Clear Sky Science · en
Associations of non-HLA gene polymorphisms with celiac disease in India
Why this matters for everyday diets
Celiac disease turns a common food staple, wheat, into a trigger for chronic gut injury. This study asks a simple but important question: beyond the best known risk genes, which other inherited traits help decide who in India gets sick from eating gluten and who stays healthy, even when both groups carry similar main risk genes?
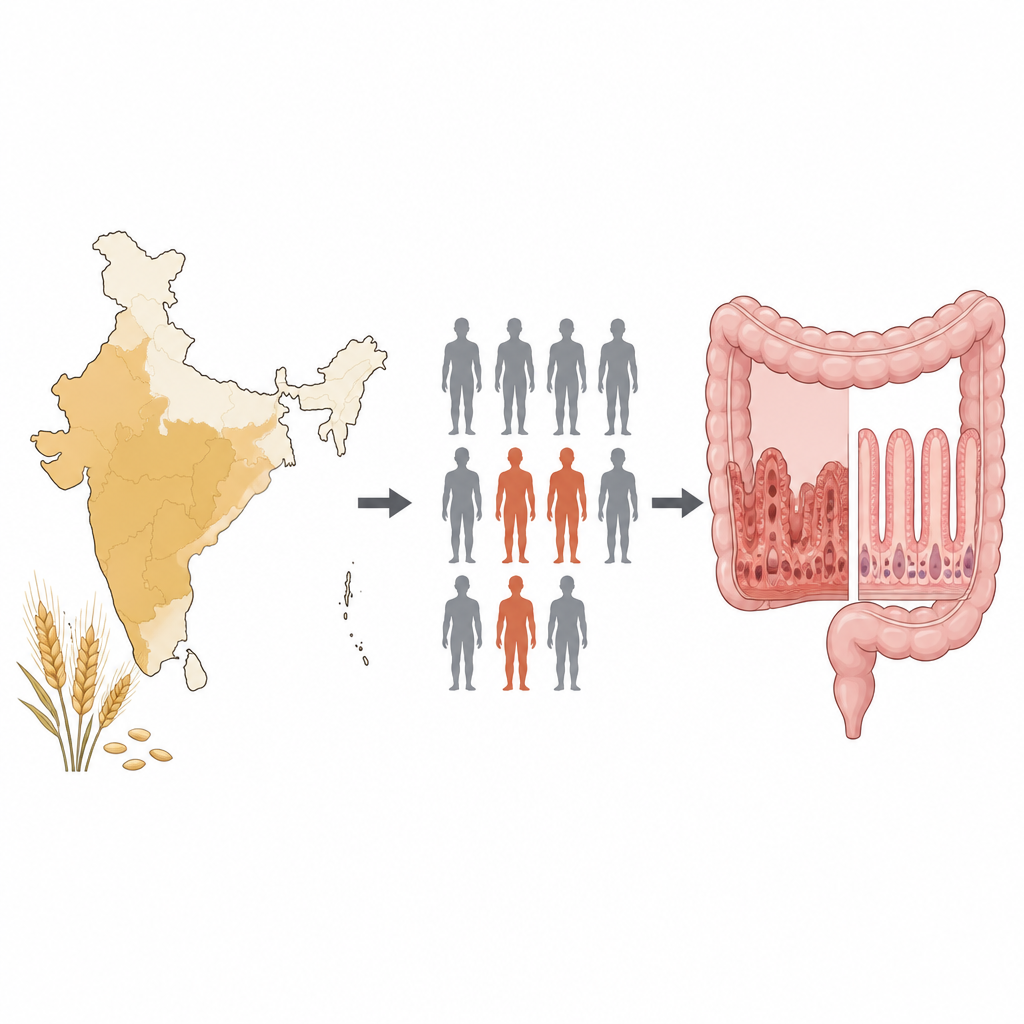
A closer look at gluten and the gut
Celiac disease is an immune condition in which eating gluten, a protein in wheat, barley and rye, damages the small intestine. It affects around 1 percent of people in Europe, North America and wheat eating regions of India. The main known genetic risk lies in a family of immune system genes called HLA DQ, which help immune cells recognize bits of gluten. People who carry certain forms, especially those called DQ2.5, DQ2.2 and DQ8, are much more likely to develop the disease. Yet many people with these genes never become ill, which suggests that other inherited factors also shape risk.
What the researchers set out to find
The team studied 376 adults with celiac disease and 736 adults without the disease from different parts of India. From earlier large genetic surveys, they chose 63 small DNA changes, known as single nucleotide polymorphisms, in genes suspected to affect the immune response. First they checked which of these changes are even present at useful levels in people of Indian ancestry. After this screening step, 51 genetic sites in both the main HLA region and many non HLA genes were tested for how often each version appeared in patients compared with healthy controls.
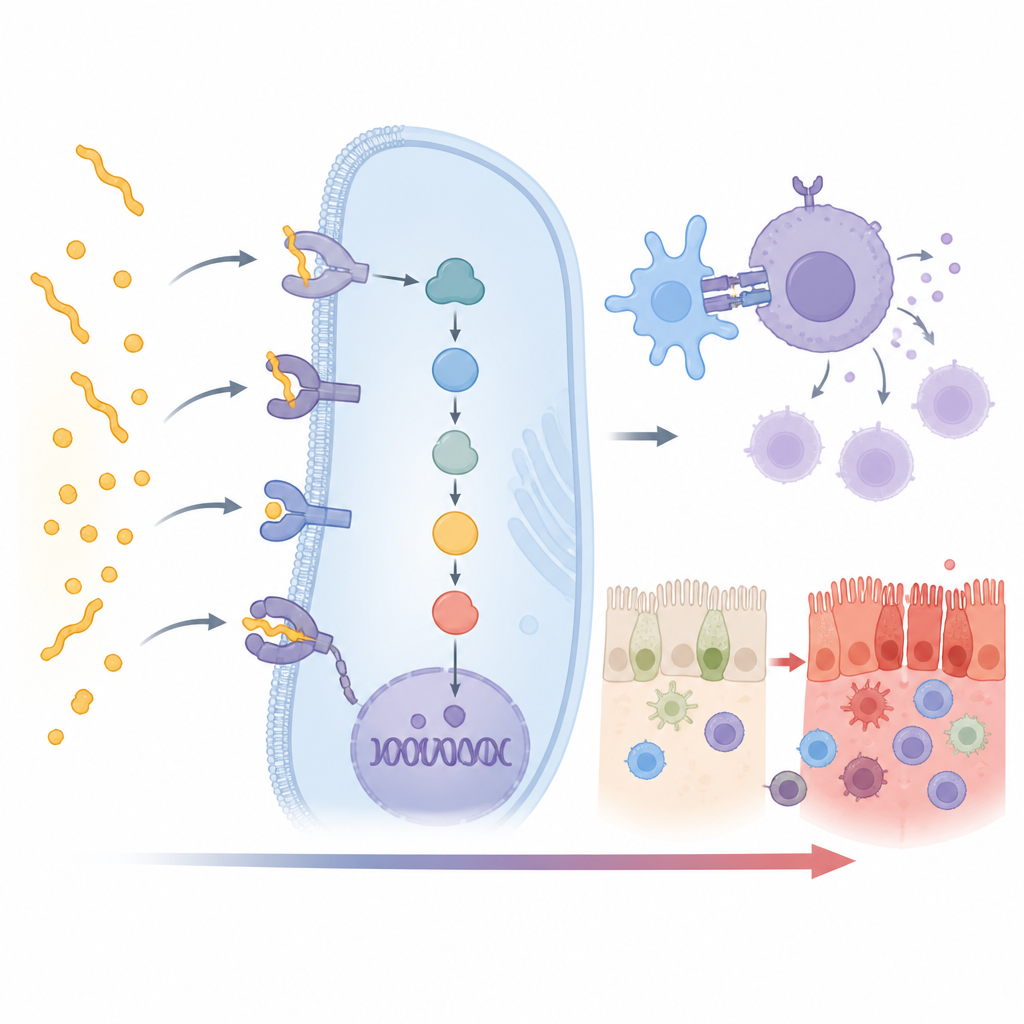
Hidden helpers in the immune response
As expected, the strongest signal came from the HLA DQ2.5 form, with weaker but still clear links to DQ2.2 and DQ8. More intriguingly, 15 additional non HLA genes also showed a strong link to celiac disease in this Indian group. Many sit directly in the chain of events that turns gluten recognition into full blown immune attack. Some genes, such as CD247, CD28, CD80 and PRKCQ, help switch on T cells, the white blood cells that drive the disease. Others, like UBASH3A and TNFAIP3, act as brakes that normally keep this activation from running out of control. Changes in these genes may tilt the balance toward either stronger or weaker immune reactions.
Protecting the gut lining and guiding immune cells
Several of the associated genes affect the health of the intestinal lining itself or the movement of immune cells. CCR3 and CCR4 help guide immune and inflammatory cells into tissues, potentially steering more such cells into the small intestine where gluten is present. GATD3 and PARK7 are linked to how mitochondria handle stress and may help gut cells cope with damage caused by inflammation and reactive oxygen. INAVA supports the tight junctions that keep the intestinal barrier sealed, while DDX6, PUS10 and a long non coding RNA called LINC01934 influence how RNA is processed and how genes are switched on and off. Together, these changes outline a chain that leads from gluten contact, through immune signaling, to injury of the gut surface.
Piecing together the genetic network
By analyzing how the different DNA changes interact with each other, the researchers found a tightly linked network of sites rather than many isolated effects. Certain variants emerged as hubs, showing many statistical connections with others. When the team mapped the corresponding genes, hub roles were taken by key immune and signaling genes, including several HLA class II genes, chemokine receptors that control cell migration, and T cell activation molecules. This network pattern supports the view that many small genetic nudges along the same pathway combine to determine whether gluten exposure tips a person into disease.
What this means for people at risk
For readers, the main takeaway is that celiac disease risk in India is not set by a single gene but by a cluster of genes that talk to each other in immune cells and in the gut lining. The classic HLA risk types remain essential, but additional inherited differences in how T cells are switched on, how signals are damped down, how immune cells move, and how the intestinal wall holds together all add to the final outcome. As these pathways are better understood, they could help refine genetic risk scores for people who already carry high risk HLA types and may eventually point to new ways to calm the immune response to gluten without relying only on strict lifelong diet changes.
Citation: Ramakrishna, B.S., Singh, A., Srinivasan, P. et al. Associations of non-HLA gene polymorphisms with celiac disease in India. Sci Rep 16, 14918 (2026). https://doi.org/10.1038/s41598-026-45159-z
Keywords: celiac disease, gluten, HLA-DQ genes, T cell activation, genetic risk