Clear Sky Science · en
Ependymoma group-specific blood-brain barrier differences uncovered by a multi-omics approach
Why the Brain’s Gatekeepers Matter for Kids with Cancer
The human brain is fiercely protected by a natural security system called the blood–brain barrier, a tight layer of cells that strictly controls which substances can enter brain tissue from the bloodstream. This barrier is vital for health, but it also makes it much harder to get cancer drugs into brain tumors—especially in children. This study asks a deceptively simple question with big clinical implications: do different subtypes of a pediatric brain tumor called ependymoma live behind different kinds of barriers, and could that help explain why many drugs fail?
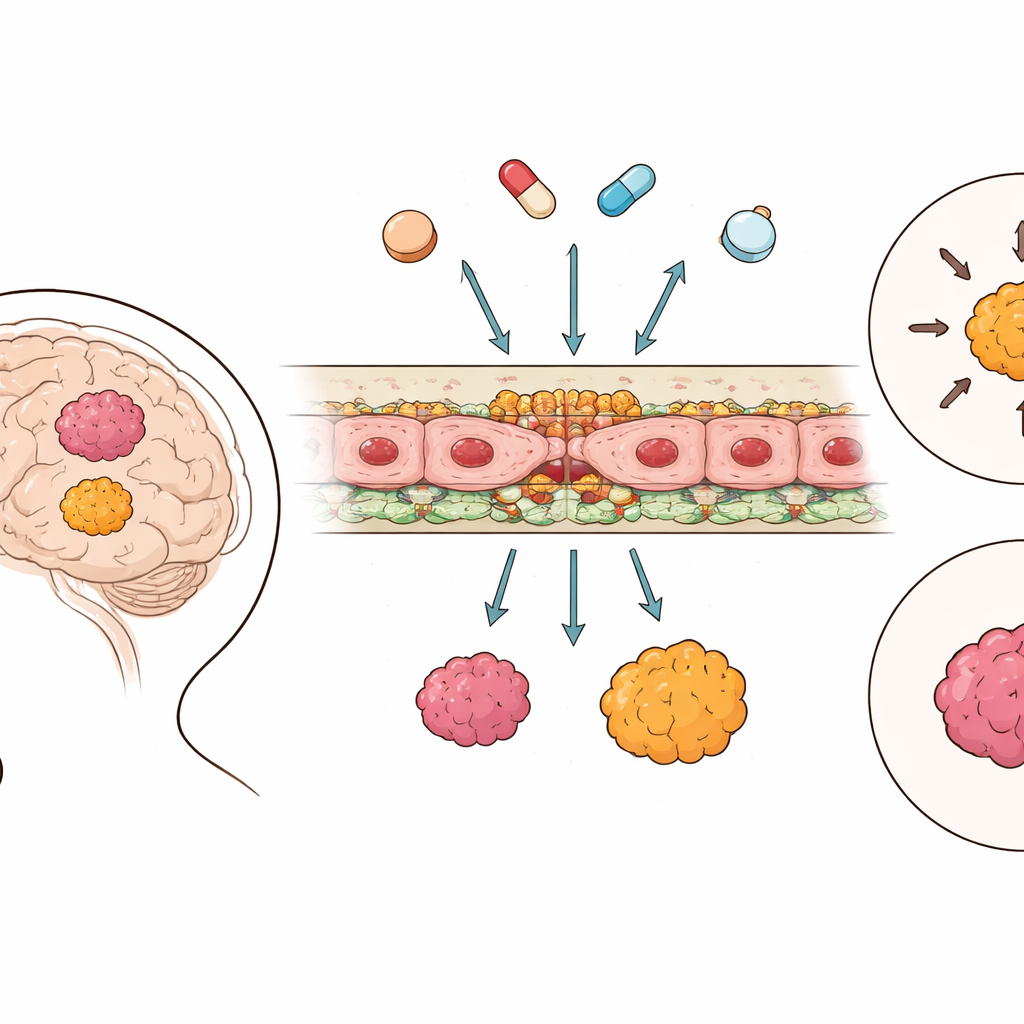
The Brain’s Protective Wall
The blood–brain barrier is built from cells lining tiny brain vessels that are locked together by specialized junctions, and equipped with molecular “pumps” that can kick drugs back into the bloodstream. Using large-scale genetic data from 364 ependymoma tumors and 225 samples of healthy brain, the researchers cataloged the activity of genes involved in these junctions and in transport systems. They found that, unlike in many other brain cancers, ependymoma tissue actually showed higher activity of key junction components than healthy brain, suggesting the barrier between blood and tumor is often tighter rather than leakier. In contrast, drug-efflux pumps and other transporters followed more mixed, group-specific patterns instead of a simple on-or-off trend.
Not Just Location or Age, but Tumor Type
Next, the team asked whether normal differences in brain development or anatomy might explain these patterns. By training neural-network scores that summarize junction and transporter gene activity, they confirmed known features in healthy brains: the barrier tightness shifts with age and differs between regions such as the cortex and cerebellum. However, these age and regional trends did not carry over into the tumors. Across the major intracranial ependymoma groups, barrier-related scores looked similar regardless of where in the brain the tumor arose or how old the patient was. This points to the molecular identity of the tumor itself—its subgroup—as the main driver of how its local barrier behaves.
Zooming in on the Barrier Cells
Bulk measurements can mask which cells are actually responsible for the observed gene activity. To resolve this, the researchers turned to single-cell and single-nucleus RNA sequencing from multiple patient cohorts and a new validation set. These finer-grained data revealed that most barrier-related genes are concentrated in a distinct subset of blood-vessel cells within the tumor microenvironment. This specialized endothelial subpopulation showed strong expression of classic junction genes such as CLDN5 and other barrier markers, while tumor cells, immune cells and support cells expressed very little. The findings were bolstered by protein measurements and advanced imaging, which showed these proteins physically lining the tumor blood vessels, confirming that the genetic signatures reflect a real structural barrier.
Testing the Barrier in Mouse Models
Because new drugs are often evaluated in mouse models before reaching patients, the team probed whether patient-derived xenografts of different ependymoma groups reproduced human barrier features. In these models, tumor cells from patients grow in the brains of mice, where the blood vessels are largely mouse-derived. Analyses of RNA and protein again showed strong junction expression in vessel cells, but with some differences between models and between tumor and surrounding brain. Imaging with contrast agents and fluorescent tracers suggested that, overall, the barrier in these tumors remained relatively intact, and that subtle differences between models and subgroups could influence how easily molecules pass from blood to tumor tissue.
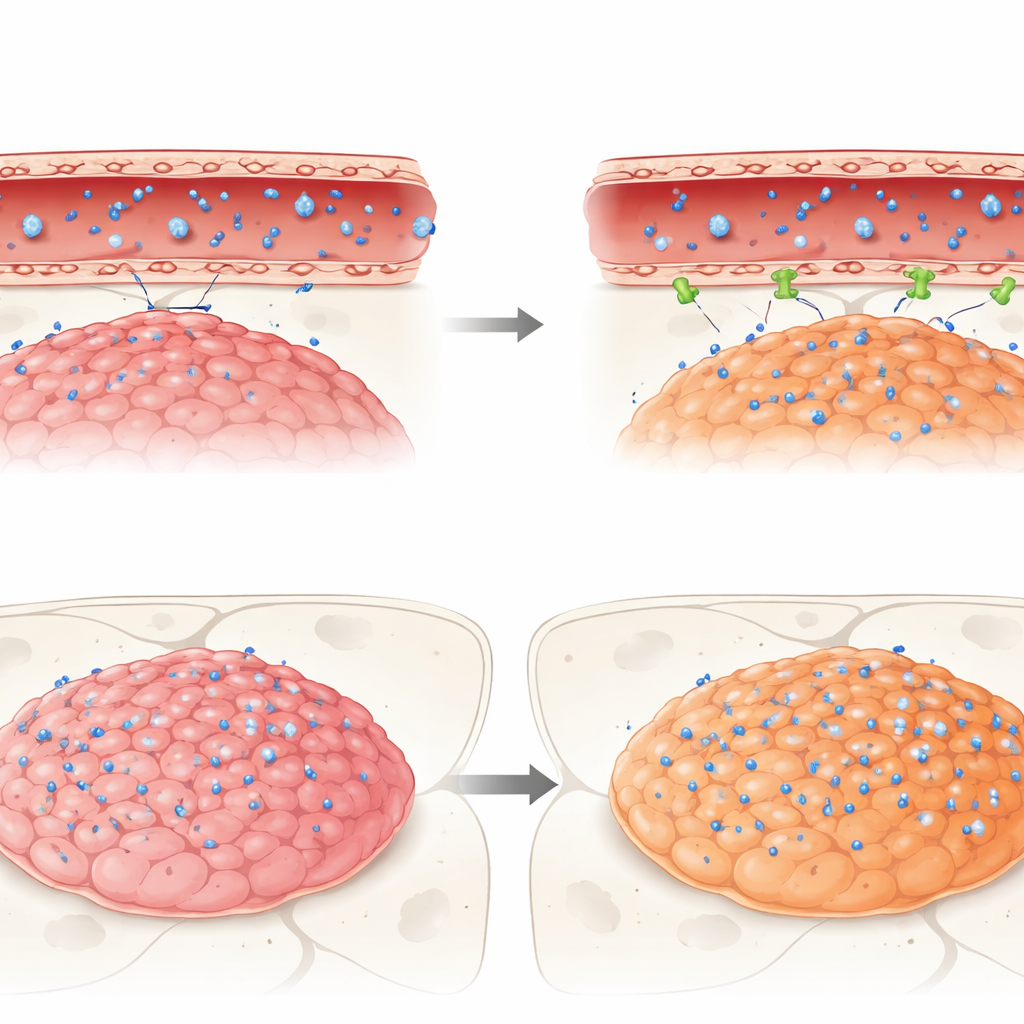
What Happens When Real Drugs Try to Cross
Finally, the researchers followed the journey of three clinically relevant anti-cancer drugs—idasanutlin, temsirolimus and etoposide—into these mouse tumors. All three were active against ependymoma cells in dishes, but in living brains they faced the barrier. Idasanutlin, although chemically suited to cross cell membranes, reached very low levels in both tumor and normal brain compared to blood, hinting at strong exclusion by the barrier. In contrast, temsirolimus and etoposide, both known to be expelled by certain pumps, accumulated more in tumors belonging to one aggressive subgroup (ZFTA fusion–positive) than in another group (PFA) or the adjacent healthy brain. Even so, none of the drugs reached concentrations in tumors that matched their effective levels seen in vitro, underscoring how the barrier can quietly undermine promising therapies.
What This Means for Future Treatments
To a layperson, the central message is straightforward: ependymoma tumors do not all sit behind the same kind of brain gate. Instead, each molecular subgroup comes with its own version of the blood–brain barrier—often surprisingly tight—which shapes how well different drugs can reach their targets. By combining multiple layers of data from patients and mouse models, this work shows that understanding and measuring these subgroup-specific barrier traits is essential for designing smarter clinical trials. Rather than assuming that a drug that kills tumor cells in a dish will work in a child’s brain, future therapies may need to be chosen—and possibly paired with barrier-modifying strategies—based on how each tumor’s gatekeepers are built.
Citation: Sundheimer, J.K., Benzel, J., Federico, A. et al. Ependymoma group-specific blood-brain barrier differences uncovered by a multi-omics approach. Sci Rep 16, 12061 (2026). https://doi.org/10.1038/s41598-026-47499-2
Keywords: ependymoma, blood-brain barrier, pediatric brain tumors, drug delivery, multi-omics