Clear Sky Science · en
Pan-cancer multi-omic ERBB2-HER2 characterization using next-generation sequencing and quantitative proteomics
Why this matters for cancer patients
Cancer drugs that home in on a molecule called HER2 have transformed treatment for many people with breast and other cancers. Yet doctors still struggle to predict exactly who will benefit. This study shows that the usual genetic and staining tests can miss patients whose tumors truly depend on HER2‑related signaling—and mistakenly flag others who are unlikely to respond. By directly measuring how HER2 and its partner EGFR are switched on inside tumors, the researchers outline a more precise way to match patients with targeted therapies.
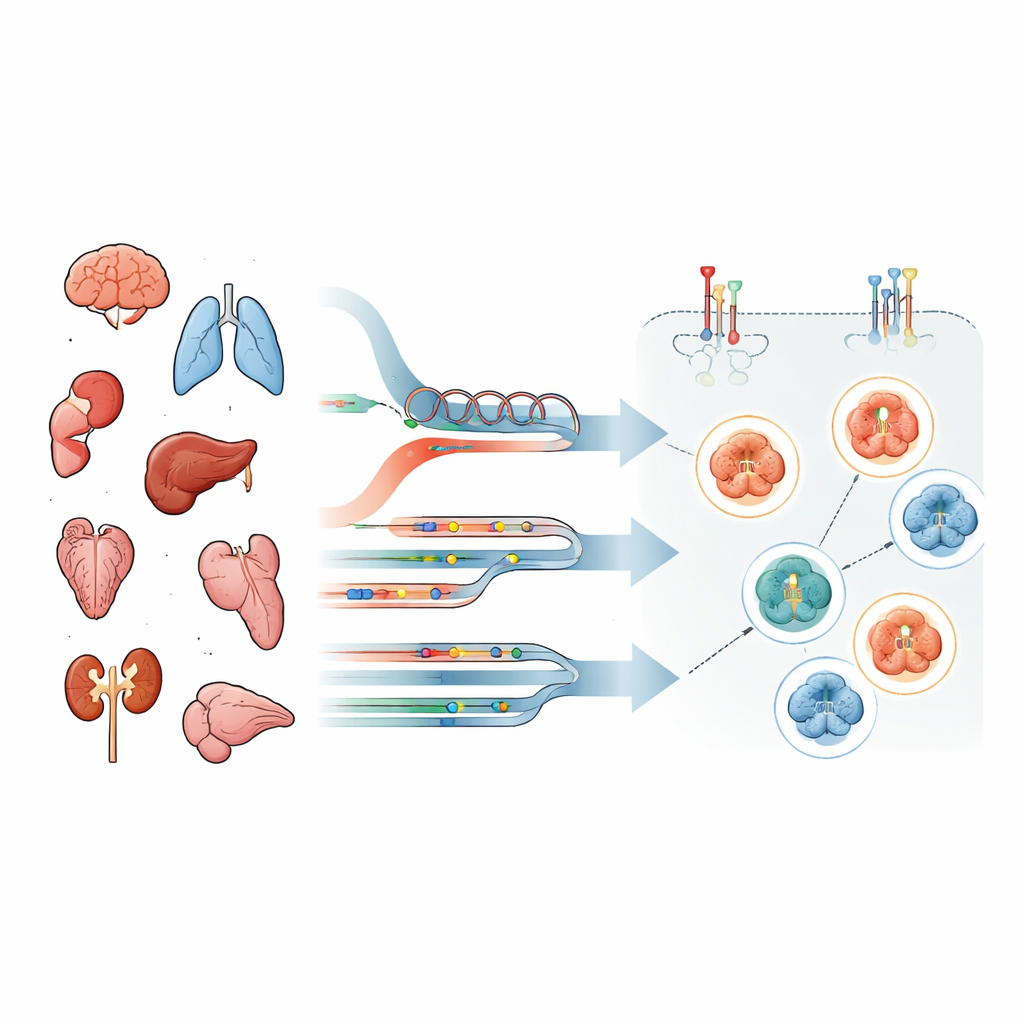
Looking beyond standard lab tests
Most hospitals judge whether a tumor is “HER2‑positive” using two routine tools: a microscope stain that shows how much HER2 protein sits on the cell surface, and DNA tests that detect extra copies or mutations of the HER2 gene (called ERBB2). These approaches focus on how much unmodified HER2 is present, not on whether it is actively sending growth signals. However, targeted drugs fall into different classes. Some antibody–drug conjugates mainly need HER2 to be present as a landing pad, while tyrosine kinase inhibitors work only if the HER2/EGFR signaling switch is truly flipped on. That means abundance alone can be misleading.
A multi‑layer view of tumors
To tackle this problem, the team profiled tumors from 69 people with advanced cancers of many types, including brain, lung, and gastrointestinal tumors. From each patient, they analyzed DNA and RNA with clinical next‑generation sequencing and, crucially, used a highly sensitive technique called reverse‑phase protein arrays on laser‑microdissected tumor cells. This allowed them to quantify not just total HER2 protein but also specific “on” modifications—phosphorylation marks—on HER2 and EGFR and on key signaling proteins downstream. Patients were sorted into groups based on whether their tumors showed HER2 gene changes, RNA overexpression, both, or neither.
When genes and signals disagree
The results revealed a striking mismatch between gene‑level changes and actual pathway activity. Many patients with HER2 or EGFR mutations or extra gene copies showed little or no activating phosphorylation on HER2. Conversely, some patients with no detectable HER2 gene or RNA abnormalities had strongly activated HER2 and EGFR, especially at two key sites that define a pattern called the HARPS signature. Tumors with this co‑activation showed clear engagement of downstream pathways that drive cell growth and survival. In brain cancers, for example, more than half of the glioblastomas showed this activated signature despite lacking classic HER2 DNA alterations, while meningiomas generally did not.
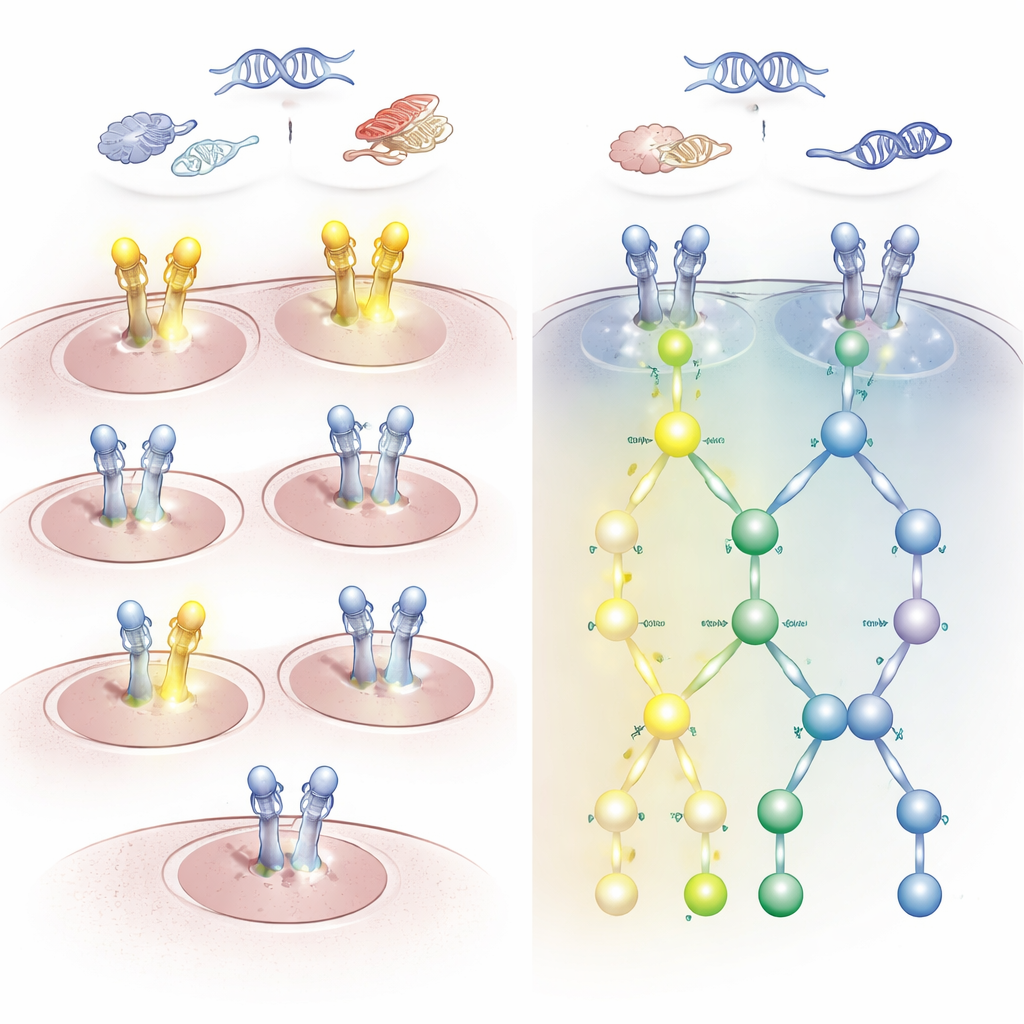
Implications for choosing the right drug
These discrepancies are not just academic. Some patients whose tumors would have been labeled HER2‑negative by standard tests might still respond to HER2‑ or EGFR‑targeted drugs if their signaling pathways are activated. Others who look strongly positive by staining or DNA tests may, in fact, have quiet HER2 signaling and are unlikely to benefit from such treatments. The study also highlighted cases where other driver mutations, such as in KRAS or BRAF, coincided with a lack of HER2/EGFR activation; these patients would probably be better served by therapies aimed at those alternative drivers rather than HER2.
Toward smarter precision oncology
The authors conclude that to truly personalize HER2‑directed therapy, doctors must go beyond DNA sequencing and simple protein abundance. Direct measurements of functional protein activity—especially phosphorylation of HER2 and EGFR—should be integrated into precision oncology workflows. By adding this layer of information, clinicians can more accurately identify who is likely to benefit from kinase inhibitors, antibodies, and antibody–drug conjugates that target the HER family. In everyday terms, it is not enough to know that the switch is installed in the tumor; we must know whether it is actually turned on.
Citation: Hunt, A.L., Randall, J., Ogata, J.D. et al. Pan-cancer multi-omic ERBB2-HER2 characterization using next-generation sequencing and quantitative proteomics. npj Precis. Onc. 10, 153 (2026). https://doi.org/10.1038/s41698-026-01351-y
Keywords: HER2, EGFR, precision oncology, phosphoproteomics, targeted therapy