Clear Sky Science · en
RNA isoform diversity, splicing variants and switching in single cells of the Alzheimer’s disease brain
Why tiny RNA differences matter in Alzheimer’s
Alzheimer’s disease is usually described in terms of plaques, tangles, and dying nerve cells. But beneath those microscope images lies an even finer layer of complexity: the RNA messages that tell each brain cell which proteins to make. This study peeks into that hidden layer at single-cell resolution, showing that not just genes, but the exact "versions" of each gene’s RNA—called isoforms—are extensively altered in Alzheimer’s brains. Understanding these subtle message changes could help explain why certain brain cells falter and may uncover new biomarkers or drug targets.
Looking inside individual brain cells
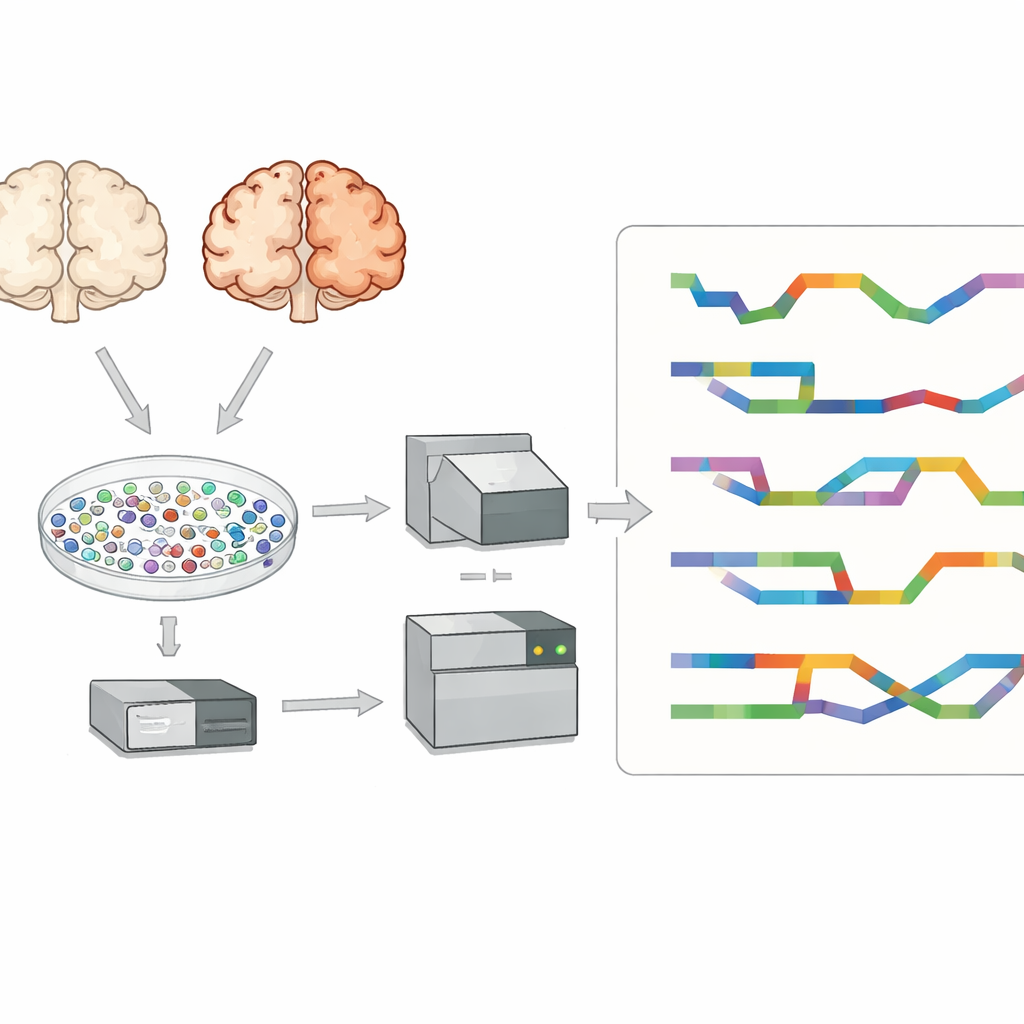
The researchers focused on the prefrontal cortex, a brain region important for thinking and memory that is strongly affected in Alzheimer’s disease. They examined tissue from eight people who had Alzheimer’s and seven who did not. Instead of studying blended tissue, they isolated tens of thousands of individual cell nuclei and captured the RNA within each one. This allowed them to identify major brain cell types—such as excitatory and inhibitory neurons, astrocytes, microglia, oligodendrocytes, and their precursors—and compare Alzheimer’s and non-diseased brains cell type by cell type, rather than as a single mixed signal.
Combining two kinds of sequencing power
Most single-cell studies use short-read sequencing, which reads small snippets of RNA well but struggles to reconstruct full-length transcripts and their exact splice patterns. To overcome this, the team used a long-read technology called PacBio Kinnex alongside standard short-read methods. Kinnex works by linking many cDNA molecules end to end so that a single long-read run captures many full-length RNA copies. After sequencing, these long reads were digitally cut back into individual molecules, then matched to the original cell barcodes. This approach produced around 70,000 single nuclei with both cell-type identity and detailed isoform information, providing an unusually rich view of RNA diversity in the Alzheimer’s brain.
A vast landscape of RNA variants
From these data, the scientists detected more than 850,000 RNA isoforms and then applied strict filters to define about 53,500 "confident" isoforms that were robustly observed across multiple brains. Most belonged to known transcript types, but nearly 4,000 were clearly novel, revealing that even well-studied human cortex still hides unexpected RNA variants. Many genes produced multiple isoforms, with some—such as the dystonin gene—showing dozens of distinct versions. The team also observed rare isoforms with unusual internal junctions, some potentially linked to a process called somatic gene recombination previously described in Alzheimer’s, underscoring how diverse and dynamic brain RNA can be.
Hidden changes that genes alone can’t show
When the scientists compared Alzheimer’s and non-diseased brains at the gene level using short-read data, they saw familiar patterns: inflammatory pathways were more active in microglia and astrocytes, and genes related to synapses and amyloid processing were altered in neurons and oligodendrocytes. But the long-read isoform analysis revealed changes that gene-level averages completely missed. Some genes showed little or no overall difference in expression, yet specific isoforms were strongly increased or decreased or even "switched" in relative abundance in Alzheimer’s. For example, the gene SEPTIN4, involved in the cell’s internal scaffolding and found in protein aggregates in neurodegenerative disease, had one isoform reduced and another increased, canceling one another out at the gene level. Another gene, CHI3L1—encoding the inflammation-linked protein YKL-40—was upregulated in astrocytes mainly through a single novel isoform with retained segments, while the known transcript stayed unchanged. Similar isoform-level shifts were seen for AD-related genes such as APOE, BIN1, APP, and MAPT, often in a cell-type-specific manner.
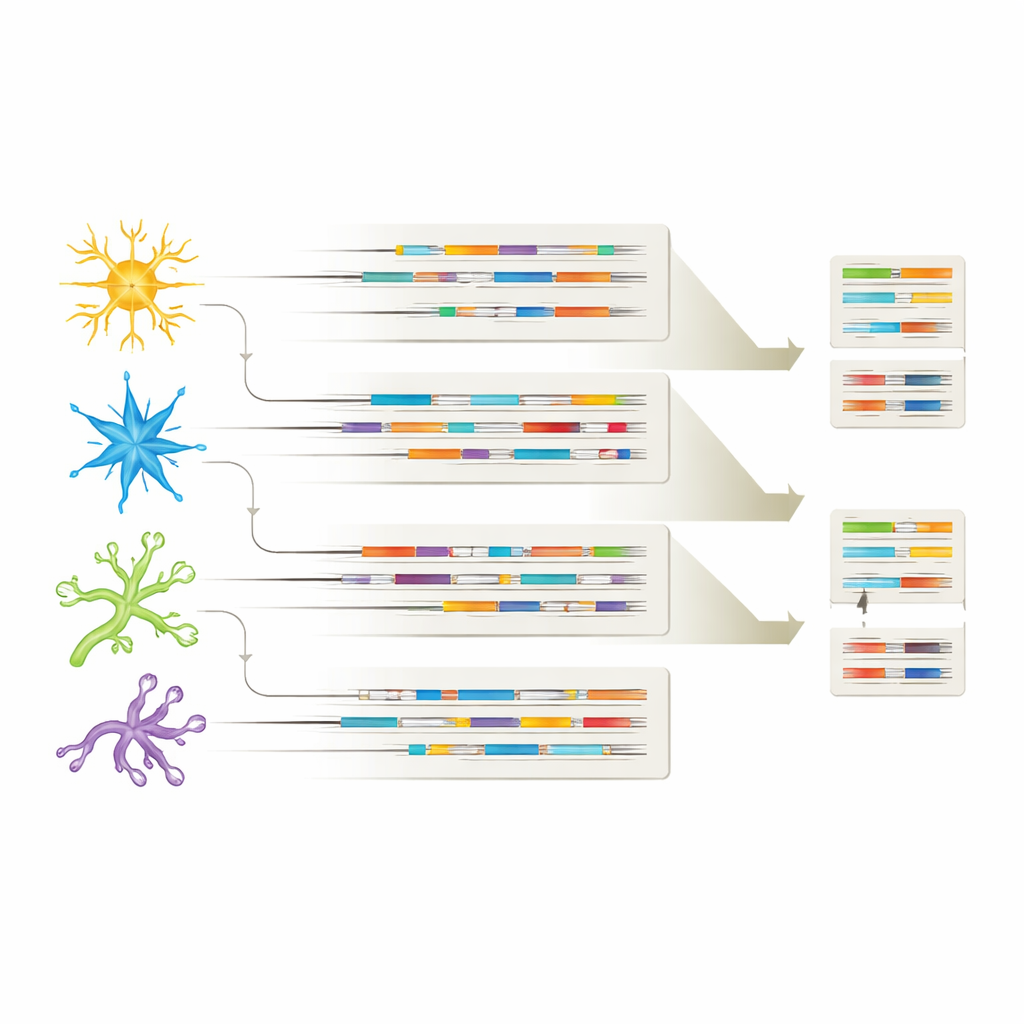
Different cell types, different RNA signatures
By linking each isoform to its cell-type of origin, the study showed that every major brain cell population carries its own distinctive isoform "fingerprint." Many RNA variants were shared across neurons and glial cells, but each cell type also expressed unique isoforms, hinting at specialized roles in health and disease. In some cases, certain isoforms appeared only in Alzheimer’s brains or only in particular cell types, such as specific APP variants in inhibitory neurons or a CHI3L1 isoform limited to astrocytes. These patterns suggest that not only which genes are on or off, but which specific RNA versions are used in each cell type, may shape how Alzheimer’s develops and which cells are most vulnerable.
What this means for understanding and treating Alzheimer’s
This work shows that the Alzheimer’s brain is not just misusing genes, but also remixing the fine structure of its RNA messages in ways that differ across cell types. Long-read single-cell sequencing exposes disease-linked isoform changes that would be invisible in conventional gene-level studies. Some of these newly identified RNA variants, such as altered isoforms of SEPTIN4 and CHI3L1, may play direct roles in brain damage or serve as more precise biomarkers of inflammation and degeneration. While more research is needed to test their functions and clinical usefulness, this study makes clear that future efforts to understand—and eventually treat—Alzheimer’s will need to consider not just genes, but the full diversity of their RNA isoforms in individual brain cells.
Citation: Shahnaee, A., Liu, C.S., Ngo, T. et al. RNA isoform diversity, splicing variants and switching in single cells of the Alzheimer’s disease brain. Commun Biol 9, 506 (2026). https://doi.org/10.1038/s42003-026-09759-9
Keywords: Alzheimer’s disease, RNA splicing, single-cell sequencing, RNA isoforms, neuroinflammation