Clear Sky Science · en
The impact of PD-1 mutations on pembrolizumab binding: insights from molecular dynamics and MM-GBSA analysis
Why this matters for cancer treatment
Immunotherapy drugs like pembrolizumab have transformed care for many cancers, yet only a fraction of patients benefit—and some who respond at first later relapse. This study asks a focused question behind that clinical mystery: what happens if subtle changes occur in PD-1, the immune “off switch” that pembrolizumab is designed to block? By using advanced computer simulations rather than patient samples, the authors map how rare genetic alterations in PD-1 could weaken or strengthen the drug’s grip, offering clues to why treatment sometimes fails and how future tests or drugs might do better.
The immune brake that tumors exploit
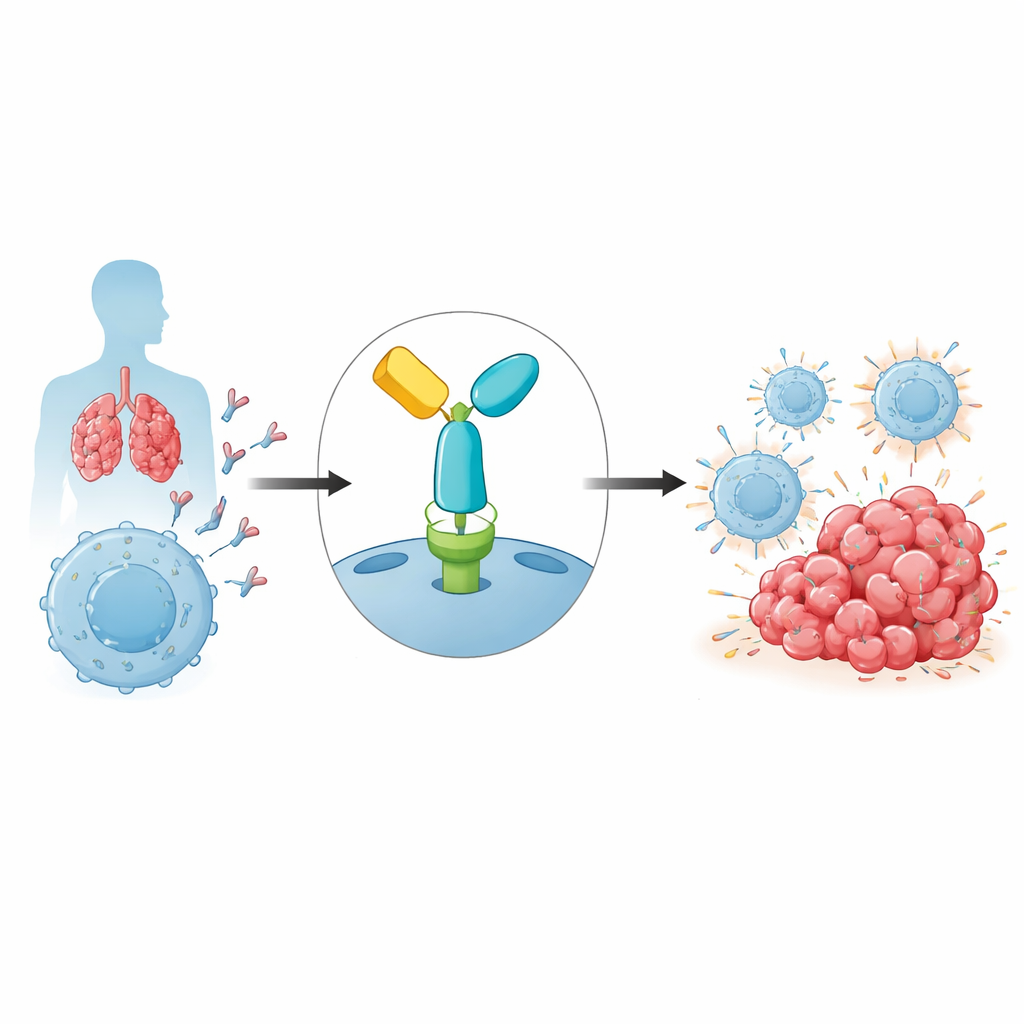
Our immune system relies on T cells to recognize and attack abnormal cells, including cancer. To avoid collateral damage, T cells carry safety switches called immune checkpoints. One of the most important is PD-1, a receptor on T cells that, when engaged by its partner PD-L1 on other cells, dampens immune activity. Tumors frequently exploit this pathway by displaying PD-L1, effectively telling nearby T cells to stand down. Pembrolizumab is an antibody drug that binds to PD-1 and blocks this inhibitory interaction, reactivating T cells so they can attack tumor cells more effectively. Although this strategy has led to long-lasting remissions in cancers such as melanoma and lung cancer, resistance is common, and the reasons are still being unraveled.
Using computers as a molecular microscope
Instead of running lab experiments on cells, the researchers turned to detailed computer models of the PD-1 protein locked together with pembrolizumab. They began with a high-resolution crystal structure of this complex and identified the key contact points—individual amino acids on PD-1 that form hydrogen bonds, grease-like hydrophobic contacts, or ionic “salt bridges” with the drug. Then, using molecular dynamics simulations, they watched how these contacts behaved over 50 billionths of a second, and calculated how much each residue contributed to keeping the complex intact. This allowed them to pinpoint 16 “hot spot” positions on PD-1 that are especially important for strong binding.
Rare genetic changes with outsized effects
Next, the team searched a large protein database for naturally occurring changes (mutations) in those hot spot positions and found 40 variants, plus one additional mutation previously reported to disrupt pembrolizumab binding in lab tests. After filtering out variants predicted to damage PD-1’s overall structure, they were left with 28 “likely benign” mutations—changes that should leave PD-1 functional but might subtly alter how it interacts with drugs. For each of these, they built a three-dimensional mutant model and ran the same type of molecular dynamics simulation, followed by an energy calculation method (MM-GBSA) to estimate how strongly pembrolizumab would bind compared with the normal version.
When key anchors are lost
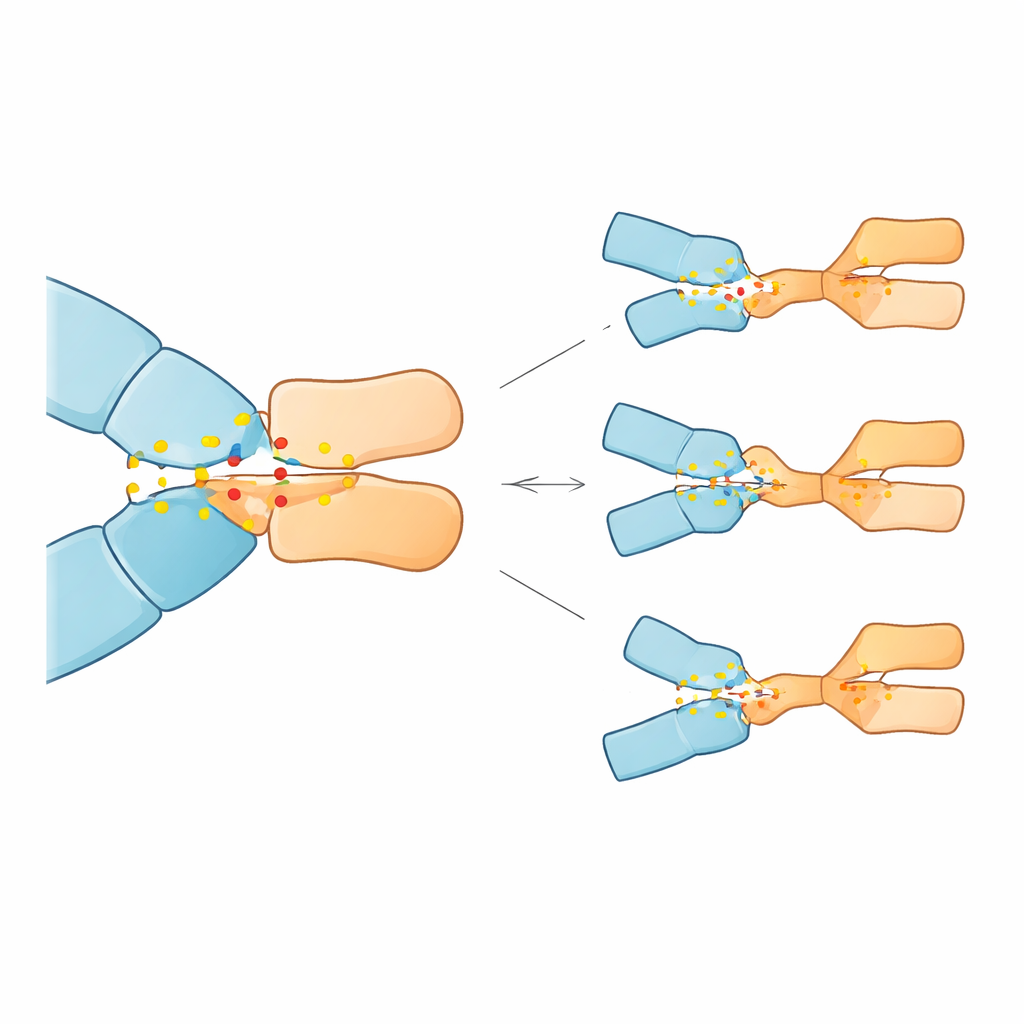
The simulations revealed that 23 of the 28 mutations weakened pembrolizumab binding, some quite dramatically, while five actually strengthened it. Three mutations stood out for causing the largest predicted loss of affinity: substitutions at positions 85 and 89 of PD-1 (Asp85Gly, Asp85Asn, and Pro89Arg). In the normal protein, Asp85 acts as an electrostatic anchor, forming a robust salt bridge and hydrogen bonds with an arginine on the antibody; Pro89 helps form a cluster of snug hydrophobic contacts with several antibody residues. The Asp85 mutations removed the negative charge or hydrogen-bonding ability at this site, effectively breaking the anchor and destabilizing the surrounding network of contacts. The Pro89Arg change replaced a neutral, shape-fitting residue with a positively charged one, disrupting the hydrophobic cluster and reducing its stabilizing contribution. In contrast, a few other mutations at different positions subtly reshaped the interface in ways that could enhance binding, at least in the model.
What this could mean for patients
These findings suggest that even very rare changes in the PD-1 gene, if they occur in crucial contact points, could reduce how well pembrolizumab latches onto its target and thus blunt its ability to unleash T cells against cancer. While the work is entirely computational and needs experimental and clinical confirmation, it offers a roadmap: specific PD-1 variants such as those at positions 85 and 89 might someday serve as biomarkers to flag patients at higher risk of resistance or to guide the choice among different PD-1–targeting drugs. More broadly, the study illustrates how in-depth computer simulations can illuminate the fine-grained molecular reasons why powerful immunotherapies succeed in some people but falter in others.
Citation: Bouricha, E.M., Hakmi, M., Batlamous, B. et al. The impact of PD-1 mutations on pembrolizumab binding: insights from molecular dynamics and MM-GBSA analysis. Sci Rep 16, 10773 (2026). https://doi.org/10.1038/s41598-026-45077-0
Keywords: PD-1 mutations, pembrolizumab resistance, cancer immunotherapy, immune checkpoint inhibitors, molecular dynamics simulation