Clear Sky Science · en
Identification of a LolB-like protein in Porphyromonas gingivalis reveals selective LolA–LolB pairing
Why This Mouth Bacterium Matters
Most people know Porphyromonas gingivalis as a troublemaker in gum disease, but this microbe has also been linked to conditions like Alzheimer’s and rheumatoid arthritis. To survive, it depends on a sophisticated shipping system that moves certain proteins, called lipoproteins, to its outer surface. These lipoproteins help the bacterium build its protective shell and interact with our bodies. Understanding how this transport system works not only illuminates basic bacterial biology, it can also reveal new weak spots for antibiotics that spare our helpful microbes.
The Bacterial Protein Highway
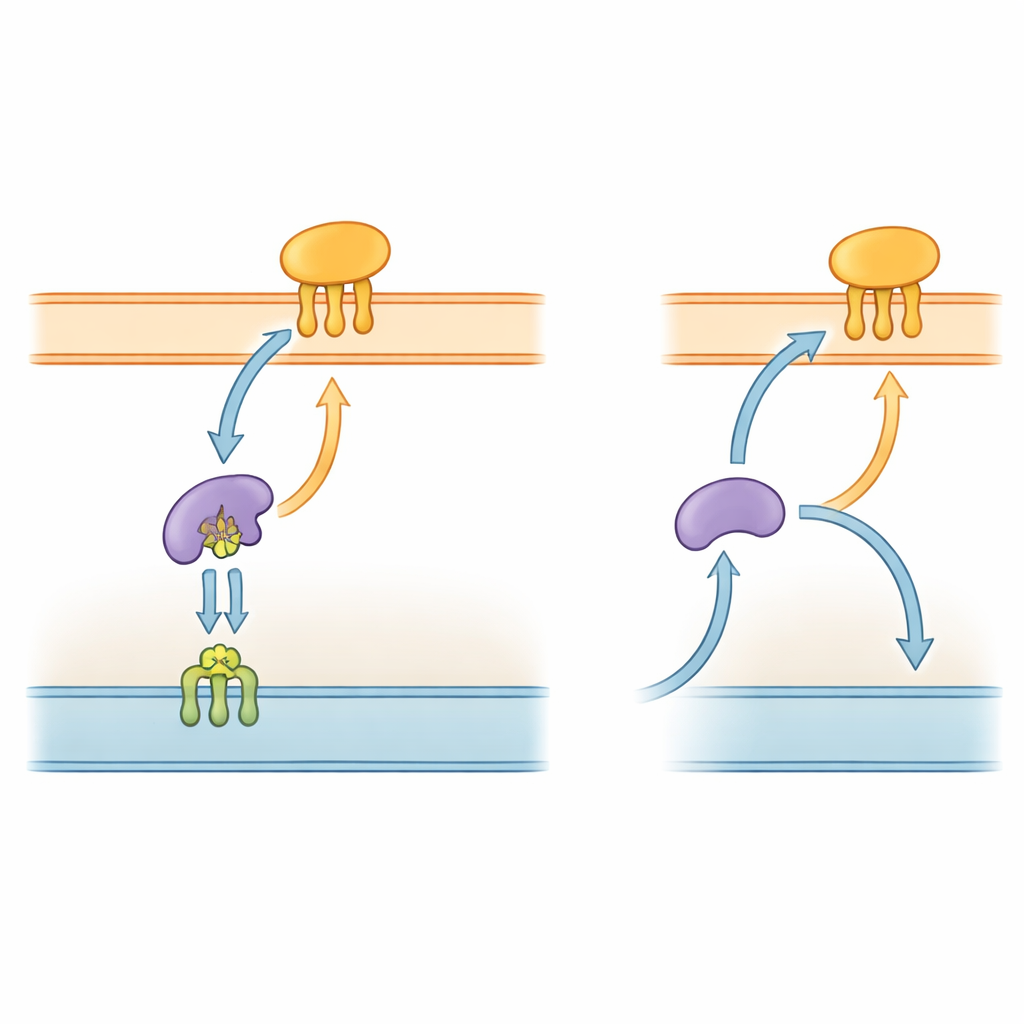
Gram-negative bacteria, including P. gingivalis, are wrapped in a complex envelope with two membranes and a watery space in between. Many of their key tools are lipoproteins—proteins anchored to membranes by short fat-like chains. In well-studied species such as Escherichia coli, a five-part “Lol” system hauls these lipoproteins from the inner membrane, across the periplasmic space, and into the outer membrane. One component, LolA, acts like a shuttle in the periplasm, while LolB in the outer membrane receives the cargo and inserts it into the surface layer. Because this delivery pathway is essential for building the outer membrane, parts of the Lol system are already being explored as targets for new antibiotics.
A Missing Part That Wasn’t Really Missing
For years, researchers thought that P. gingivalis and many of its relatives in the Bacteroidota group lacked LolB altogether. That raised a puzzle: how did these bacteria manage to insert lipoproteins into their outer membrane? Using modern structure-prediction tools (such as AlphaFold) and follow-up experiments, the authors identified a long-sought candidate in P. gingivalis, a protein encoded by the gene pgn0994. They showed that this protein, which they call LolB-PG, folds in almost the same way as known LolB proteins from other bacteria, even though its amino acid sequence is quite different. Its three-dimensional structure revealed a curved sheet forming a deep groove, a shape well suited to cradle the fatty tail of a lipoprotein.
A Tailor-Made Molecular Partnership
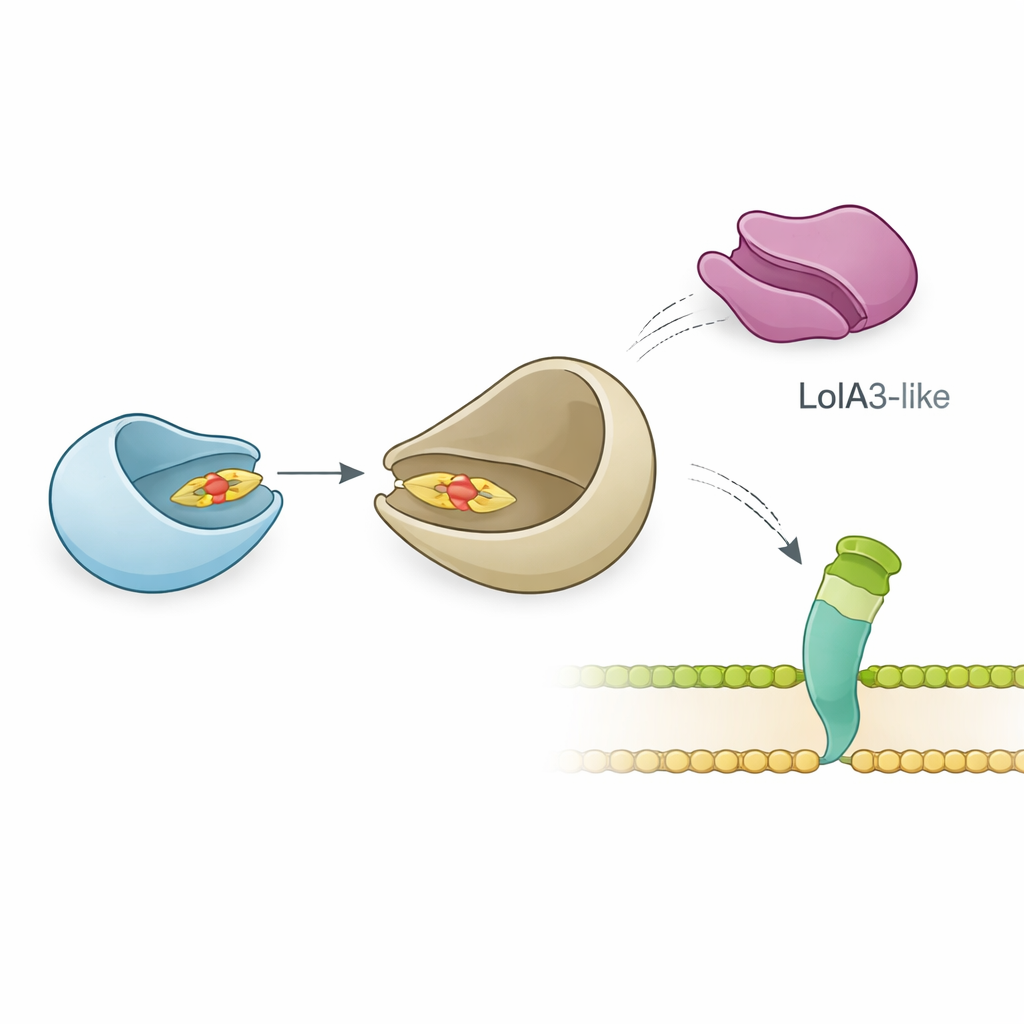
The team went on to examine how LolB-PG interacts with its likely partner, LolA, which ferries lipoproteins through the periplasm. Precise biophysical measurements demonstrated that LolB-PG binds strongly and specifically to the main LolA protein of P. gingivalis, with an affinity similar to classic LolA–LolB pairs in E. coli and other bacteria. In contrast, LolB-PG barely interacts with a second LolA-like protein in P. gingivalis, called LolA3, or with LolA from other species. Structural modeling suggested why: LolB-PG and the primary LolA dock in a “mouth-to-mouth” fashion, aligning their grooves so the lipid tail can slide from one to the other, whereas LolA3 presents a mismatched surface. These results point to a selective pairing where each protein has a dedicated role rather than a one-size-fits-all interaction.
Surprising Flexibility in Surface Assembly
Given this clear partnership, the researchers expected that removing LolB-PG would seriously disrupt the bacterium’s outer membrane. To test this, they deleted the pgn0994 gene and examined whether the bacterium could still build its long surface fibers, called fimbriae, which are partly made from lipoproteins. Surprisingly, the mutant strain grew normally and formed fimbriae that looked and behaved much like those in the unmodified bacteria. Key fimbrial components were still properly processed, transported to the envelope, and assembled into polymers. This suggests that, at least under the tested laboratory conditions, P. gingivalis can use alternative routes to move these lipoproteins, or that other systems step in to compensate when LolB-PG is missing.
What This Means for Future Treatments
This work shows that P. gingivalis does, in fact, carry a functional LolB-like protein that forms a specific partnership with its main LolA chaperone, preserving a fundamental lipoprotein transport step seen across many bacteria. At the same time, the enlarged binding groove of LolB-PG, its selective pairing, and the lack of obvious defects when the protein is removed all point to a more flexible and diversified transport network than previously thought. For non-specialists, the take-home message is that even highly conserved bacterial systems can be reworked in different species, which may influence how well antibiotic strategies translate from one pathogen to another. Mapping these variations in detail will be crucial for designing drugs that can disarm harmful bacteria like P. gingivalis without broadly harming the beneficial microbes we rely on.
Citation: Jaiman, D., Hirohata, M., Hasegawa, Y. et al. Identification of a LolB-like protein in Porphyromonas gingivalis reveals selective LolA–LolB pairing. Sci Rep 16, 13157 (2026). https://doi.org/10.1038/s41598-026-49975-1
Keywords: lipoprotein transport, Porphyromonas gingivalis, LolA LolB system, bacterial outer membrane, antibiotic targets