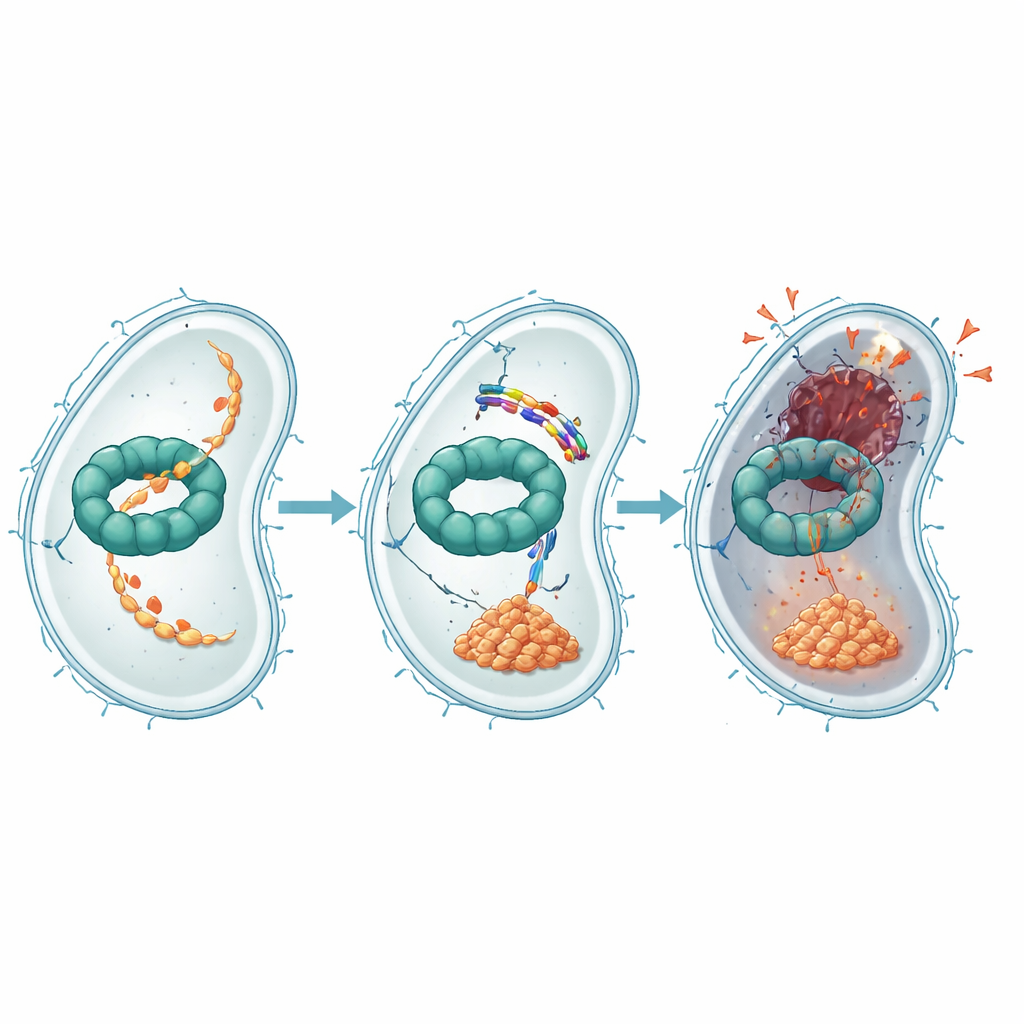
What This Means for Fighting Resistant Infections
In simple terms, this study shows that a small worm-derived peptide can sneak into bacteria and jam a vital internal machine, while also punching holes in their outer shell. By blocking the system that normally clears a key stress regulator, EWAMP-R traps E. coli in a self-destructive state where it cannot properly repair its DNA. Even when bacteria evolve some resistance, they pay a price in slower growth or potentially reduced ability to cause disease. Together, these findings suggest that EWAMP-R, and molecules modeled on it, could inspire a new generation of multi-pronged antibacterial agents that are harder for pathogens to outsmart.
Citation: Wu, Y., Tang, H., Hao, P. et al. Identification of intracellular target of antimicrobial peptide EWAMP-R in bacteria. Commun Biol 9, 564 (2026). https://doi.org/10.1038/s42003-026-09822-5
Keywords: antimicrobial peptides, antibiotic resistance, bacterial stress response, protein quality control, ClpX protease